Abstract
Climate change is expected to impact the functioning of the entire Earth system. However, detecting changes in ecosystem dynamics and attributing such change to anthropogenic climate change has proved difficult. Here we analyse the vegetation dynamics of 100 sites representative of the diversity of terrestrial ecosystem types using remote-sensing data spanning the past 40 years and a dynamic model of plant growth, forced by climate reanalysis data. We detect a change in vegetation activity for all ecosystem types and find these changes can be attributed to trends in climate-system parameters. Ecosystems in dry and warm locations responded primarily to changes in soil moisture, whereas ecosystems in cooler locations responded primarily to changes in temperature. We find that the effects of CO2 fertilization on vegetation are limited, potentially due to masking by other environmental drivers. Observed trend switching is widespread and dominated by shifts from greening to browning, suggesting many of the ecosystems studied are accumulating less carbon. Our study reveals a clear fingerprint of climate change in the change exhibited by terrestrial ecosystems over recent decades.
Similar content being viewed by others
Main
Climate change is impacting the structure and functioning of terrestrial ecosystems1,2,3,4,5,6. Detecting this change and attributing it to climate change is a major research challenge in Earth systems science7. Detection is difficult because signals in the data can be diluted by the stochasticity inherent in the system, the short length of available time series, errors in the observation process and the often overriding effect of land-use change. Attribution is difficult because ecosystem dynamics can be nonlinear and asynchronous and the dynamics are co-limited by environmental drivers that are often highly correlated with one another. For example, biomass increases that are expected as a consequence of enhanced carbon assimilation by elevated atmospheric CO2 concentrations may be constrained by water and nutrient availability8,9,10,11. These difficulties mean that convincing detection and attribution of climate change impacts on ecosystems is limited to a small number of well-studied systems7.
The most convincing evidence to date for attributing changes in ecosystems to climate change comes from the high latitudes where temperatures are increasing from the lower thermal limits of multiple ecological processes7,12. In such examples, the case for attribution has been constructed from experimental, modelling and observational lines of evidence derived from multiple studies. Yet it has emerged that even in these high-latitude ecosystems, where there is convincing evidence for change, not all instances of these ecosystems have changed in the same way13,14,15. Some cases exhibited greening (an increase in vegetation activity), others browning (a decrease in vegetation activity), and for yet others, the initially reported trends had to be revised as the observation window has expanded in time and space10,14,15,16,17. This suggests that robust detection and attribution require longer time series and assessments at multiple instances of each of the world’s major ecosystem types.
Earth observation satellite programmes provide multi-decade time series of vegetation activity and allow the monitoring of many sites. Previous studies have used Earth observation data to detect change in vegetation activity across the entire land surface and identified that vegetation change is to some extent sensitive to climate-system parameters18,19,20,21,22,23. However, such complete analyses of the land surface include pervasive land-use impacts in the data24, which may confound climate change and land use as drivers of vegetation change. Interpretation is further complicated by the fact that previous studies used model-based interpretations of the radiometric data collected by satellites such as net primary productivity (NPP18,21,25), gross primary productivity (GPP23,25) and leaf area index (LAI20).
A state-space method for detection and attribution
Deepening our understanding of how ecosystems respond to climate change requires a robust detection and attribution methodology that addresses the problems highlighted in the previous paragraphs. In this study, we develop and apply a new method for detecting and attributing climate change impacts on terrestrial ecosystems that addresses confounding effects of land-use change, biases associated with short time series, limitations of correlative methods and ambiguities with interpreting NPP, GPP and LAI modelled from satellite-derived data. We detect change using high-quality, long-term remotely sensed normalized difference vegetation index (NDVI) and enhanced vegetation index (EVI) time series. NDVI and EVI data products are sensitive indicators of leaf area and chlorophyll content, and unlike NPP, GPP and LAI products, they are based purely on the radiometrically and geometrically corrected measurements made by satellites. The NDVI time series we use spans 1981–2015 and was derived from the Advanced Very High Resolution Radiometer (AVHRR) satellite programme as part of the Global Inventory Modeling and Mapping Studies (GIMMS) project26,27. The EVI dataset we use spans 2000–2019 and is a product of the Moderate Resolution Imaging Spectroradiometer (MODIS) programme28. GIMMS NDVI provides a longer record, while MODIS EVI offers the opportunity to use a vegetation index that does not saturate at high chlorophyll situations and is less sensitive to atmospheric and soil contamination28.
To avoid the confounding effects of direct human impacts on vegetation, we carefully selected 100 sites, distributed across the major ecosystems of the world, where direct land-use impacts are absent (Methods). We considered only grid cells (the GIMMS NDVI grid cells are 1/12° in size, ~9 km) that show no signs of anthropogenic land transformation in the observation period. We further included only sites where the vegetation is relatively homogeneous within the grid cell. The MODIS EVI data were resampled to the GIMMS NDVI grid (Methods). Our final sample is stratified to include at least five examples of each of the major ecosystems of the world. These criteria yielded a total of 100 sites (Extended Data Figs. 2 and 3) covering tropical evergreen forest (RF, n = 16), boreal forest (BF, n = 10), temperate forest (TF, n = 12), savannah (SA, n = 18), shrubland (SH, n = 16), grassland (GR, n = 14), tundra (TU, n = 9) and Mediterranean-type ecosystems (MT, n = 5).
At each site, we attributed the detected vegetation index change to climate-system parameters (soil temperature, air temperature, soil moisture, solar radiation, atmospheric CO2 concentration) by using a process-based model of plant growth29,30 forced by weekly climate-system data. The plant growth model allows us to interpret how temporal variation in satellite observations of vegetation indices relate to variation in climate-system parameters31,32. A state-space modelling framework33 is used to estimate the parameters of the growth model from the vegetation index time series. The state-space analysis assumes that the remotely sensed NDVI or EVI observations arise from an unobserved underlying dynamic process that is represented by the plant growth model (Methods). The analysis furthermore treats process and observation uncertainty separately, allowing us to account for how observation uncertainty related to cloud and snow cover15,18,19 structures uncertainty in the analysis.
The process model we use29 simulates a single plant that represents a virtual phytometer exposed to different environmental conditions; it therefore differs from the Dynamic Global Vegetation Models34 used in Earth system model attribution studies20,21,23. The simulations focus on modelling the plant’s carbon and nitrogen assimilation and, in a separate process, how these assimilates influence growth29. Assimilation and growth processes in the model are co-limited by combinations of air and soil temperature, soil moisture, solar radiation and atmospheric CO230,35 (Methods). The model’s structure thereby represents a hypothesis of how environmental factors co-limit the growth of a phytometer.
