Abstract
Social science genetics is concerned with understanding whether, how and why genetic differences between human beings are linked to differences in behaviours and socioeconomic outcomes. Our review discusses the goals, methods, challenges and implications of this research endeavour. We survey how the recent developments in genetics are beginning to provide social scientists with a powerful new toolbox they can use to better understand environmental effects, and we illustrate this with several substantive examples. Furthermore, we examine how medical research can benefit from genetic insights into social-scientific outcomes and vice versa. Finally, we discuss the ethical challenges of this work and clarify several common misunderstandings and misinterpretations of genetic research on individual differences.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
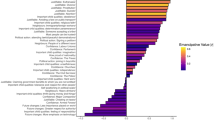
The genetic correlations reported in Fig. 1 are based on publicly available GWAS summary statistics on LDHub (http://ldsc.broadinstitute.org/ldhub/). The Health and Retirement Study data in Fig. 3 can be accessed via dbGaP and the University of Michigan.
References
Definition of social science. Merriam Webster Dictionary https://www.merriam-webster.com/dictionary/social%20science (Accessed 1 November 2018).
Turkheimer, E. Three laws of behavior genetics and what they mean. Curr. Dir. Psychol. Sci. 9, 160–164 (2000).
Polderman, T. J. C. et al. Meta-analysis of the heritability of human traits based on fifty years of twin studies. Nat. Genet. 47, 702–709 (2015).
Benjamin, D. J. et al. The promises and pitfalls of genoeconomics. Annu. Rev. Econ. 4, 627–662 (2012).
Visscher, P. M. et al. 10 years of GWAS discovery: Biology, function, and translation. Am. J. Hum. Genet. 101, 5–22 (2017).
Buniello, A. et al. The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 47D1, D1005–D1012 (2019).
Freese, J. The arrival of social science genomics. Contemp. Sociol. 47, 524–536 (2018).
Comfort, N. Nature still battles nurture in the haunting world of social genomics. Nature 553, 278–280 (2018).
Turkheimer, E. & Paige Harden, K. Behavior Genetic Research Methods. in Handbook of Research Methods in Social and Personality Psychology 159–187 (Cambridge University Press, 2014).
Kong, A. et al. The nature of nurture: Effects of parental genotypes. Science 359, 424–428 (2018).
Koellinger, P. D. & Harden, K. P. Using nature to understand nurture: Genetic associations show how parenting matters for children’s education. Science 359, 386–387 (2018).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Servick, K. Can 23andMe have it all? Science 349, 1472–1474, 1476–1477 (2015).
Duncan, L. E., Pollastri, A. R. & Smoller, J. W. Mind the gap: why many geneticists and psychological scientists have discrepant views about gene-environment interaction (G×E) research. Am. Psychol. 69, 249–268 (2014).
Reich, D. E. et al. Linkage disequilibrium in the human genome. Nature 411, 199–204 (2001).
Abecasis, G. R. et al. The 1000 Genomes Project Consortium. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Visscher, P. M., Brown, M. A., McCarthy, M. I. & Yang, J. Five years of GWAS discovery. Am. J. Hum. Genet. 90, 7–24 (2012).
Chabris, C. F., Lee, J. J., Cesarini, D., Benjamin, D. J. & Laibson, D. I. The fourth law of behavior genetics. Curr. Dir. Psychol. Sci. 24, 304–312 (2015).
Rapid GWAS of thousands of phenotypes for 337,000 samples in the UK Biobank. Neale Lab http://www.nealelab.is/blog/2017/7/19/rapid-gwas-of-thousands-of-phenotypes-for-337000-samples-in-the-uk-biobank (Accessed 6 November 2018).
Kyoko Watanabe, D.P. Atlas of GWAS Summary Statistics. GWAS Atlas (2017). http://atlas.ctglab.nl/ (Accessed 6 November 2019).
Global Biobank Engine. https://biobankengine.stanford.edu/ (Accessed 6 November 2019).
Manolio, T. A. et al. Finding the missing heritability of complex diseases. Nature 461, 747–753 (2009).
