Abstract
Hunting can fundamentally alter wildlife population dynamics but the consequences of hunting on pathogen transmission and evolution remain poorly understood. Here, we present a study that leverages a unique landscape-scale quasi-experiment coupled with pathogen-transmission tracing, network simulation and phylodynamics to provide insights into how hunting shapes feline immunodeficiency virus (FIV) dynamics in puma (Puma concolor). We show that removing hunting pressure enhances the role of males in transmission, increases the viral population growth rate and increases the role of evolutionary forces on the pathogen compared to when hunting was reinstated. Changes in transmission observed with the removal of hunting could be linked to short-term social changes while the male puma population increased. These findings are supported through comparison with a region with stable hunting management over the same time period. This study shows that routine wildlife management can have impacts on pathogen transmission and evolution not previously considered.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
DNA sequences are GenBank accessions MN563193–MN563239. All other data and code to perform the analysis are available on Github at https://github.com/nfj1380/Transmission-dynamics_huntingPumaFIV65
Code availability
The code and data to perform these operations as well as the transmission tree analysis above can be found here: https://github.com/nfj1380/Transmission-dynamics_huntingPumaFIV65
References
Packer, C. et al. Sport hunting, predator control and conservation of large carnivores. PLoS ONE 4, e5941 (2009).
Whitman, K., Starfield, A. M., Quadling, H. S. & Packer, C. Sustainable trophy hunting of African lions. Nature 428, 175–178 (2004).
Treves, A. Hunting for large carnivore conservation. J. Appl. Ecol. 46, 1350–1356 (2009).
Milner-Gulland, E. J. et al. Reproductive collapse in saiga antelope harems. Nature 422, 135 (2003).
Bischof, R. et al. Implementation uncertainty when using recreational hunting to manage carnivores. J. Appl. Ecol. 49, 824–832 (2012).
Booth, V. R., Masonde, J., Simukonda, C. & Cumming, D. H. M. Managing hunting quotas of African lions (Panthera leo): a case study from Zambia. J. Nat. Conserv. 55, 125817 (2020).
Potapov, A., Merrill, E. & Lewis, M. A. Wildlife disease elimination and density dependence. Proc. R. Soc. B 279, 3139–3145 (2012).
Lloyd-Smith, J. O. et al. Should we expect population thresholds for wildlife disease? Trends Ecol. Evol. 20, 511–519 (2005).
Beeton, N. & McCallum, H. Models predict that culling is not a feasible strategy to prevent extinction of Tasmanian devils from facial tumour disease. J. Appl. Ecol. 48, 1315–1323 (2011).
Choisy, M. & Rohani, P. Harvesting can increase severity of wildlife disease epidemics. Proc. R. Soc. B 273, 2025–2034 (2006).
Allendorf, F. W. & Hard, J. J. Human-induced evolution caused by unnatural selection through harvest of wild animals. Proc. Natl Acad. Sci. USA 106, 9987–9994 (2009).
Hamede, R. K., Bashford, J., McCallum, H. & Jones, M. Contact networks in a wild Tasmanian devil (Sarcophilus harrisii) population: using social network analysis to reveal seasonal variability in social behaviour and its implications for transmission of devil facial tumour disease. Ecol. Lett. 12, 1147–1157 (2009).
Woodroffe, R. et al. Culling and cattle controls influence tuberculosis risk for badgers. Proc. Natl Acad. Sci. USA 103, 14713–14717 (2006).
Carr, A. N. et al. Wildlife Management Practices Associated with Pathogen Exposure in Non-native Wild Pigs in Florida, U.S. (USDA National Wildlife Research Center, 2019).
Woodroffe, R., Cleaveland, S., Courtenay, O., Laurenson, M. K. & Artois, M. in The Biology and Conservation of Wild Canids 123–142 (Oxford Univ. Press, 2004).
Carter, S. P. et al. Culling-induced social perturbation in Eurasian badgers Meles meles and the management of TB in cattle: an analysis of a critical problem in applied ecology. Proc. R. Soc. B 274, 2769–2777 (2007).
