Abstract
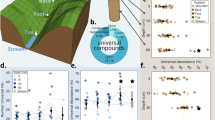
Dissolved organic matter affects fundamental biogeochemical processes in the soil such as nutrient cycling and organic matter storage. The current paradigm is that processing of dissolved organic matter converges to recalcitrant molecules (those that resist degradation) of low molecular mass and high molecular diversity through biotic and abiotic processes. Here we demonstrate that the molecular composition and properties of dissolved organic matter continuously change during soil passage and propose that this reflects a continual shifting of its sources. Using ultrahigh-resolution mass spectrometry and nuclear magnetic resonance spectroscopy, we studied the molecular changes of dissolved organic matter from the soil surface to 60 cm depth in 20 temperate grassland communities in soil type Eutric Fluvisol. Applying a semi-quantitative approach, we observed that plant-derived molecules were first broken down into molecules containing a large proportion of low-molecular-mass compounds. These low-molecular-mass compounds became less abundant during soil passage, whereas larger molecules, depleted in plant-related ligno-cellulosic structures, became more abundant. These findings indicate that the small plant-derived molecules were preferentially consumed by microorganisms and transformed into larger microbial-derived molecules. This suggests that dissolved organic matter is not intrinsically recalcitrant but instead persists in soil as a result of simultaneous consumption, transformation and formation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The compiled dataset used in our analyses is available at https://doi.org/10.17617/3.28 and root standing biomass at https://doi.org/10.1594/PANGAEA.880324. The raw data are available from the corresponding author (M.L.) on request.
Code availability
The codes used for this study are available on request.
References
Battin, T. J. et al. The boundless carbon cycle. Nat. Geosci. 2, 598–600 (2009).
Roulet, N. & Moore, T. R. Environmental chemistry. Browning the waters. Nature 444, 283–284 (2006).
Kalbitz, K., Solinger, S., Park, J. H., Michalzik, B. & Matzner, E. Controls on the dynamics of dissolved organic matter in soils: a review. Soil Sci. 165, 277–304 (2000).
Kaiser, K. & Kalbitz, K. Cycling downwards—dissolved organic matter in soils. Soil Biol. Biochem. 52, 29–32 (2012).
Brantley, S. L., Goldhaber, M. B. & Ragnarsdottir, K. V. Crossing disciplines and scales to understand the Critical Zone. Elements 3, 307–314 (2007).
Li, L. et al. Expanding the role of reactive transport models in critical zone processes. Earth Sci. Rev. 165, 280–301 (2017).
Sanderman, J., Baldock, J. A. & Amundson, R. Dissolved organic carbon chemistry and dynamics in contrasting forest and grassland soils. Biogeochemistry 89, 181–198 (2008).
Marschner, B. et al. How relevant is recalcitrance for the stabilization of organic matter in soils? J. Plant Nutr. Soil Sc. 171, 91–110 (2008).
von Luetzow, M. et al. Stabilization of organic matter in temperate soils: mechanisms and their relevance under different soil conditions—a review. Eur. J. Soil. Sci. 57, 426–445 (2006).
Dungait, J. A. J., Hopkins, D. W., Gregory, A. S. & Whitmore, A. P. Soil organic matter turnover is governed by accessibility not recalcitrance. Glob. Change Biol. 18, 1781–1796 (2012).
Don, A., Roedenbeck, C. & Gleixner, G. Unexpected control of soil carbon turnover by soil carbon concentration. Environ. Chem. Lett. 11, 407–413 (2013).
Leinemann, T. et al. Multiple exchange processes on mineral surfaces control the transport of dissolved organic matter through soil profiles. Soil Biol. Biochem. 118, 79–90 (2018).
Marschner, B. & Kalbitz, K. Controls of bioavailability and biodegradability of dissolved organic matter in soils. Geoderma 113, 211–235 (2003).
Fontaine, S. et al. Stability of organic carbon in deep soil layers controlled by fresh carbon supply. Nature 450, 277–280 (2007).
Schmidt, M. W. I. et al. Persistence of soil organic matter as an ecosystem property. Nature 478, 49–56 (2011).
Steinbeiss, S., Temperton, V. M. & Gleixner, G. Mechanisms of short-term soil carbon storage in experimental grasslands. Soil Biol. Biochem. 40, 2634–2642 (2008).
Kaiser, K., Guggenberger, G. & Haumaier, L. Changes in dissolved lignin-derived phenols, neutral sugars, uronic acids, and amino sugars with depth in forested Haplic Arenosols and Rendzic Leptosols. Biogeochemistry 70, 135–151 (2004).