The first step in our analysis is to use the state-space approach to estimate the model parameters for each vegetation index time series (Fig. 1a). We then use time-series decomposition methods to identify anomalies in the vegetation indices (Fig. 1b) and to generate detrended time series of the climate-system forcing data (air and soil temperature, soil moisture, solar radiation, atmospheric CO2; Methods). The fitted model is then used with the full and detrended climate data to predict the observed anomalies in the vegetation indices. To attribute vegetation trends to trends in the climate-system data, we use the slope of the regression (with intercept 0) between observed and modelled anomalies (Fig. 1c). Attribution is diagnosed if the slope of this regression from model runs forced by the full climate data (red line in Fig. 1c) is positive and clearly higher (no overlap in the credible intervals of the slopes; Extended Data Figs. 9a and 10a) than the regression slope from model runs forced by climate data with trends removed (blue line in Fig. 1c). That is, attribution is diagnosed if trends in climate-system variables statistically enhance the ability to predict the observed anomalies in the vegetation indices. For cases where attribution is supported, we estimate the relative importance of trends in each climate-system variable in explaining the anomalies. This was achieved by assessing the change in regression slope induced by separately removing trends in each climate-system variable (Fig. 1d). The same analyses were repeated using the EVI time series (Extended Data Figs. 4, 9b and 10b).
a, GIMMS NDVI time series of vegetation activity (Data) and the process model’s (Model) fit to these data for a savannah site in the Burkina Faso National Park. The blue shaded envelope shows the 95% credible intervals around the mean model predictions, which includes parameter, process and observation uncertainty. b, Anomalies in the NDVI data (blue bars) and the fit of a bent-cable regression to these anomalies. The shaded envelope shows the 95% credible intervals of the bent-cable regression predictions. c, Zero-intercept regression showing the model’s ability to predict observed anomalies, with full and detrended climate-forcing data. Shaded envelopes show the 95% credible intervals of the mean regression line. d, Posterior density of the change in the full model’s regression slope (as shown in c) caused by removing trends in a climate-forcing variable from the full model; in this example, the ability to predict anomalies was sensitive to soil-moisture (Msoil) and soil-temperature (Tsoil) anomalies. Extended Data Fig. 4 shows an analogous plot using MODIS EVI data. Tair, air temperature; Srad, solar radiation; CO2, atmospheric CO2 concentration.
Widespread shifts from greening to browning
The anomaly trends observed in the satellite vegetation indices revealed qualitatively different patterns when interpreted using trend analyses. To quantify these different patterns, we used both quadratic polynomials and a bent-cable piecewise linear regression. Both approaches allow one to test whether there is a long-term trend and whether there has been a shift in this long-term trend. We classified the response trends as being (1) hat shaped where trends in the vegetation indices increased at first (‘greening’) but later declined (‘browning’), (2) cup shaped where trends decreased initially and later increased or (3) linear where no change in the sign of the trend was detected. The majority of sites illustrated hat-shaped trends, fewer sites illustrated cup-shaped trends and only a minority of sites illustrated linear trends (Fig. 2 and Extended Data Fig. 5). In the analyses that used the shorter EVI time series, the cup-shaped trends were more common than in the analyses that used NDVI data (Extended Data Fig. 5). In the NDVI data, overall increasing and decreasing vegetation greenness trends were almost equally distributed, while in the EVI data, more sites revealed increasing trends. Analysing these trends for the common (15 yr) time window for which both data products are available revealed similar distributions although the EVI data revealed more increasing trends than the NDVI data (Extended Data Fig. 6). Despite uncertainty associated with vegetation index datasets and time-series window (1982–2015 or 2000–2019), both analyses agreed that nonlinearity dominated and that greening trends switching to browning trends was the most common pattern in the data.
The frequency of cup-shaped (initial decrease, subsequent increase), hat-shaped (initial increase, subsequent decrease) and linear time trends in the anomalies in NDVI vegetation activity. The number of anomaly trends that showed an overall increase or decrease over the time series is indicated by different colours. Overall increases or decreases less than 2% in NDVI were classified as small. The response types were classified from bent-cable regression models fitted to the anomaly data from each site (for example, Fig. 1b). Similar results were found when using a polynomial regression model and EVI data (Extended Data Fig. 5).
The dominance of shifts from greening to browning trends in vegetation activity detected here contradicts the predominance of positive trends previously reported20 but is supported by studies that have shown negative NDVI and EVI trends36. Hat-shaped trend switching has been reported in high-latitude ecosystems where initial greening trends attributed to warming have now been replaced by browning trends attributed to seasonal water deficits10,37. Positive to negative trend switching has also been reported in Amazonian forests, and predictions are that African tropical forests will follow suit38. Switching of the kind observed here is in all likelihood driven by interactions between climate-system drivers and internal ecosystem dynamics39, which makes the prediction of future trends highly context dependent.
Attribution of vegetation activity change
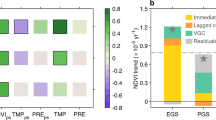
In 80 (NDVI data) or 75 (EVI data) of the 100 study sites, the model was able to explain variation in the observed trends in the vegetation indices and attribute this to trends in the climate-forcing data (Extended Data Fig. 9). This demonstrates a clear fingerprint of climate change on the activity of the world’s ecosystems. A classification of sites according to the relative importance of climate-system variables driving vegetation anomalies yielded two groups that were qualitatively consistent between the NDVI and EVI analyses (Fig. 3 and Extended Data Fig. 7). In the first group, moisture was the dominant factor explaining anomalies in the vegetation data; this group consisted primarily of grassland, savannah and shrubland sites. In the second group, vegetation index anomalies were explained by a combination of factors, with temperature often playing a central role; this group consisted primarily of boreal forest, temperate forest and tundra sites. It is notable that trends in CO2 and solar radiation seldom had a dominant influence on the model’s ability to predict the observed anomalies in the vegetation indices (Fig. 3 and Extended Data Fig. 7). The lack of evidence for dominating CO2 effects contradicts a previous study of LAI20 but is consistent with analyses that conclude that CO2 enrichment effects on vegetation biomass are conditional on nutrient and water supply8,11, mycorrhiza9 and successional stage25 and with studies that suggest that CO2 enrichment effects on GPP and biomass accumulation may be weakening as global warming progresses10,18,23,38.
The sensitivity is quantified as the effect of each of five forcing factors (Tair, Tsoil, Msoil, Srad and CO2) on the slope, describing the ability of the model to predict anomalies in the NDVI vegetation activity time series (compare Fig. 1). The slope of the full model is represented by the red colour ramp. Shown in the matrix are the 80 of 100 sites where anomalies in vegetation activity could be attributed to the environmental-forcing factors. The coloured circles indicate the response groups the sites are assigned to by an unsupervised classification of the sites by the effects of the five forcing factors on the slope. The site codes indicate ecosystem type (BF = boreal forest, GR = grassland, MT = Mediterranean-type ecosystems, RF = tropical evergreen forest, SA = savanna, SH = shrubland, TF = temperate forest, TU = tundra.), ISO 3166-1 alpha-3 country codes and a unique site identifier code. See Extended Data Fig. 7 for the same analysis using EVI data.