Sohail, M. et al. Polygenic adaptation on height is overestimated due to uncorrected stratification in genome-wide association studies. eLife 8, e39702 (2019).
Berg, J. J. et al. Reduced signal for polygenic adaptation of height in UK Biobank. eLife 8, e39725 (2019).
Haworth, S. et al. Apparent latent structure within the UK Biobank sample has implications for epidemiological analysis. Nat. Commun. 10, 333 (2019).
Lawson, D. J. et al. Is population structure in the genetic biobank era irrelevant, a challenge, or an opportunity? Hum. Genet. 139, 23–41 (2020).
Loh, P.-R. et al. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat. Genet. 47, 284–290 (2015).
Yang, J., Zaitlen, N. A., Goddard, M. E., Visscher, P. M. & Price, A. L. Advantages and pitfalls in the application of mixed-model association methods. Nat. Genet. 46, 100–106 (2014).
Davies, N. M. et al. Within family Mendelian randomization studies. Hum. Mol. Genet. 28R2 R170–R179, https://doi.org/10.1093/hmg/ddz204 (2019).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Lee, J. J., McGue, M., Iacono, W. G. & Chow, C. C. The accuracy of LD Score regression as an estimator of confounding and genetic correlations in genome-wide association studies. Genet. Epidemiol. 42, 783–795 (2018).
Zheng, J. et al. LD Hub: a centralized database and web interface to perform LD score regression that maximizes the potential of summary level GWAS data for SNP heritability and genetic correlation analysis. Bioinformatics 33, 272–279 (2017).
Lee, J. J. et al. Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals. Nat. Genet. 50, 1112–1121 (2018).
Gage, S. H., Davey Smith, G., Ware, J. J., Flint, J. & Munafò, M. R. G = E: what GWAS can tell us about the environment. PLoS Genet. 12, e1005765 (2016).
Turley, P. et al. Multi-trait analysis of genome-wide association summary statistics using MTAG. Nat. Genet. 50, 229–237 (2018).
Grotzinger, A. D. et al. Genomic structural equation modelling provides insights into the multivariate genetic architecture of complex traits. Nat. Hum. Behav. 3, 513–525 (2019).
Baselmans, B. M. L. et al. Multivariate genome-wide analyses of the well-being spectrum. Nat. Genet. 51, 445–451 (2019).
Loehlin, J. C. The Cholesky approach: A cautionary note. Behav. Genet. 26, 65–69 (1996).
Torkamani, A., Wineinger, N. E. & Topol, E. J. The personal and clinical utility of polygenic risk scores. Nat. Rev. Genet. 19, 581–590 (2018).
Dudbridge, F. Power and predictive accuracy of polygenic risk scores. PLoS Genet. 9, e1003348 (2013).
Wray, N. R. et al. Research review: Polygenic methods and their application to psychiatric traits. J. Child Psychol. Psychiatry 55, 1068–1087 (2014).
de Vlaming, R. et al. Meta-GWAS accuracy and power (MetaGAP) calculator shows that hiding heritability is partially due to imperfect genetic correlations across studies. PLoS Genet. 13, e1006495 (2017).
Rietveld, C. A. et al. GWAS of 126,559 individuals identifies genetic variants associated with educational attainment. Science 340, 1467–1471 (2013).
Okbay, A. et al. Genome-wide association study identifies 74 loci associated with educational attainment. Nature 533, 539–542 (2016).
Wood, A. R. et al. Defining the role of common variation in the genomic and biological architecture of adult human height. Nat. Genet. 46, 1173–1186 (2014).
Yengo, L. et al. Meta-analysis of genome-wide association studies for height and body mass index in ∼700000 individuals of European ancestry. Hum. Mol. Genet. 27, 3641–3649 (2018).
Belsky, D. W. & Harden, K. P. Phenotypic annotation: Using polygenic scores to translate discoveries from genome-wide association studies from the top down. Curr. Dir. Psychol. Sci. 28, 82–90 (2019).
Barcellos, S. H., Carvalho, L. S. & Turley, P. Education can reduce health differences related to genetic risk of obesity. Proc. Natl Acad. Sci. USA 115, E9765–E9772 (2018).