Gleixner, G., Poirier, N., Bol, R. & Balesdent, J. Molecular dynamics of organic matter in a cultivated soil. Org. Geochem. 33, 357–366 (2002).
Gleixner, G. Soil organic matter dynamics: a biological perspective derived from the use of compound-specific isotopes studies. Ecol. Res. 28, 683–695 (2013).
Klotzbücher, T., Kalbitz, K., Cerli, C., Hernes, P. J. & Kaiser, K. Gone or just out of sight? The apparent disappearance of aromatic litter components in soils. Soil 2, 325–335 (2016).
Waggoner, D. C., Chen, H., Willoughby, A. S. & Hatcher, P. G. Formation of black carbon-like and alicyclic aliphatic compounds by hydroxyl radical initiated degradation of lignin. Org. Geochem. 82, 69–76 (2015).
DiDonato, N., Chen, H., Waggoner, D. & Hatcher, P. G. Potential origin and formation for molecular components of humic acids in soils. Geochim. Cosmochim. Acta 178, 210–222 (2016).
Saidy, A. R., Smernik, R. J., Baldock, J. A., Kaiser, K. & Sanderman, J. The sorption of organic carbon onto differing clay minerals in the presence and absence of hydrous iron oxide. Geoderma 209–210, 15–21 (2013).
Keiluweit, M. et al. Mineral protection of soil carbon counteracted by root exudates. Nat. Clim. Change 5, 588–595 (2015).
Lange, M. et al. Plant diversity increases soil microbial activity and soil carbon storage. Nat. Commun. 6, 6707 (2015).
Liang, C., Schimel, J. P. & Jastrow, J. D. The importance of anabolism in microbial control over soil carbon storage. Nat. Microbiol. 2, 17105 (2017).
Miltner, A., Bombach, P., Schmidt-Bruecken, B. & Kaestner, M. SOM genesis: microbial biomass as a significant source. Biogeochemistry 111, 41–55 (2012).
Koch, B. P. & Dittmar, T. From mass to structure: an aromaticity index for high-resolution mass data of natural organic matter. Rapid Commun. Mass Spectrom. 20, 926–932 (2006); erratum 30, 250 (2016).
Hertkorn, N. et al. High-precision frequency measurements: indispensable tools at the core of the molecular-level analysis of complex systems. Anal. Bioanal. Chem. 389, 1311–1327 (2007).
Magurran A. E. Measuring Biological Diversity (Blackwell, 2004).
Seidel, M. et al. Molecular-level changes of dissolved organic matter along the Amazon river-to-ocean continuum. Mar. Chem. 177, 218–231 (2015). Part 2.
Flerus, R. et al. A molecular perspective on the ageing of marine dissolved organic matter. Biogeosciences 9, 1935–1955 (2012).
Bandowe, B. A. M. et al. Plant diversity enhances the natural attenuation of polycyclic aromatic compounds (PAHs and oxygenated PAHs) in grassland soils. Soil Biol. Biochem. 129, 60–70 (2019).
Hertkorn, N. et al. Characterization of a major refractory component of marine dissolved organic matter. Geochim. Cosmochim. Acta 70, 2990–3010 (2006).
Einsiedl, F. et al. Rapid biotic molecular transformation of fulvic acids in a karst aquifer. Geochim. Cosmochim. Acta 71, 5474–5482 (2007).
Fellman, J. B., D’Amore, D. V. & Hood, E. Fluorescence characteristics and biodegradability of dissolved organic matter in forest and wetland soils from coastal temperate watersheds in southeast Alaska. Biogeochemistry 88, 169–184 (2008).
Sanderman, J., Maddern, T. & Baldock, J. Similar composition but differential stability of mineral retained organic matter across four classes of clay minerals. Biogeochemistry 121, 409–424 (2014).
Rasmussen, C. et al. Beyond clay: towards an improved set of variables for predicting soil organic matter content. Biogeochemistry 137, 297–306 (2018).
Jap, B. & Walian, P. Structure and functional mechanism of porins. Physiol. Rev. 76, 1073–1088 (1996).
Nikaido, H. Transport across the bacterial outer membrane. J. Bioenerg. Biomembr. 25, 581–589 (1993).
Lehmann, J. & Kleber, M. The contentious nature of soil organic matter. Nature 528, 60–68 (2015).