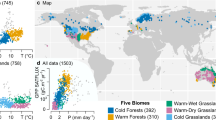
Which environmental factors dominated the anomaly response was clearly structured in climate space. A discriminant analysis revealed that the major groups in Fig. 3 and Extended Data Fig. 7 can be separated using mean annual temperature and mean annual soil moisture (Fig. 4a and Extended Data Fig. 8a). Sites where anomalies in vegetation indices were attributed to anomalies in soil moisture were located in warm and dry locations. Sites where a mixture of factors, often dominated by temperature, explained anomalies in vegetation indices were centred in cooler and moister locations. The 20 (NDVI data) and 25 (EVI data) sites where anomalies in the forcing data could not explain vegetation index anomalies were centred in warmer and moister locations (sites labelled 0 in Fig. 4 and Extended Data Fig. 8); indeed, the majority of tropical rainforest sites were in this group.
a, The distribution of attribution classes (Fig. 3) in bivariate temperature and moisture climate space. Points represent the locations of the 100 study sites in climate space. Attribution classes are the two major groups classified in Fig. 3. Sites labelled 0 are sites excluded from Fig. 3 because the model could not explain observed anomalies. The colours of the sites labelled 1 and 2 correspond to the groups classified in Fig. 3. The ellipses represent the fitted covariance estimates of the classes as estimated by discriminant analysis based on Gaussian finite mixture modelling. b,c, The shape (b) and direction (c) of the anomaly response (as defined in Fig. 2) plotted in bivariate climate space. The mean annual temperature and mean annual soil moisture were calculated over the study period using the ERA5-Land reanalysis data (European Centre for Medium-Range Weather Forecasts Reanalysis v. 5). d–f, The points in panels a (d), b (e) and c (f) plotted in geographic space. See Extended Data Fig. 8 for the same analysis using EVI data.
In general, these findings are consistent with fundamental ecological premises that warm and dry ecosystems are water limited and that cooler ecosystems are more likely to be temperature limited. What is perhaps unexpected was that changes in vegetation indices in ecosystems in warm and moist locations could not be attributed to changes in a third factor, such as atmospheric CO2. However, we would also expect our method to have lower statistical power in environments with low seasonal and interannual variation in vegetation activity simply because these situations yield vegetation index time series with lower information content. This is particularly true for the NDVI analysis since NDVI is known to saturate at the LAIs typical of tropical forests. Furthermore, moist tropical regions are often phosphorus limited40, which may constrain the ability of such ecosystems to respond to elevated CO2 (ref. 9).
The shape of the vegetation response and whether the response is increasing or decreasing over time (as summarized in Fig. 2) were not clearly structured in environmental space (Fig. 4 and Extended Data Fig. 8). The complexity in these patterns is consistent with modelling studies that predicted that transitions in ecosystem behaviour will occur at different points in time for different geographic locations41 and with remote-sensing and field studies that have reported both increasing and decreasing vegetation responses within regions and observation time windows14,16,17,38.
Co-limiting drivers of change
Overall, we have detected change in indices of vegetation activity in all terrestrial ecosystem types, and in the majority of cases studied (80% for the NDVI analysis and 75% for the EVI analysis) we were able to attribute observed changes in vegetation indices to trends in climate-system variables. Temperature and moisture trends explained most of the variation, whereas increasing CO2 and changes in surface solar radiation were seldom important. The apparent insensitivity of vegetation to CO2 is due to co-limitation processes in the growth model that allow the model to accommodate no increases in plant biomass despite increasing potential carbon assimilation caused by increasing atmospheric CO2. For example, in the model, elevated CO2 allows for higher potential rates of carbon assimilation, but decreases in soil moisture can prevent this potential carbon assimilation from being realized. In addition, the model can represent that higher carbon assimilation will not lead to enhanced growth as long as one of nitrogen, temperature and soil moisture is limiting. Analogous co-limitation pathways allow the model to simulate situations in high latitudes where increasing temperatures promote growth, yet increasing water deficits retard growth12,42.
The dominance of hat-shaped responses in the vegetation indices analysed here suggests that terrestrial ecosystems, which have been a valuable carbon sink in recent decades6, may sequester less carbon in coming decades10,18,23,38. However, NDVI and EVI provide an incomplete picture of ecosystem carbon sequestration43. This is because NDVI and EVI, although sensitive to changes in the fraction of photosynthetically active radiation that is absorbed by leaves, are less sensitive to how efficiently this fraction is translated into assimilated carbon (light use efficiency (LUE))43. It follows that both the NDVI and EVI data may poorly represent changes in LUE, which is known to increase with elevated CO244. Plants that invest the spoils of increased LUE in root growth, defence, mutualisms or reproduction may therefore go undetected in our analysis. This incomplete picture of vegetation carbon dynamics could be improved by considering additional data sources. For example, satellite-based observations of chlorophyll fluorescence provide new opportunities for estimating LUE43,45, allowing the separation of carbon assimilation from leaf area dynamics46. The state-space model used here could be expanded to simultaneously model a fraction of absorbed photosynthetically active radiation index (EVI or NDVI) from the process model’s biomass and a LUE index (for example, solar-induced fluorescence) from the model’s carbon assimilation. This would further constrain the model’s parameter estimates and allow attribution of both biomass and GPP trends to climate-system variables.
Dealing with climate change is at minimum a three-step process: detection, attribution and adaptation/mitigation. We have illustrated a generally applicable method for detection and attribution of climate change impacts on the world’s major terrestrial ecosystems. In this study, we detected that many ecosystems are on hat-shaped trajectories, where initial greening trends have switched to browning trends, suggesting that the majority of the ecosystems studied here may be accumulating less leaf biomass and potentially less carbon than they did previously. The detection and attribution methodology may help adaptation and mitigation strategists better understand the trajectories that ecosystems are on. In particular, our analyses revealed geographic coherence as to which environmental drivers are driving change, which may help the design of ecosystem restoration programmes.
Methods
Plant growth model without environmental forcing
The model without environmental forcing closely follows the original description of the Thornley transport resistance (TTR) model29. A summary of the model parameters is provided in Supplementary Table 2. The shoot and root mass pools (MS and MR, in kg structural dry matter) change as a function of growth and loss (equations (1) and (2)). The litter (kL) and maintenance respiration (r) loss rates (in kg kg−1 d−1) are treated as constants. In the original model description29 r = 0. The parameter KM (units kg) describes how loss varies with mass (MS or MR). Growth (Gs and Gr, in kg d−1) varies as a function of the carbon and nitrogen concentrations (equations (3) and (4)). CS, CR, NS and NR are the amounts (kg) of carbon and nitrogen in the roots and shoots. These assumptions yield the following equations for shoot and root dry matter,
where GS and GR are
and g is the growth coefficient (in kg kg−1 d−1).