Bansal, V. et al. Genome-wide association study results for educational attainment aid in identifying genetic heterogeneity of schizophrenia. Nat. Commun. 9, 3078 (2018).
Belsky, D. W. et al. The genetics of success: how single-nucleotide polymorphisms associated with educational attainment relate to life-course development. Psychol. Sci. 27, 957–972 (2016).
Young, A. I., Benonisdottir, S., Przeworski, M. & Kong, A. Deconstructing the sources of genotype-phenotype associations in humans. Science 365, 1396–1400 (2019).
Mostafavi, H., Harpak, A., Conley, D., Pritchard, J. K. & Przeworski, M. Variable prediction accuracy of polygenic scores within an ancestry group. eLife 9, e48376 (2020).
DiPrete, T. A., Burik, C. A. P. & Koellinger, P. D. Genetic instrumental variable regression: Explaining socioeconomic and health outcomes in nonexperimental data. Proc. Natl Acad. Sci. USA 115, E4970–E4979 (2018).
Wertz, J. et al. Genetics of nurture: A test of the hypothesis that parents’ genetics predict their observed caregiving. Dev. Psychol. 55, 1461–1472 (2019).
SSGAC Polygenic Score Data. Health and Retirement Study https://hrs.isr.umich.edu/news/ssgac-polygenic-score-data (Accessed 11 November 2019).
Young, A. I. et al. Relatedness disequilibrium regression estimates heritability without environmental bias. Nat. Genet. 50, 1304–1310 (2018).
Davey Smith, G. & Hemani, G. Mendelian randomization: genetic anchors for causal inference in epidemiological studies. Hum. Mol. Genet. 23R1 R89–R98 (2014).
Brumpton, B. et al. Within-family studies for Mendelian randomization: avoiding dynastic, assortative mating, and population stratification biases. bioRxiv https://doi.org/10.1101/602516 (2019).
O’Connor, L. J. & Price, A. L. Distinguishing genetic correlation from causation across 52 diseases and complex traits. Nat. Genet. 50, 1728–1734 (2018).
Zhu, Z. et al. Causal associations between risk factors and common diseases inferred from GWAS summary data. Nat. Commun. 9, 224 (2018).
Verbanck, M., Chen, C.-Y., Neale, B. & Do, R. Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat. Genet. 50, 693–698 (2018).
Koellinger, P. D. & de Vlaming, R. Mendelian randomization: the challenge of unobserved environmental confounds. Int. J. Epidemiol. 48, 665–671 (2019).
Black, S. E., Devereux, P. J. & Salvanes, K. G. Why the apple doesn’t fall far: Understanding intergenerational transmission of human capital. Am. Econ. Rev. 95, 437–449 (2005).
Bates, T. C. et al. The nature of nurture: Using a virtual-parent design to test parenting effects on children’s educational attainment in genotyped families. Twin Res. Hum. Genet. 21, 73–83 (2018).
Liu, H. Social and genetic pathways in multigenerational transmission of educational attainment. Am. Sociol. Rev. 83, 278–304 (2018).
Belsky, D. W. et al. Genetic analysis of social-class mobility in five longitudinal studies. Proc. Natl Acad. Sci. USA 115, E7275–E7284 (2018).
Barth, D., Papageorge, N. W. & Thom, K. Genetic endowments and wealth inequality. J. Polit. Econ. 128, 1474–1522 (2020).
Holland, P. W. Statistics and causal inference. J. Am. Stat. Assoc. 81, 945–960 (1986).
Rodgers, J. & Kohler, H.-P. The Biodemography of Human Reproduction and Fertility. (Springer, 2002).
Barban, N. et al. Genome-wide analysis identifies 12 loci influencing human reproductive behavior. Nat. Genet. 48, 1462–1472 (2016).
Day, F. R. et al. Genomic analyses identify hundreds of variants associated with age at menarche and support a role for puberty timing in cancer risk. Nat. Genet. 49, 834–841 (2017).
Day, F. R. et al. Physical and neurobehavioral determinants of reproductive onset and success. Nat. Genet. 48, 617–623 (2016).
Mehta, D. et al. Evidence for genetic overlap between schizophrenia and age at first birth in women. JAMA Psychiatry 73, 497–505 (2016).
