Abstract
Complex assemblages of microbes in the surface ocean are responsible for approximately half of global carbon fixation. The persistence of high taxonomic diversity despite competition for a small suite of relatively homogeneously distributed nutrients, that is, ‘the paradox of the plankton’, represents a long-standing challenge for ecological theory. Here we find evidence consistent with temporal niche partitioning of nitrogen assimilation processes over a diel cycle in the North Pacific Subtropical Gyre. We jointly analysed transcript abundances, lipids and metabolites and discovered that a small number of diel archetypes can explain pervasive periodic dynamics. Metabolic pathway analysis of identified diel signals revealed asynchronous timing in the transcription of nitrogen uptake and assimilation genes among different microbial groups—cyanobacteria, heterotrophic bacteria and eukaryotes. This temporal niche partitioning of nitrogen uptake emerged despite synchronous transcription of photosynthesis and central carbon metabolism genes and associated macromolecular abundances. Temporal niche partitioning may be a mechanism by which microorganisms in the open ocean mitigate competition for scarce resources, supporting community coexistence.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Sequence data for the >0.2 μm metatranscriptome have been deposited in the Sequence Read Archive through the National Center for Biotechnology Information under BioProject ID PRJNA358725. The Station ALOHA gene catalogue data are available under BioProject ID PRJNA352737 and iMicrobe (https://www.imicrobe.us/#/search/PRJNA352737). Sequence data for the >5 μm metatranscriptomes are available at the Sequence Read Archive under accession no. SRP136571 and BioProject no. PRJNA437978. Raw files for the metabolomics data are available at Metabolomics Workbench under Project ID PR000926. The lipidomics mass spectral raw data are available from the authors upon request.
Code availability
All code and feature/abundance tables used to complete this analysis are available at GitHub (https://github.com/WeitzGroup/community_scale_metabolism_NPSG) and are archived under https://zenodo.org/badge/latestdoi/262179139 (ref. 90).
References
Ottesen, E. A. et al. Pattern and synchrony of gene expression among sympatric marine microbial populations. Proc. Natl Acad. Sci. USA 110, E488–E497 (2013).
Muñoz-Marín, M. D. C. et al. The transcriptional cycle is suited to daytime N2 fixation in the unicellular cyanobacterium “Candidatus Atelocyanobacterium thalassa” (UCYN-A). mBio 10, e02495-18 (2019).
Vislova, A., Sosa, O. A., Eppley, J. M., Romano, A. E. & DeLong, E. F. Diel oscillation of microbial gene transcripts declines with depth in oligotrophic ocean waters. Front. Microbiol. 10, 2191 (2019).
Harke, M. J. et al. Periodic and coordinated gene expression between a diazotroph and its diatom host. ISME J. 13, 118–131 (2019).
Hernández Limón, M. D. et al. Transcriptional patterns of Emiliania huxleyi in the North Pacific Subtropical Gyre reveal the daily rhythms of its metabolic potential.Environ. Microbiol. 22, 381–396 (2020).
Becker, K. W. et al. Daily changes in phytoplankton lipidomes reveal mechanisms of energy storage in the open ocean. Nat. Commun. 9, 5179 (2018).
Frischkorn, K. R., Haley, S. T. & Dyhrman, S. T. Coordinated gene expression between Trichodesmium and its microbiome over day–night cycles in the North Pacific Subtropical Gyre. ISME J. 12, 997–1007 (2018).
Ottesen, E. A. et al. Ocean microbes. Multispecies diel transcriptional oscillations in open ocean heterotrophic bacterial assemblages. Science 345, 207–212 (2014).
Wilson, S. T. et al. Coordinated regulation of growth, activity and transcription in natural populations of the unicellular nitrogen-fixing cyanobacterium Crocosphaera. Nat. Microbiol. 2, 17118 (2017).
Saito, M. A. et al. Iron conservation by reduction of metalloenzyme inventories in the marine diazotroph Crocosphaera watsonii. Proc. Natl Acad. Sci. USA 108, 2184–2189 (2011).
Strenkert, D. et al. Multiomics resolution of molecular events during a day in the life of Chlamydomonas. Proc. Natl Acad. Sci. USA 116, 2374–2383 (2019).
Boysen, A. K. et al. Particulate metabolites and transcripts reflect diel oscillations of microbial activity in the surface ocean. mSystems 6, e00896-20 (2021).
White, A. E., Barone, B., Letelier, R. M. & Karl, D. M. Productivity diagnosed from the diel cycle of particulate carbon in the North Pacific Subtropical Gyre: optically derived productivity. Geophys. Res. Lett. 44, 3752–3760 (2017).