Carbon uptake UC is determined by the net photosynthetic rate (a, in kg kg−1 d−1) and the shoot mass (equation (5)). Similarly, nitrogen uptake (UN) is determined by the nitrogen uptake rate (b, in kg kg−1 d−1) and the root mass. The parameter KA (units kg) forces both photosynthesis and nitrogen uptake to be asymptotic with mass. The second terms in the denominators of equations (5) and (6) model product inhibitions of carbon and nitrogen uptake, respectively; that is, the parameters JC and JN (in kg kg−1) mimic the inhibition of source activity when substrate concentrations are high,
The substrate transport fluxes of C and N (τC and τN, in kg d−1) between roots and shoots are determined by the concentration gradients between root and shoot and by the resistances. In the original model description29, these resistances are defined flexibly, but we simplify and assume that they scale linearly with plant mass,
The changes in mass of carbon and nitrogen in the roots and shoots are then
where fC and fN (in kg kg−1) are the fractions of structural carbon and nitrogen in dry matter.
Adding environmental forcing to the plant growth model
In this section, we describe how the net photosynthetic rate (a), the nitrogen uptake rate (b), the growth rate (g) and the respiration rate (r) are influenced by environmental-forcing factors. These environmental-forcing effects are described in equations (13)–(17) and summarized graphically in Extended Data Fig. 1. All other model parameters are treated as constants. Previous work that implemented the TTR model as a species distribution model30 is used as a starting point for adding environmental forcing. As in this previous work30, we assume that parameters a, b and g are co-limited by environmental factors in a manner analogous to Liebig’s law of the minimum, which is a crude but pragmatic abstraction. The implementation here differs in some details.
Unlike previous work30, we use the Farquhar model of photosynthesis47,48 to represent how solar radiation, atmospheric CO2 concentration and air temperature co-limit photosynthesis35. We assume that the Farquhar model parameters are universal and that all vegetation in our study uses the C3 photosynthetic pathway. The Farquhar model photosynthetic rates are rescaled to [0,amax] to yield afqr. The effects of soil moisture (Msoil) on photosynthesis are represented as an increasing step function \({{{{S}}}}(M_{\mathrm{soil}},{\beta }_{1},{\beta }_{2})=\max \left\{\min \left(\frac{M_{\mathrm{soil}}-{\beta }_{1}}{{\beta }_{2}-{\beta }_{1}},1\right),0\right\}\). This allows us to redefine a as,
The processes influencing nitrogen availability are complex, and global data products on plant available nitrogen are uncertain. We therefore assume that nitrogen uptake will vary with soil temperature and soil moisture. That is, the nitrogen uptake rate b is assumed to have a maximum rate (bmax) that is co-limited by soil temperature Tsoil and soil moisture Msoil,
In equation (14), we have assumed that the nitrogen uptake rate is a simple increasing and saturating function of temperature. We have also assumed that the nitrogen uptake rate is a trapezoidal function of soil moisture with low uptake rates in dry soils, higher uptake rates at intermediate moisture levels and lower rates once soils are so moist as to be waterlogged. The trapezoidal function is \({{{{Z}}}}(M_{\mathrm{soil}},{\beta }_{5},{\beta }_{6},{\beta }_{7},{\beta }_{8})=\max \left\{\min \left(\frac{M_{\mathrm{soil}}-{{{{{\beta }}}}}_{5}}{{{{{{\beta }}}}}_{6}-{{{{{\beta }}}}}_{5}},1,\frac{{{{{{\beta }}}}}_{8}-M_{\mathrm{soil}}}{{\beta }_{8}-{\beta }_{7}}\right),0\right\}\).
The previous sections describe how the assimilation of carbon and nitrogen by a plant are influenced by environmental factors. The TTR model describes how these assimilate concentrations influence growth (equations (3) and (4)). In our implementation, we additionally allow the growth rate to be co-limited by temperature (soil temperature, Tsoil) and soil moisture (Msoil),
We use Tsoil since we assume that growth is more closely linked to soil temperature, which varies slower than air temperature. The respiration rate (r, equations (1) and (2)) increases as a function of air temperature (Tair) to a maximum rmax,
The parameter r is best interpreted as a maintenance respiration. Growth respiration is not explicitly considered; it is implicitly incorporated in the growth rate parameter (g, equation (15)), and any temperature dependence in growth respiration is therefore assumed to be accommodated by equation (15).
Fire can reduce the structural shoot mass MS as follows,
where F is an indicator of fire severity at a point in time (for example, burnt area) and the function S(F, β17, β18) allows MS to decrease when the fire severity indicator F is high. If F = 0, this process plays no role. This fire impact equation was used in preliminary analyses, but the data on fire activity did not provide sufficient information to estimate β17 and β18; we therefore excluded this process from the final analyses.
We further estimate two additional β parameters (βa and βb) so that each site can have unique maximum carbon and nitrogen uptake rates. Specifically, we redefine a as \({a}^{{\prime} }={\beta }_{a}\ a\) and b as \({b}^{{\prime} }={\beta }_{b}\ b\).
Data sources and preparation
To describe vegetation activity, we use the GIMMS 3g v.1 NDVI data26,27 and the MODIS EVI28 data. The GIMMS data product has been derived from the AVHRR satellite programme and controls for atmospheric contamination, calibration loss, orbital drift and volcanic eruptions26,27. The data provide 24 NDVI raster grids for each year, starting in July 1981 and ending in December 2015. The spatial resolution is 1/12° (~9 × 9 km). The EVI data used are from the MODIS programme’s Terra satellite; it is a 1 km data product provided at a 16-day interval. We use data from the start of the record (February 2000) to December 2019. The MODIS data product (MOD13A2) uses a temporal compositing algorithm to produce an estimate every 16 days that filters out atmospheric contamination. The EVI is designed to reduce the effects of atmospheric, bare-ground and surface water on the vegetation index28.
For environmental forcing, we use the ERA5-Land data31,32 (European Centre for Medium-Range Weather Forecasts Reanalysis v. 5; hereafter, ERA5). The ERA5 products are global reanalysis products based on hourly estimates of atmospheric variables and extend from present back to 1979. The data products are supplied at a variety of spatial and temporal resolutions. We used the monthly averages from 1981 to 2019 at a 0.1° spatial resolution (~11 km). The ERA5 data provide air temperature (2 m surface air temperature), soil temperature (0–7 cm soil depth), surface solar radiation and volumetric soil water (0–7 cm soil depth). Fire data were taken from the European Space Agency Fire Disturbance Climate Change Initiative’s AVHRR Long-Term Data Record Grid v.1.0 product49. This product provides gridded (0.25° resolution) data of monthly global (from 1982 to 2017) burned area derived from the AVHRR satellite programme. As mentioned, the fire data did not enrich our analysis, and the analyses we present here therefore exclude further consideration of the fire data.