Ni, G., Gratten, J., Wray, N. R. & Lee, S. H., Schizophrenia Working Group of the Psychiatric Genomics Consortium. Age at first birth in women is genetically associated with increased risk of schizophrenia. Sci. Rep. 8, 10168 (2018).
Harden, K. P. et al. A behavior genetic investigation of adolescent motherhood and offspring mental health problems. J. Abnorm. Psychol. 116, 667–683 (2007).
Beauchamp, J. P. Genetic evidence for natural selection in humans in the contemporary United States. Proc. Natl Acad. Sci. USA 113, 7774–7779 (2016).
Kong, A. et al. Selection against variants in the genome associated with educational attainment. Proc. Natl Acad. Sci. USA 114, E727–E732 (2017).
Rodgers, J. L. et al. Education and cognitive ability as direct, mediating, or spurious influences on female age at first birth: behavior genetic models fit to Danish twin data. Am. J.Sociol. 114, S202–S232 (2008). Suppl.
Tropf, F. C. & Mandemakers, J. J. Is the association between education and fertility postponement causal? The role of family background factors. Demography 54, 71–91 (2017).
Jocklin, V., McGue, M. & Lykken, D. T. Personality and divorce: a genetic analysis. J. Pers. Soc. Psychol. 71, 288–299 (1996).
D’Onofrio, B. M., Eaves, L. J., Murrelle, L., Maes, H. H. & Spilka, B. Understanding biological and social influences on religious affiliation, attitudes, and behaviors: a behavior genetic perspective. J. Pers. 67, 953–984 (1999).
Pilling, L. C. et al. Human longevity: 25 genetic loci associated in 389,166 UK biobank participants. Aging (Albany NY) 9, 2504–2520 (2017).
Abdellaoui, A. et al. Genetic correlates of social stratification in Great Britain. Nat. Hum. Behav. 3, 1332–1342 (2019).
Hill, W. D. et al. Molecular genetic contributions to social deprivation and household income in UK Biobank. Curr. Biol. 26, 3083–3089 (2016).
Belsky, D. W. et al. Genetics and the geography of health, behaviour and attainment. Nat. Hum. Behav. 3, 576–586 (2019).
van der Sluis, S., Posthuma, D. & Dolan, C. V. A note on false positives and power in G × E modelling of twin data. Behav. Genet. 42, 170–186 (2012).
Duncan, L. E. & Keller, M. C. A critical review of the first 10 years of candidate gene-by-environment interaction research in psychiatry. Am. J. Psychiatry 168, 1041–1049 (2011).
Ceci, S. J. & Papierno, P. B. The rhetoric and reality of gap closing: when the “have-nots” gain but the “haves” gain even more. Am. Psychol. 60, 149–160 (2005).
Fletcher, J. M. Why have tobacco control policies stalled? Using genetic moderation to examine policy impacts. PLoS One 7, e50576 (2012).
Goldberger, A. S. Heritability. Economica 46, 327–347 (1979).
Jencks, C. Heredity, environment, and public policy reconsidered. Am. Sociol. Rev. 45, 723–736 (1980).
Bronfenbrenner, U. & Ceci, S. J. Nature-nurture reconceptualized in developmental perspective: a bioecological model. Psychol. Rev. 101, 568–586 (1994).
Dickens, W. T. & Flynn, J. R. Heritability estimates versus large environmental effects: the IQ paradox resolved. Psychol. Rev. 108, 346–369 (2001).
Lykken, D. T., Bouchard, T. J. Jr., McGue, M. & Tellegen, A. Heritability of interests: a twin study. J. Appl. Psychol. 78, 649–661 (1993).
Tucker-Drob, E. M. & Harden, K. P. Early childhood cognitive development and parental cognitive stimulation: evidence for reciprocal gene-environment transactions. Dev. Sci. 15, 250–259 (2012).
Klahr, A. M., Thomas, K. M., Hopwood, C. J., Klump, K. L. & Burt, S. A. Evocative gene-environment correlation in the mother-child relationship: a twin study of interpersonal processes. Dev. Psychopathol. 25, 105–118 (2013).