Silk, M. J. et al. Contact networks structured by sex underpin sex-specific epidemiology of infection. Ecol. Lett. 21, 309–318 (2018).
Silk, M. J. et al. The application of statistical network models in disease research. Methods Ecol. Evol. 8, 1026–1041 (2017).
Morters, M. K. et al. Evidence-based control of canine rabies: a critical review of population density reduction. J. Anim. Ecol. 82, 6–14 (2013).
Lachish, S., McCallum, H., Mann, D., Pukk, C. E. & Jones, M. E. Evaluation of selective culling of infected individuals to control Tasmanian Devil facial tumor disease. Conserv. Biol. 24, 841–851 (2010).
Grubaugh, N. D. et al. Tracking virus outbreaks in the twenty-first century. Nat. Microbiol. 4, 10–19 (2019).
Didelot, X., Fraser, C., Gardy, J. & Colijn, C. Genomic infectious disease epidemiology in partially sampled and ongoing outbreaks. Mol. Biol. Evol. 34, msw075 (2017).
Smith, M. D. et al. Less is more: an adaptive branch-site random effects model for efficient detection of episodic diversifying selection. Mol. Biol. Evol. 32, 1342–1353 (2015).
Logan, K. A. & Runge, J. P. et al. Effects of hunting on a puma population in Colorado. Wildl. Monogr. 209, 1–35 (2020).
Biek, R., Pybus, O. G., Lloyd-Smith, J. O. & Didelot, X. Measurably evolving pathogens in the genomic era. Trends Ecol. Evol. 30, 306–313 (2015).
Pybus, O. G., Tatem, A. J. & Lemey, P. Virus evolution and transmission in an ever more connected world. Proc. R. Soc. B 282, 20142878 (2015).
Woolhouse, M. E. J., Adair, K. & Brierley, L. RNA viruses: a case study of the biology of emerging infectious diseases. Microbiol. Spectr. https://doi.org/10.1128/microbiolspec.oh-0001-2012 (2021).
Pybus, O. G. & Rambaut, A. Evolutionary analysis of the dynamics of viral infectious disease. Nat. Rev. Genet. 10, 540–550 (2009).
Fountain-Jones, N. M. et al. Towards an eco-phylogenetic framework for infectious disease ecology. Biol. Rev. 93, 950–970 (2018).
Webb, C. O. Exploring the phylogenetic structure of ecological communities: an example for rain forest trees. Am. Nat. 156, 145–155 (2000).
Biek, R. et al. Epidemiology, genetic diversity, and evolution of endemic feline immunodeficiency virus in a population of wild cougars. J. Virol. 77, 9578–9589 (2003).
Pedersen, N. C., Yamamoto, J. K., Ishida, T. & Hansen, H. Feline immunodeficiency virus infection. Vet. Immunol. Immunopathol. 21, 111–129 (1989).
Malmberg, J. L. et al. Altered lentiviral infection dynamics follow genetic rescue of the Florida panther. Proc. R. Soc. B 286, 20191689 (2019).
Elbroch, L. M., Levy, M., Lubell, M., Quigley, H. & Caragiulo, A. Adaptive social strategies in a solitary carnivore. Sci. Adv. 3, e1701218 (2017).
Sweanor, L. L., Logan, K. A. & Hornocker, M. G. Cougar dispersal patterns, metapopulation dynamics, and conservation. Conserv. Biol. 14, 798–808 (2000).
Fountain-Jones, N. M. et al. Linking social and spatial networks to viral community phylogenetics reveals subtype-specific transmission dynamics in African lions. J. Anim. Ecol. 86, 1469–1482 (2017).
Gilbertson, M. L. J. et al. Transmission of one predicts another: apathogenic proxies for transmission dynamics of a fatal virus. Preprint at bioRxiv https://doi.org/10.1101/2021.01.09.426055 (2021).
Fountain-Jones, N. M. et al. Host relatedness and landscape connectivity shape pathogen spread in a large secretive carnivore. Commun. Biol. 4, 12 (2021).