All data were resampled to the GIMMS grid. The mean pixel EVI was then calculated for each GIMMS cell for each time point in the MODIS EVI data. We used linear interpolation on the NDVI, EVI and ERA5 environmental-forcing data to estimate each variable on a weekly time step. This served to set the time step of the TTR difference equations to one week and to synchronize the different time series.
Site selection
The GIMMS grid cells define the spatial resolution of our sample points. GIMMS grid cells are large (1/12°, ~9 km), meaning that most grid cells contain multiple land-cover types. We focused on wilderness landscapes, and our aim was to find multiple grid cells for the major ecosystems of the world. We used the following classification of ecosystem types to guide the stratification of our grid-cell selection: tropical evergreen forest (RF), boreal forest (BF), temperate evergreen and temperate deciduous forest (TF), savannah (SA), shrubland (SH), grassland (GR), tundra (TU) and Mediterranean-type ecosystems (MT).
We used the following criteria to select grid cells. (1) Selected grid cells should contain relatively homogeneous vegetation. Small-scale heterogeneity was allowed (for example, catenas, drainage lines, peatlands) as long as many of these elements are repeated in the scene (for example, rolling hills were accepted, but elevation gradients were rejected). (2) Sites should have no signs of transformative human activity (for example, tree harvesting, crop cultivation, paved surfaces). We used the Time Tool in Google Earth Pro, which provides annual satellite imagery of the Earth from 1984 onwards, to ensure that no such activity occurred during the observation period (note that the GIMMS record starts in July 1981; however, it is likely that evidence of transformative activity between July 1981 and 1984 would be visible in 1984). Grid cells with extensive livestock holding on non-improved pasture were included. In some cases, a small agricultural field or pasture was present, and such grid cells were used as long as the field or pasture was small and remained constant in size. (3) Grid cells should not include large water bodies, but small drainage lines or ponds were accepted as long as they did not violate the first criterion. (4) Grid cells should be independent (neighbouring grid cells were not selected) and cover the major ecosystems of the world. Using these criteria, we were able to include 100 sites in the study (Extended Data Figs. 2 and 3 and Supplementary Table 4).
State-space model
We used a Bayesian state-space approach. Conceptually, the analysis takes the form,
Here M[t] is the plant growth model’s prediction of biomass (M = MS + MR) at time t, and ϵt−1 is the process error associated with the state variables. In the model, each underlying state variable (MS, MR, CS, CR, NS and NR) has an associated process error term. The function f(M[t − 1], β, θt−1, ϵt−1) summarizes that the development of M is influenced by the state variables MS, MR, CS, CR, NS and NR, the environmental-forcing data θt−1 and the β parameters. The observation equation (equation (19)) uses the parameter m to link the VI (vegetation index, either NDVI or EVI) observations to the modelled state M. The parameter η is the observation error. Equation (19) assumes that there is a linear relationship between modelled biomass (M) and VI, which is a simplification of reality50,51,52. The observation error η is structured by our simplification of the data products quality scores (coded Q = 0, 1, 2, with 0 being good and 2 being poor; Supplementary Table 3) to allow the error to increase with each level of the quality score. Specifically, we define η = e0 + e1 × Q.
The model was formulated using the R package LaplacesDemon53. All β parameters are given vague uniform priors. The parameter m is given a vague normal prior (truncated to be >0). The process error terms are modelled using normal distributions, and the variances of the error terms are given vague half-Cauchy priors. The ex parameters are given vague normal priors. The model also requires the parameterization of M[0], the initial vegetation biomass; M[0] is given a vague uniform prior. We used the twalk Markov chain Monte Carlo (MCMC) algorithm as implemented in LaplacesDemon53 and its default control parameters to estimate the posterior distributions of the model parameters. We further fitted the model using DEoptim54,55, which is a robust genetic algorithm that is known to perform stably on high-dimensional and multi-modal problems56, to verify that the MCMC algorithm had not missed important regions of the parameter space. The models estimated with MCMC had significantly lower log root-mean-square error than models estimated with DEoptim (paired t-test NDVI analysis: t = –2.9806, degrees of freedom (d.f.) = 99, P = 0.00362; EVI analysis: t = –4.6229, d.f. = 99, P = 1.144 × 10–5), suggesting that the MCMC algorithm performed well compared with the genetic algorithm.
Anomaly extraction and trend estimation
We use the ‘seasonal and trend decomposition using Loess’ (STL57) as implemented in the R58 base function stl. STL extracts the seasonal component s of a time series using Loess smoothing. What remains after seasonal extraction (the anomaly or remainder, r) is the sum of any long-term trend and stochastic variation. We estimate the trend in two ways. First, we estimate the trend by fitting a quadratic polynomial (r = a + bx + cx2) to the remainder (r is the remainder, x is time and a, b and c are regression coefficients). The use of polynomials allows the data to specify whether a trend exists, whether the trend is linear, cup or hat shaped and whether the overall trend is increasing or decreasing. As a second method, we estimate the trend by fitting a bent-cable regression to the remainder. Bent-cable regression is a type of piecewise linear regression for estimating the point of transition between two linear phases in a time series59,60. The model takes the form r = b0 + b1x + b2 q(x, τ, γ)60. Here r is the remainder, x is time, b0 is the initial intercept, b1 is the slope in phase 1, the slope in phase 2 is b2 − b1 and q is a function that defines the change point: \(q(x,\tau ,\gamma )=\frac{{(x-\tau +\gamma )}^{2}}{4\gamma }I(\tau -\gamma < \tau +\gamma )+(x-\tau )I(x > \tau +\gamma )\); τ represents the location of the change point and γ the span of the bent cable that joins the two linear phases; I(A) is an indicator function that returns 1 if A is true and 0 if A is false. The bent-cable model allows the data to specify whether a trend exists and whether there has been a switch in the trend, thereby allowing the identification of whether the trend is linear, cup or hat shaped and whether the overall trend is increasing or decreasing. Both the polynomial and bent-cable models were estimated using LaplacesDemon’s53 Adaptive Metropolis MCMC algorithm and vague priors, although for the bent-cable model we constrained τ to be in the middle 70% of the time series and γ to be at most two years.
The STL extraction of the seasonal components in the air temperature, soil temperature, soil moisture and solar radiation data (there is no stochasticity or seasonal trend in the CO2 data we used) allows us to simulate detrended time series d of these forcing variables as \(d=\bar{y}+s+{{{{N}}}}(\mu ,\sigma )\) where N(μ, σ) is a normally distributed random variable with mean and standard deviation estimated from the remainder r (we verified that r was well described by the normal distribution), \(\bar{y}\) is the mean of the data over the time series and s is the seasonal component extracted by STL. For CO2, the detrended time series is simply the average CO2 over the time series.