Osterholz, H., Niggemann, J., Giebel, H.-A., Simon, M. & Dittmar, T. Inefficient microbial production of refractory dissolved organic matter in the ocean. Nat. Commun. 6, 7422 (2015).
Amon, R. M. W. & Benner, R. Bacterial utilization of different size classes of dissolved organic matter. Limnol. Oceanogr. 41, 41–51 (1996).
Riedel, T., Zak, D., Biester, H. & Dittmar, T. Iron traps terrestrially derived dissolved organic matter at redox interfaces. Proc. Natl Acad. Sci. USA 110, 10101–10105 (2013).
Benk, S. A., Li, Y., Roth, V.-N. & Gleixner, G. Lignin dimers as potential markers for 14C-young terrestrial dissolved organic matter in the Critical Zone. Front. Earth Sci. 6, 168 (2018).
Neff, J. C. & Asner, G. P. Dissolved organic carbon in terrestrial ecosystems: synthesis and a model. Ecosystems 4, 29–48 (2001).
Roscher, C. et al. The role of biodiversity for element cycling and trophic interactions: an experimental approach in a grassland community. Basic Appl. Ecol. 5, 107–121 (2004).
Scheffer F. & Schachtschabel P. Lehrbuch der Bodenkunde (Spektrum Akademischer Verlag, 2002).
Bauriegel, A., Kühn, D., Schmidt, R., Hering, J. & Hannemann, J. Bodenübersichtskarte des Landes Brandenburg im Maßstab 1:300 000 (Kleinmachnow, Landesamt für Geowissenschaften und Rohstoffe, 2001).
Dittmar, T., Koch, B., Hertkorn, N. & Kattner, G. A simple and efficient method for the solid-phase extraction of dissolved organic matter (SPE-DOM) from seawater. Limnol. Oceanogr. Methods 6, 230–235 (2008).
Steinbeiss, S. et al. Plant diversity positively affects short-term soil carbon storage in experimental grasslands. Glob. Change Biol. 14, 2937–2949 (2008).
Ravenek, J. M. et al. Long-term study of root biomass in a biodiversity experiment reveals shifts in diversity effects over time. Oikos 123, 1528–1536 (2014).
Bligh, E. G. & Dyer, W. J. A rapid method of total lipid extraction and purification. Can. J. Biochem. Physiol. 37, 911–917 (1959).
Kramer, C. & Gleixner, G. Variable use of plant- and soil-derived carbon by microorganisms in agricultural soils. Soil Biol. Biochem. 38, 3267–3278 (2006).
Mellado-Vázquez, P. G., Lange, M. & Gleixner, G. Soil microbial communities and their carbon assimilation are affected by soil properties and season but not by plants differing in their photosynthetic pathways (C3 vs. C4). Biogeochemistry 142, 175–187 (2019).
Frostegard, A. & Baath, E. The use of phospholipid fatty acid analysis to estimate bacterial and fungal biomass in soil. Biol. Fertil. Soils 22, 59–65 (1996).
Zelles, L. Identification of single cultured micro-organisms based on their whole-community fatty acid profiles, using an extended extraction procedure. Chemosphere 39, 665–682 (1999).
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K. & Schloss, P. D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 79, 5112–5120 (2013).
Muyzer, G., Dewaal, E. C. & Uitterlinden, A. G. Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl. Environ. Microbiol. 59, 695–700 (1993).
Yu, Y., Lee, C., Kim, J. & Hwang, S. Group-specific primer and probe sets to detect methanogenic communities using quantitative real-time polymerase chain reaction. Biotechnol. Bioeng. 89, 670–679 (2005).
Ihrmark, K. et al. New primers to amplify the fungal ITS2 region—evaluation by 454-sequencing of artificial and natural communities. FEMS Microbiol. Ecol. 82, 666–677 (2012).
Gweon, H. S. et al. PIPITS: an automated pipeline for analyses of fungal internal transcribed spacer sequences from the Illumina sequencing platform. Methods Ecol. Evol. 6, 973–980 (2015).
Oksanen J. et al. vegan: Community ecology package. R package version 2.5-3 https://cran.r-project.org/web/packages/vegan/index.html (2015).
Malik, A. A. et al. Linking molecular size, composition and carbon turnover of extractable soil microbial compounds. Soil Biol. Biochem. 100, 66–73 (2016).
Pohlabeln, A. M. & Dittmar, T. Novel insights into the molecular structure of non-volatile marine dissolved organic sulfur. Mar. Chem. 168, 86–94 (2015).