Hornocker, M. G. & Negri, S. Cougar: Ecology and Conservation (Univ. Chicago Press, 2010).
Moss, W. E., Alldredge, M. W. & Pauli, J. N. Quantifying risk and resource use for a large carnivore in an expanding urban-wildland interface. J. Appl. Ecol. 53, 371–378 (2016).
Trumbo, D. et al. Urbanization impacts apex predator gene flow but not genetic diversity across an urban-rural divide. Mol. Ecol. 28, 4926–4940 (2019).
VandeWoude, S. & Apetrei, C. Going wild: lessons from naturally occurring T-lymphotropic lentiviruses. Clin. Microbiol. Rev. 19, 728–762 (2006).
Logan, K. A. & Sweanor, L. L. Desert Puma: Evolutionary Ecology and Conservation of an Enduring Carnivore (Island Press, 2001).
Krakoff, E., Gagne, R. B., VandeWoude, S. & Carver, S. Variation in intra-individual lentiviral evolution rates: a systematic review of human, nonhuman primate, and felid species. J. Virol. https://doi.org/10.1128/JVI.00538-19 (2019).
Volz, E. M. & Didelot, X. Modeling the growth and decline of pathogen effective population size provides insight into epidemic dynamics and drivers of antimicrobial resistance. Syst. Biol. 67, 719–728 (2018).
Murrell, B. et al. Detecting individual sites subject to episodic diversifying selection. PLoS Genet. 8, e1002764 (2012).
Kenyon, J. C. & Lever, A. M. L. The molecular biology of feline immunodeficiency virus (FIV). Viruses 3, 2192–2213 (2011).
Tamuri, A. U., dos Reis, M., Hay, A. J. & Goldstein, R. A. Identifying changes in selective constraints: host shifts in influenza. PLoS Comput. Biol. 5, e1000564 (2009).
Forni, D., Cagliani, R., Clerici, M. & Sironi, M. Molecular evolution of human coronavirus genomes. Trends Microbiol. 25, 35–48 (2017).
Fountain-Jones, N. M. et al. Urban landscapes can change virus gene flow and evolution in a fragmentation-sensitive carnivore. Mol. Ecol. 26, 6487–6498 (2017).
Kozakiewicz, C. P. et al. Pathogens in space: advancing understanding of pathogen dynamics and disease ecology through landscape genetics. Evol. Appl. 11, 1763–1778 (2018).
McDonald, J. L., Smith, G. C., McDonald, R. A., Delahay, R. J. & Hodgson, D. Mortality trajectory analysis reveals the drivers of sex-specific epidemiology in natural wildlife–disease interactions. Proc. R. Soc. B 281, 20140526 (2014).
Gilbertson, M. L. J., Fountain-Jones, N. M. & Craft, M. E. Incorporating genomic methods into contact networks to reveal new insights into animal behaviour and infectious disease dynamics. Behaviour 155, 759–791 (2018).
Alldredge, M. W., Blecha, T. & Lewis, J. H. Less invasive monitoring of cougars in Colorado’s front range. Wildl. Soc. Bull. 43, 222–230 (2019).
Lewis, J. S. et al. The effects of urbanization on population density, occupancy, and detection probability of wild felids. Ecol. Appl. 25, 1880–1895 (2015).
Csárdi, G. & Nepusz, T. The igraph software package for complex network research. InterJournal Complex Syst. 1695, 1–9 (2006).
Didelot, X., Kendall, M., Xu, Y., White, P. J. & McCarthy, N. Genomic epidemiology analysis of infectious disease outbreaks using TransPhylo. Curr. Protoc. 1, e60 (2021).
Handcock, M. S., Hunter, D. R., Butts, C. T., Goodreau, S. M. & Morris, M. statnet: software tools for the representation, visualization, analysis and simulation of network data. J. Stat. Softw. 24, 1548 (2008).
Wertheim, J. O., Murrell, B., Smith, M. D., Pond, S. L. K. & Scheffler, K. RELAX: detecting relaxed selection in a phylogenetic framework. Mol. Biol. Evol. 32, 820–832 (2015).