Data availability
The NDVI data used in this study are from the GIMMS 3g v.1 NDVI data product26,27, downloaded from https://ecocast.arc.nasa.gov/data/pub/gimms/3g.v1/ on 8 January 2019. The EVI data are from the MODIS Vegetation Indices data product28, downloaded from https://lpdaac.usgs.gov/products/mod13a2v006/ on 1 April 2020. The climate-system data were from the ERA5-Land monthly averaged data from 1950 to present32 downloaded from the Climate Data Store (https://doi.org/10.24381/cds.68d2bb3) on 3 July 2020. Annual historical atmospheric CO2 concentrations were taken from ISIMIP, downloaded 3 June 2020. In preliminary analyses we used fire data from the ESA Fire Disturbance Climate Change Initiative’s AVHRR LTDR Grid v.1.0 product49 from https://doi.org/10.5285/4f377defc2454db9b2a6d032abfd0cbd downloaded on 2 July 2020.
Code availability
A C version of the TTR-SDM growth model described in Methods that can be called from R is available at https://github.com/pfloek-bt/TTRcodeAttribution.
Change history
09 May 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41561-023-01196-1
References
Cramer, W. et al. Global response of terrestrial ecosystem structure and function to CO2 and climate change: results from six dynamic global vegetation models. Glob. Change Biol. 7, 357–373 (2001).
Parmesan, C. & Yohe, G. A globally coherent fingerprint of climate change impacts across natural systems. Nature 421, 37–42 (2003).
Menzel, A. et al. European phenological response to climate change matches the warming pattern. Glob. Change Biol. 12, 1969–1976 (2006).
Nolan, C. et al. Past and future global transformation of terrestrial ecosystems under climate change. Science 361, 920–923 (2018).
Zhang, T., Niinemets, U., Sheffield, J. & Lichstein, J. W. Shifts in tree functional composition amplify the response of forest biomass to climate. Nature 556, 99–102 (2018).
Bonan, G. B. & Doney, S. C. Climate, ecosystems, and planetary futures: the challenge to predict life in Earth system models. Science 359, eaam8328 (2018).
Settele, J. et al. in Climate Change 2014: Impacts, Adaptation, and Vulnerability (eds Fields, C. B. et al.) Ch. 4 (Cambridge Univ. Press, 2014).
Reich, P. B., Hobbie, S. E. & Lee, T. D. Plant growth enhancement by elevated CO2 eliminated by joint water and nitrogen limitation. Nat. Geosci. 7, 920–924 (2014).
Terrer, C., Vicca, S., Hungate, B. A., Phillips, R. P. & Prentice, I. C. Mycorrhizal association as a primary control of the CO2 fertilization effect. Science 353, 72–74 (2016).
Peñuelas, J. et al. Shifting from a fertilization-dominated to a warming-dominated period. Nat. Ecol. Evol. 1, 1438–1445 (2017).
Hovenden, M. J. et al. Globally consistent influences of seasonal precipitation limit grassland biomass response to elevated CO2. Nat. Plants 5, 167–173 (2019).
Bjorkman, A. D. et al. Plant functional trait change across a warming tundra biome. Nature 562, 57–62 (2018).
McManus, K. M. et al. Satellite-based evidence for shrub and graminoid tundra expansion in northern Quebec from 1986 to 2010. Glob. Change Biol. 18, 2313–2323 (2012).
Pan, N. et al. Increasing global vegetation browning hidden in overall vegetation greening: insights from time-varying trends. Remote Sens. Environ. 214, 59–72 (2018).
Myers-Smith, I. H. et al. Complexity revealed in the greening of the Arctic. Nat. Clim. Change 10, 106–117 (2020).
de Jong, R., Verbesselt, J., Schaepman, M. E. & de Bruin, S. Trend changes in global greening and browning: contribution of short-term trends to longer-term change. Glob. Change Biol. 18, 642–655 (2012).
Sulla-Menashe, D., Woodcock, C. E. & Friedl, M. A. Canadian boreal forest greening and browning trends: an analysis of biogeographic patterns and the relative roles of disturbance versus climate drivers. Environ. Res. Lett. 13, 014007 (2018).
Zhao, M. & Running, S. W. Drought-induced reduction in global terrestrial net primary production from 2000 through 2009. Science 329, 940–943 (2010).
Seddon, A. W. R., Macias-Fauria, M., Long, P. R., Benz, D. & Willis, K. J. Sensitivity of global terrestrial ecosystems to climate variability. Nature 531, 229–232 (2016).
Zhu, Z. C. et al. Greening of the Earth and its drivers. Nat. Clim. Change 6, 791–795 (2016).
Smith, W. K. et al. Large divergence of satellite and Earth system model estimates of global terrestrial CO2 fertilization. Nat. Clim. Change 6, 306–310 (2016).
Buitenwerf, R., Rose, L. & Higgins, S. I. Three decades of multi-dimensional change in global leaf phenology. Nat. Clim. Change 5, 364–368 (2015).
Wang, S. et al. Recent global decline of CO2 fertilization effects on vegetation photosynthesis. Science 370, 1295–1300 (2020).
Song, X.-P. et al. Global land change from 1982 to 2016. Nature 560, 639–643 (2018).
Jiang, M. et al. The fate of carbon in a mature forest under carbon dioxide enrichment. Nature 580, 227–231 (2020).
Tucker, C. J. et al. An extended AVHRR 8 km NDVI dataset compatible with MODIS and SPOT vegetation NDVI data. Int. J. Remote Sens. 26, 4485–4498 (2005).
Pinzon, J. E. & Tucker, C. J. A non-stationary 1981–2012 AVHRR NDVI3g time series. Remote Sens. 6, 6929–6960 (2014).
Didan, K. Mod13q1 V006 MODIS/Terra Vegetation Indices 16-Day l3 Global 1km SIN Grid V006. (NASA EOSDIS Land Processes DAAC, 2015). https://doi.org/10.5067/MODIS/MOD13A2.006
Thornley, J. H. Modelling shoot:root relations: the only way forward? Ann. Bot. 81, 165–171 (1998).
Higgins, S. I. et al. A physiological analogy of the niche for projecting the potential distribution of plants. J. Biogeogr. 39, 2132–2145 (2012).
Hersbach, H. et al. The ERA5 global reanalysis. Q. J. R. Meteorol. Soc. 146, 1999–2049 (2020).
Muñoz Sabater, J. et al. Era5-Land: a state-of-the-art global reanalysis dataset for land applications. Earth Syst. Sci. Data 13, 4349–4383 (2021).
Cressie, N. & Wikle, C. K. Statistics for Spatio-Temporal Data (Wiley & Sons, 2011).
Prentice, I. C. et al. in Terrestrial Ecosystems in a Changing World (eds Canadell, J. G. et al.) 175–192 (Springer, 2007).