Koch, B. P., Dittmar, T., Witt, M. & Kattner, G. Fundamentals of molecular formula assignment to ultrahigh resolution mass data of natural organic matter. Anal. Chem. 79, 1758–1763 (2007).
Stenson, A. C., Marshall, A. G. & Cooper, W. T. Exact masses and chemical formulas of individual Suwannee River fulvic acids from ultrahigh resolution electrospray ionization Fourier transform ion cyclotron resonance mass spectra. Anal. Chem. 75, 1275–1284 (2003).
R Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2016).
Bray, J. R. & Curtis, J. T. An ordination of the upland forest communities of southern Wisconsin. Ecol. Monogr. 27, 326–349 (1957).
Pinheiro, J., Bates, D., DebRoy, S., Sarkar, D. & R Development Core Team. nlme: Linear and nonlinear mixed effects models. R package version 3.1-137 https://cran.r-project.org/web/packages/nlme (2016).
Legendre, P. & Legendre, L. Numerical Ecology Vol. 20 (Elsevier, 1998).
Micallef, L. & Rodgers, P. eulerAPE: Drawing area-proportional 3-Venn diagrams using ellipses. PLOS ONE 9, e101717 (2014).
Hunt, J. F. & Ohno, T. Characterization of fresh and decomposed dissolved organic matter using excitation-emission matrix fluorescence spectroscopy and multiway analysis. J. Agric. Food Chem. 55, 2121–2128 (2007).
Merritt, K. A. & Erich, M. S. Influence of organic matter decomposition on soluble carbon and its copper-binding capacity. J. Environ. Qual. 32, 2122–2131 (2003).
Simon, C., Roth, V.-N., Dittmar, T. & Gleixner, G. Molecular signals of heterogeneous terrestrial environments identified in dissolved organic matter: a comparative analysis of orbitrap and ion cyclotron resonance mass spectrometers. Front. Earth Sci. 6, 138 (2018).
Chambers, M. C. et al. A cross-platform toolkit for mass spectrometry and proteomics. Nat. Biotechnol. 30, 918–920 (2012).
Strohalm, M., Kavan, D., Novak, P., Volny, M. & Havlicek, V. mMass 3: A cross-platform software environment for precise analysis of mass spectrometric data. Anal. Chem. 82, 4648–4651 (2010).
Acknowledgements
We thank U. Gerighausen for sampling and K. Klapproth for technical support with FT-ICR-MS measurements. This work was supported by the Zwillenberg-Tietz Stiftung and the Deutsche Forschungsgemeinschaft as part of the Critical Zone Observatory ‘AquaDiva’ (CRC 1076) and the Jena Experiment (FOR 1451, GL 262/14 and GL 262/19). The International Max Planck Research School for Global Biogeochemical Cycles (IMPRS-gBGC) provided the funding for the PhD scholarships of P.G.M.-V. and C.S.
Author information
Authors and Affiliations
Contributions
V.-N.R. and G.G. conceived and designed the study. M.L., V.-N.R. and G.G. wrote the main text. V.-R.N. and T.D. measured and processed MS data, N.H. obtained the NMR data. M.L. and V.-R.N. analysed the data. V.-N.R. and S.B. performed the supplementary decomposition experiment and C.S. measured, processed and analysed the data from the supplementary sites. L.M., N.J.O. and A.W. provided root standing biomass data, P.G.M.-V. provided data on microbial biomass, and R.I.G. and T.G. provided data on microbial diversity. All authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1−6 and Tables 1−4.
Rights and permissions
About this article
Cite this article
Roth, VN., Lange, M., Simon, C. et al. Persistence of dissolved organic matter explained by molecular changes during its passage through soil. Nat. Geosci. 12, 755–761 (2019). https://doi.org/10.1038/s41561-019-0417-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41561-019-0417-4
This article is cited by
-
Variations in the quantity and chemical composition of soil dissolved organic matter along a chronosequence of wolfberry plantations in an arid area of Northwest China
Chemical and Biological Technologies in Agriculture (2024)
-
Universal microbial reworking of dissolved organic matter along environmental gradients
Nature Communications (2024)
-
Humic Substances-Induced Changes in the Properties of Sb-Contaminated Soil and Effects on Sb Forms
Water, Air, & Soil Pollution (2024)
-
Soil carbon storage and accessibility drive microbial carbon use efficiency by regulating microbial diversity and key taxa in intercropping ecosystems
Biology and Fertility of Soils (2024)
-
Warming and flooding have different effects on organic carbon stability in mangrove soils
Journal of Soils and Sediments (2024)