Kosakovsky Pond, S. L. et al. A random effects branch-site model for detecting episodic diversifying selection. Mol. Biol. Evol. 28, 3033–3043 (2011).
Weaver, S. et al. Datamonkey 2.0: a modern web application for characterizing selective and other evolutionary processes. Mol. Biol. Evol. 35, 773–777 (2018).
Gill, M. S., Lemey, P., Bennett, S. N., Biek, R. & Suchard, M. A. Understanding past population dynamics: Bayesian coalescent-based modeling with covariates. Syst. Biol. 65, 1041–1056 (2016).
Faith, D. P. Conservation evaluation and phylogenetic diversity. Biol. Conserv. 61, 1–10 (1992).
Tsirogiannis, C. & Sandel, B. PhyloMeasures: a package for computing phylogenetic biodiversity measures and their statistical moments. Ecography 39, 709–714 (2016).
Fountain-Jones, N. nfj1380/Transmission-dynamics_huntingPumaFIV: (Puma-FIV_transmissionDynamics) (Zenodo, 2021); https://doi.org/10.5281/zenodo.5602162
Fountain-Jones, N. et al. Emerging phylogenetic structure of the SARS-CoV-2 pandemic. Virus Evol. 6, veaa082 (2020).
Karcher, M. D., Palacios, J. A., Bedford, T., Suchard, M. A. & Minin, V. N. Quantifying and mitigating the effect of preferential sampling on phylodynamic inference. PLoS Comput. Biol. 12, e1004789 (2016).
Acknowledgements
This project was funded by the National Science Foundation Ecology of Infectious Diseases research programme grants (DEB 1413925) (S.V., W.C.F., M.E.C., K.C. and S.C.) and an Australian Research Council Discovery Project Grant (DP190102020) (M.C., S.C., M.E.C. and S.V.). M.L.J.G. was supported by the Office of the Director, National Institutes of Health (NIH) under award no. NIH T32OD010993. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH. S.D. is supported by the Fonds National de la Recherche Scientifique (FNRS, Belgium). G.B. acknowledges support from the Interne Fondsen KU Leuven/Internal Funds KU Leuven under grant agreement C14/18/094 and the Research Foundation—Flanders (‘Fonds voor Wetenschappelijk Onderzoek—Vlaanderen’, G0E1420N). M.E.C. was funded by the National Science Foundation (DEB 1654609 and 2030509) and the College of Veterinary Medicine Research Office UMN Ag Experiment Station General Ag Research Funds. X.D. was supported by the National Institute for Health Research Health Protection Research Unit in Genomics and Enabling Data. Any use of trade, firm or product names is for descriptive purposes only and does not imply endorsement by the US Government.
Author information
Authors and Affiliations
Contributions
N.M.F.J. conducted the analysis and wrote the initial draft of the paper to which all authors contributed. K.L. and M.A. studied the puma populations in the field and provided the blood samples. S.K., D.R.T., P.S., R.G. and S.V. collected virus and host genetic data. S.D., G.B., M.C. and X.D. contributed to the phylogenetic and transmission tree analyses. M.L.J.G. contributed to the spatial analysis. M.E.C., S.V., K.C., W.C.F. and S.C. conceived of the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Animal care statement
Puma samples were collected as part of ongoing studies by CPW between 2006 and 2014. We handled all pumas in accordance with approved CPW ACUC capture and handling protocols (ACUC file no. 08-2004, ACUC protocol nos. 03-2007 and 16‐2008). Samples were provided to Colorado State University for diagnostic evaluation. Colorado State University and CPW Institutional Animal Care and Use Committees reviewed and approved this work before initiation (CSU IACUC protocol 05-061A).
Peer review information
Nature Ecology & Evolution thanks Pauline Kamath, Nichola Hill and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Probabilities of transmission between pairs of individuals in both regions.
The left panel (a) shows the treatment region and the right panel (b) shows the stable region.
Extended Data Fig. 2 Realized generation time distributions (time from infection to onward transmission) by region.