Conradi, T. et al. An operational definition of the biome for global change research. N. Phytol. 227, 1294–1306. (2020).
Zhao, M. & Running, S. W. Response to comments on “Drought-induced reduction in global terrestrial net primary production from 2000 through 2009". Science 333, 1093–1093 (2011).
Reich, P. B. et al. Effects of climate warming on photosynthesis in boreal tree species depend on soil moisture. Nature 562, 263–267 (2018).
Hubau, W. et al. Asynchronous carbon sink saturation in African and Amazonian tropical forests. Nature 579, 80–87 (2020).
Odum, E. P. The strategy of ecosystem development. Science 164, 262–270 (1969).
McGroddy, M. E., Daufresne, T. & Hedin, L. O. Scaling of C:N:P stoichiometry in forests worldwide: implications of terrestrial Redfield-type ratios. Ecology 85, 2390–2401 (2004).
Higgins, S. I. & Scheiter, S. Atmospheric CO2 forces abrupt vegetation shifts locally, but not globally. Nature 488, 209–212 (2012).
Buermann, W. et al. Widespread seasonal compensation effects of spring warming on northern plant productivity. Nature 562, 110–114 (2018).
Smith, W. K., Fox, A. M., MacBean, N., Moore, D. J. P. & Parazoo, N. C. Constraining estimates of terrestrial carbon uptake: new opportunities using long-term satellite observations and data assimilation. N. Phytol. 225, 105–112 (2020).
Ainsworth, E. A. & Long, S. P. What have we learned from 15 years of free-air CO2 enrichment (FACE)? A meta-analytic review of the responses of photosynthesis, canopy properties and plant production to rising CO2. N. Phytol. 165, 351–372 (2005).
Porcar-Castell, A. et al. Linking chlorophyll a fluorescence to photosynthesis for remote sensing applications: mechanisms and challenges. J. Exp. Bot. 65, 4065–4095 (2014).
Magney, T. S. et al. Mechanistic evidence for tracking the seasonality of photosynthesis with solar-induced fluorescence. Proc. Natl Acad. Sci. USA 116, 11640–11645 (2019).
Farquhar, G. D., von Caemmerer, S. & Berry, J. A. A biochemical model of photosynthetic CO2 assimilation in leaves of C3 species. Planta 149, 78–90 (1980).
von Caemmerer, S. Biochemical Models of Leaf Photosynthesis (CSIRO, 2000).
Chuvieco, E., Pettinari, M., Otón, G., Storm, T. & Padilla Parellada, M. ESA Fire Climate Change Initiative (Fire-cci): AVHRR-LTDR Burned Area Grid Product Version 1.0. (Centre for Environmental Data Analysis, 2019); https://doi.org/10.5285/4f377defc2454db9b2a6d032abfd0cbd
Box, E. O., Holben, B. N. & Kalb, V. Accuracy of the AVHRR vegetation index as a predictor of biomass, primary productivity and net CO2 flux. Vegetatio 80, 71–89 (1989).
Wessels, K. J. et al. Relationship between herbaceous biomass and 1 km2 advanced very high resolution radiometer (AVHRR) NDVI in Kruger National Park, South Africa. Int. J. Remote Sens. 27, 951–973 (2006).
Zhu, X. & Liu, D. Improving forest aboveground biomass estimation using seasonal Landsat NDVI time-series. ISPRS J. Photogramm. Remote Sens. 102, 222–231 (2015).
LaplacesDemon: Complete Environment for Bayesian Inference R package v.16.1.4 (Statisticat-LLC, 2020).
Storn, R. & Price, K. Differential evolution–a simple and efficient heuristic for global optimization over continuous spaces. J. Glob. Optim. 11, 341–359 (1997).
Mullen, K., Ardia, D., Gil, D., Windover, D. & Cline, J. DEoptim: an R package for global optimization by differential evolution. J. Stat. Softw. https://doi.org/10.18637/jss.v040.i06 (2011).
Ardia, D., Boudt, K., Carl, P., Mullen, K. M. & Peterson, B. G. Differential evolution with DEoptim: an application to non-convex portfolio optimization. R J. 3, 27–34 (2011).
Cleveland, R. B., Cleveland, W. S., McRae, J. E. & Terpenning, I. STL: a seasonal-trend decomposition procedure based on Loess (with discussion). J. Off. Stat. 6, 3–73 (1990).
R Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2020).
Chiu, G., Lockhart, R. & Routledge, R. Bent-cable regression theory and applications. J. Am. Stat. Assoc. 101, 542–553 (2006).
Khan, S. A. & Kar, S. C. Generalized bent-cable methodology for changepoint data: a Bayesian approach. J. Appl. Stat. 45, 1799–1812 (2018).
Whittaker, R. H. Communities and Ecosystems (Macmillan, 1975).
Acknowledgements
S.I.H. acknowledges funding support from the BMBF SPACES project EMSAfrica grant number 01LL1801A and from the DFG project HI 1106/11-1. E.M. acknowledges a DAAD doctoral scholarship.
Funding
Open Access funding provided by Universität Bayreuth.
Author information
Authors and Affiliations
Contributions
S.I.H. designed the study, performed the analyses and led the writing. T.C. and E.M. performed the site selection and contributed to study design, interpretation of the analyses and writing.
Corresponding author
Ethics declarations
Competing interests
The authors no conflict of interest.
Peer review
Peer review information
Nature Geoscience thanks Sara Vicca, Steven Running and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Tamara Goldin and Simon Harold, in collaboration with the Nature Geoscience team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Graphical description of β parameters.
Graphical representation of how the β parameters define the influence of environmental forcing variables on rates in the plant growth model as described by equations 13 to 17. All units are rescaled to the range 0-1 and the β parameters represent positions on the x-axes. The effect of photosynthetically active radiation, air temperature and atmospheric CO2 on carbon assimilation is described by the Farquhar photosynthesis model. Two additional β parameters (βa and βb) which describe the the influence of site fertility on carbon and nitrogen assimilation are not represented in this diagram. In addition, for all analyses we set fire severity to zero, thereby ignoring the effects of fire.
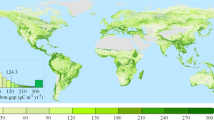
Extended Data Fig. 2 Distribution of the 100 study sites.
The different colours indicate the biome assignments made using the authors assessment of the Google Earth imagery. BF=boreal forest, GR=grassland, MT=Mediterranean type ecosystems, RF=tropical evergreen forest, SA=savanna, SH=shrubland, TF=temperate forest, TU=tundra.
Extended Data Fig. 3 Distribution of study sites in climate space.
The distribution of the 100 study sites in the temperature and mean annual precipitation climate space used by Whittaker61. The data points are superimposed on Whittaker’s61 biome scheme. BF=boreal forest, GR=grassland, MT=Mediterranean type ecosystems, RF=tropical evergreen forest, SA=savanna, SH=shrubland, TF=temperate forest, TU=tundra.