Scarr, S. & McCartney, K. How people make their own environments: a theory of genotype greater than environment effects. Child Dev. 54, 424–435 (1983).
Wray, N. R. et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat. Genet. 50, 668–681 (2018).
Wehby, G. L., Domingue, B. W. & Wolinsky, F. D. Genetic risks for chronic conditions: Implications for long-term wellbeing. J. Gerontol. A Biol. Sci. Med. Sci. 73, 477–483 (2018).
Mallard, T. T., Harden, K. P. & Fromme, K. Genetic risk for schizophrenia is associated with substance use in emerging adulthood: an event-level polygenic prediction model. Psychol. Med. 49, 2027–2035 (2019).
Okbay, A. et al. Genetic variants associated with subjective well-being, depressive symptoms, and neuroticism identified through genome-wide analyses. Nat. Genet. 48, 624–633 (2016).
Wootton, R. E. et al. Evaluation of the causal effects between subjective wellbeing and cardiometabolic health: mendelian randomisation study. Br. Med. J. 362, k3788 (2018).
Kevles, D.J. In the Name of Eugenics: Genetics and the Uses of Human Heredity. (Harvard University Press, 1995).
Rawls, J. A Theory of Justice. (Belknap Press, 1999).
Herrnstein, R.J. & Murray, C. The Bell Curve: Intelligence and Class Structure in American Life (Free Press, 1996).
Jensen, A. How much can we boost IQ and scholastic achievement. Harv. Educ. Rev. 39, 1–123 (1969).
Yudell, M., Roberts, D., DeSalle, R. & Tishkoff, S. Taking race out of human genetics. Science 351, 564–565 (2016).
Smedley, A. & Smedley, B. D. Race as biology is fiction, racism as a social problem is real: Anthropological and historical perspectives on the social construction of race. Am. Psychol. 60, 16–26 (2005).
Henrich, J., Heine, S. J. & Norenzayan, A. The weirdest people in the world? Behav. Brain Sci. 33, 61–83 (2010). discussion 83–135.
Martin, A. R. et al. Clinical use of current polygenic risk scores may exacerbate health disparities. Nat. Genet. 51, 584–591 (2019).
Martin, A. R. et al. Human demographic history impacts genetic risk prediction across diverse populations. Am. J. Hum. Genet. 100, 635–649 (2017).
Panofsky, A. & Donovan, J. Genetic ancestry testing among white nationalists: From identity repair to citizen science. Soc. Stud. Sci. https://doi.org/10.1177/0306312719861434 (2019).
Novembre, J. & Barton, N. H. Tread lightly interpreting polygenic tests of selection. Genetics 208, 1351–1355 (2018).
Acknowledgements
We thank C. Burik for preparing Fig. 1 and the Social Science Genetic Association Consortium (https://www.thessgac.org/) for Fig. 3. P.D.K. was financially supported by an ERC consolidator grant (647648 EdGe). K.P.H. was supported by the Jacobs Foundation, the Templeton Foundation and Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD) grants R01-HD083613 and 5-R24-HD042849 (to the Population Research Center at the University of Texas at Austin). The funders had no role in the conceptualization, preparation or decision to publish this work.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Editor recognition statement: Primary handling editor: Stavroula Kousta.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Harden, K.P., Koellinger, P.D. Using genetics for social science. Nat Hum Behav 4, 567–576 (2020). https://doi.org/10.1038/s41562-020-0862-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41562-020-0862-5
This article is cited by
-
Infrastructuring Educational Genomics: Associations, Architectures, and Apparatuses
Postdigital Science and Education (2024)
-
Could interventions on physical activity mitigate genomic liability for obesity? Applying the health disparity framework in genetically informed studies
European Journal of Epidemiology (2023)
-
The impact of digital media on children’s intelligence while controlling for genetic differences in cognition and socioeconomic background
Scientific Reports (2022)
-
Delayed tracking and inequality of opportunity: Gene-environment interactions in educational attainment
npj Science of Learning (2022)
-
Genetics of cognitive performance, education and learning: from research to policy?
npj Science of Learning (2022)