The top panel (a) shows the treatment region and the bottom panel (b) shows the stable region. In both regions, onward transmission events for FIVpco were most likely in the first two years after infection.
Extended Data Fig. 3 Estimated number of unsampled vs. sampled cases by region.
The top panel (a) shows the treatment region and the bottom panel (b) shows the stable region.
Extended Data Fig. 4 Histograms showing the expected homophily weighted degree distribution from our simulated networks by region (that is, just including edges from males-males) compared to observed homophily weight degree values.
The left panel (a) shows the treatment region and the right panel (b) shows the stable region. (see Methods -simulation modelling for details).
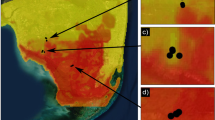
Extended Data Fig. 5 Time distributions for individuals involved in a putative transmission chain with the likely direction (red arrows) and the spatial context on each transmission event.
See Fig. 2 for other putative transmission events in the treatment region. Light yellow background: hunting pressure relieved, red background: hunting pressure resumed. White arrows in the maps indicate the likely transmission direction. Birth, death and sampling date is provided under each silhouette and estimated birth year is indicated by the black arrow. Colour of the boxes reflects transmission chain identity. M114 and F136 had overlapping home ranges and potentially transmitted FIVpco during mating event(s). M55 and F94 also had overlapping home ranges, and M55 was likely the sire of F94’s kittens; the pair consorted on 15 April 2010 and kittens were born on 15 July 2010. In addition, M55 associated with this family when the kittens were nurslings (K. Logan observation). Maps Data: Google ©2020.
Extended Data Fig. 6 Infection time distributions in the stable region for individuals involved in a putative transmission chain with the likely direction (red arrows) and the spatial context on each transmission event.
Colour of the boxes reflects transmission chain identity and grey boxes indicates sampling period. Sex, sampling date, birth date and death date of each individual are provided in each box when known. Maps Data: Google ©2020.
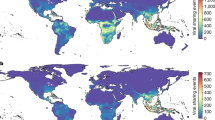
Extended Data Fig. 7 Map of our study regions.
Top panel: the location of all individuals sampled in 2005–2009 (no hunting in the treatment region). Bottom panel: the location of all individuals sampled (2005–2014 including the years when hunting was resumed in the treatment region). White diagonal lines show the broad extent of the Denver metropolitan area. Maps Data: Google ©2020.
Extended Data Fig. 8 Skyline plots showing the effective population size through time of dominant FIVpco lineages in each region.
The top panel (a) shows the treatment region and the bottom panel (b) shows the stable region. Grey shading provides the 95% high posterior density (HPD) estimates. a) Light yellow: hunting pressure relieved, red: hunting period, grey background: stable region. c) skygrowth plot from the management stable region showing FIVpco growth rate through time (see Fig. 3b in the main text for the corresponding plot from the treatment region). The dashed horizontal line reflects the 0-growth line.
Extended Data Fig. 9 FIVpco prevalence through time for each region. The top panel (a) shows the treatment region and the bottom panel (b) shows the stable region.
Numbers next to the points indicate how many samples were screened using qPCR each year. Confidence intervals were calculated using a binomial distribution and are only shown for total population estimates rather than for each sex (to aid interpretation). a) Light yellow: hunting pressure relieved, red: hunting period.
Extended Data Fig. 10 There were only insignificant relationships between FIVpco prevalence, growth rate and population size estimates.
(a) FIVpco growth rate (b) estimated puma population size, (c) female and (d) male population sizes in the treatment region. All data are scaled. Similar data was not available for the stable region. R: R2.
Supplementary information
Supplementary Information
Supplementary Tables 1–3 and references.
Rights and permissions
About this article
Cite this article
Fountain-Jones, N.M., Kraberger, S., Gagne, R.B. et al. Hunting alters viral transmission and evolution in a large carnivore. Nat Ecol Evol 6, 174–182 (2022). https://doi.org/10.1038/s41559-021-01635-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-021-01635-5
This article is cited by
-
Managing host-parasite interactions in humans and wildlife in times of global change
Parasitology Research (2022)