Extended Data Fig. 4 EVI time series analysis of vegetation activity for a savanna site in the Burkina Faso National Park.
(A) MODIS EVI time series of vegetation activity (data) and the state space model’s fit to these data (model). The blue polygon shows the 95% credible intervals around the mean model predictions which includes parameter, process and observation uncertainty. (B) Anomalies in the NDVI data (blue bars) and the fit of a bent-cable regression to these anomalies. The polygons show the 95 % credible intervals of the bent-cable regression predictions. (C) Zero intercept regression showing the model’s ability to predict observed anomalies, with full and detrended climate forcing data. Polygons show the 95% credible intervals of the mean regression line. (D) Posterior density of the change in the full model’s regression slope (as shown in panel C) caused by removing trends in a climate forcing variable from the full model; in this example the ability to predict anomalies was sensitive to soil moisture and soil temperature anomalies. Figure 1 shows an analogous plot using GIMMS NDVI data.
Extended Data Fig. 5 Distribution of NDVI and EVI anomaly response types.
The frequency of cup (initial decrease, subsequent increase), hat (initial increase, subsequent decrease), and linear time trends in the anomalies in NDVI and EVI. The number of anomaly trends that showed an overall increase or decrease over the time series are indicated by different colours. Overall increases or decreases less than 2 % in EVI were classified as small. The response types were classified from bent-cable and polynomial regression models fitted to the anomaly data from each site. Panels show results using polynomical regression on the NDVI data (a), bent-cable regression on the EVI data (b) and polynomial regression on the EVI data. Figure 2 shows results for the bent-cable regression on the NDVI data.
Extended Data Fig. 6 Comparison of anomaly response types for 2000-2015 for NDVI and EVI data.
Repeat of the analyses shown in Fig. 2 and Extended Data Fig. 5 using the common time window (years 2000-2015) for which both EVI and NDVI data are available. Shown are the frequencies of cup (initial decrease, subsequent increase), hat (initial increase, subsequent decrease), and linear time trends in the anomalies in vegetation indices. The number of anomaly trends that showed an overall increase or decrease over the time series are indicated by different colours (the category “small" as defined in Fig. 2 and Extended Data Fig. 5 was not detected in these analyses). These plots reveal that although the analysis time window influences the trends detected, that the EVI and NDVI data reveal similar trends, albeit with more decreasing trends in the NDVI data and more cup shaped trends when using bent-cable regression. The EVI and NDVI response shapes of individual sites agreed for 52% and 45% of cases (respectively for polynomial and bent-cable analyses). The EVI and NDVI response trends of individual sites agreed for 67% and 59% of cases (respectively for polynomial and bent-cable analyses).
Extended Data Fig. 7 Sensitivity of EVI vegetation anomalies to climate anomalies.
The sensitivity is quantified as the effect of each of five forcing factors (T air = air temperature, T soil = soil temperature, M soil = soil moisture, S rad = solar radiation and CO2 = atmospheric CO2 concentration) on the slope describing the ability of the model to predict anomalies in the vegetation activity time series (Extended Data Fig. 4). The slope with the full model is represented by the red colour ramp. Shown in the matrix are the 75 of 100 sites where anomalies in vegetation activity could be attributed to the environmental forcing factors. The coloured circles indicate the response groups the sites are assigned to by an unsupervised classification. The site names are codes indicating ecosystem type (BF=boreal forest, GR=grassland, MT=Mediterranean type ecosystems, RF=tropical evergreen forest, SA=savanna, SH=shrubland, TF=temperate forest, TU=tundra), country and site name. Red colours indicate high sensitivity, blue colours indicate low sensitivity. See Fig. 3 for the same analysis using NDVI data.
Extended Data Fig. 8 Ecosystem sensitivity classes in relation to climate (using EVI data).
(A) The distribution of attribution classes (Extended Data Fig. 7) in bivariate temperature and moisture climate space. Points represent the locations of the 100 study sites in climate space. Attribution classes are the two major groups classified in Extended Data Fig. 7. Sites labelled 0 are sites excluded from Extended Data Fig. 7 because the model could not explain observed anomalies. The colours of the sites labelled 1 and 2 correspond to the groups classified in Extended Data Fig. 7. The ellipses represent the fitted covariance estimates of the classes as estimated by discriminant analysis based on Gaussian finite mixture modelling. (B, C) The shape and direction of the anomaly response (as defined in Extended Data Fig. 5) plotted in bivariate climate space. The mean annual temperature and mean annual soil moisture were calculated over the study period using the ERA5 reanalysis data. Panels D, E and F plot the points in panels A, B and C in geographic space. See Fig. 4 for the same analysis using NDVI data.
Extended Data Fig. 9 Anomaly prediction with full and detrended climate data.
The slopes of the zero intercept regression of the model’s ability to predict anomalies in vegetation activity when using the full and detrended climate data (for example Fig. 1C). The points represent the mean of the posterior estimates of the slope, the tick marks span the 95 % credible intervals of the estimates. Attribution is diagnosed if the slopes are positive and clearly higher (no overlap in the credible intervals) for model runs forced by the full climate data than for model runs forced by climate data with trends removed. Panel (a) shows the results when using the NDVI data and panel (b) when using EVI data.
Extended Data Fig. 10 Summary of NDVI and EVI anomaly prediction with full and detrended climate data.
Probability density functions of the slopes of the zero intercept regression of the model’s ability to predict anomalies in vegetation activity when using the full and detrended climate data (shown in Extended Data Fig. 9). Solid lines are for all sites, broken lines are for the cases where attribution was diagnosed.Panel (a) shows the results when using the NDVI data and panel (b) when using EVI data.
Supplementary information
Supplementary Information
Supplementary Information, Tables 1–4 and the Farquhar photosynthesis model.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Higgins, S.I., Conradi, T. & Muhoko, E. Shifts in vegetation activity of terrestrial ecosystems attributable to climate trends. Nat. Geosci. 16, 147–153 (2023). https://doi.org/10.1038/s41561-022-01114-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41561-022-01114-x
This article is cited by
-
Earlier spring greening in Northern Hemisphere terrestrial biomes enhanced net ecosystem productivity in summer
Communications Earth & Environment (2024)
-
Machine Learning for Modeling Soil Organic Carbon as Affected by Land Cover Change in the Nebraska Sandhills, USA
Environmental Modeling & Assessment (2024)
-
Assessing the impact of dust events on the Holiday Climate Index in the Taklimakan Desert region
International Journal of Biometeorology (2024)
-
Increased impact of the El Niño–Southern Oscillation on global vegetation under future warming environment
Scientific Reports (2023)
-
Carbon ecological security assessment based on the decoupling relationship between carbon balance pressure and ecological quality in Xuzhou City, China
Environmental Science and Pollution Research (2023)