DeLong, E. F. et al. Community genomics among stratified microbial assemblages in the ocean’s interior. Science 311, 496–503 (2006).
Sunagawa, S. et al. Ocean plankton. Structure and function of the global ocean microbiome. Science 348, 1261359 (2015).
Coles, V. J. et al. Ocean biogeochemistry modeled with emergent trait-based genomics. Science 358, 1149–1154 (2017).
Walbauer, J. R., Rodrigue, S., Coleman, M. L. & Chisholm, S. W. Transcriptome and proteome dynamics of a light–dark synchronized bacterial cell cycle.PLoS ONE 7, e43432 (2012).
Steiner, P. A. et al. Highly variable mRNA half-life time within marine bacterial taxa and functional genes. Environ. Microbiol. 21, 3873–3884 (2019).
Moran, M. A. et al. Sizing up metatranscriptomics. ISME J. 7, 237–243 (2013).
Tamames, J., Cobo-Simón, M. & Puente-Sánchez, F. Assessing the performance of different approaches for functional and taxonomic annotation of metagenomes. BMC Genomics 20, 960 (2019).
DiTullio, G. R. & Laws, E. A. Diel periodicity of nitrogen and carbon assimilation in five species of marine phytoplankton: accuracy of methodology for predicting N-assimilation rates and N/C composition ratios. Mar. Ecol. Prog. Ser. 32, 123–132 (1986).
Granum, E., Kirkvold, S. & Myklestad, S. M. Cellular and extracellular production of carbohydrates and amino acids by the marine diatom Skeletonema costatum: diel variations and effects of N depletion. Mar. Ecol. Prog. Ser. 242, 83–94 (2002).
Lacour, T., Sciandra, A., Talec, A., Mayzaud, P. & Bernard, O. Diel variations of carbohydrates and neutral lipids in nitrogen-sufficient and nitrogen-starved cyclostat cultures of Isochrysis sp. J. Phycol. 48, 966–975 (2012).
Follett, C. L., Dutkiewicz, S., Karl, D. M., Inomura, K. & Follows, M. J. Seasonal resource conditions favor a summertime increase in North Pacific diatom–diazotroph associations. ISME J. 12, 1543–1557 (2018).
Chen, W.-N. U. et al. Diel rhythmicity of lipid-body formation in a coral-Symbiodinium endosymbiosis. Coral Reefs 31, 521–534 (2012).
Zhou, X. & Mopper, K. Photochemical production of low-molecular-weight carbonyl compounds in seawater and surface microlayer and their air-sea exchange. Mar. Chem. 56, 201–213 (1997).
Durham, B. P. et al. Sulfonate-based networks between eukaryotic phytoplankton and heterotrophic bacteria in the surface ocean.Nat. Microbiol. 4, 1706–1715 (2019).
Lambert, S. et al. Rhythmicity of coastal marine picoeukaryotes, bacteria and archaea despite irregular environmental perturbations. ISME J. 13, 388–401 (2019).
Kolody, B. C. et al. Diel transcriptional response of a California Current plankton microbiome to light, low iron, and enduring viral infection. ISME J. 13, 2817–2833 (2019).
Aylward, F. O. et al. Microbial community transcriptional networks are conserved in three domains at ocean basin scales. Proc. Natl Acad. Sci. USA 112, 5443–5448 (2015).
Rusch, D. B. et al. The Sorcerer II Global Ocean Sampling expedition: northwest Atlantic through eastern tropical Pacific. PLoS Biol. 5, e77 (2007).
Bork, P. et al. Tara Oceans studies plankton at planetary scale. Science 348, 873 (2015).
Delmont, T. O. et al. Nitrogen-fixing populations of Planctomycetes and Proteobacteria are abundant in surface ocean metagenomes. Nat. Microbiol. 3, 804–813 (2018).
Fuhrman, J. A. et al. Annually reoccurring bacterial communities are predictable from ocean conditions. Proc. Natl Acad. Sci. USA 103, 13104–13109 (2006).
Morris, R. M. et al. Temporal and spatial response of bacterioplankton lineages to annual convective overturn at the Bermuda Atlantic Time‐series Study site. Limnol. Oceanogr. 50, 1687–1696 (2005).
Mende, D. R. et al. Environmental drivers of a microbial genomic transition zone in the ocean’s interior. Nat. Microbiol. 2, 1367–1373 (2017).
Keeling, P. J. et al. The Marine Microbial Eukaryote Transcriptome Sequencing Project (MMETSP): illuminating the functional diversity of eukaryotic life in the oceans through transcriptome sequencing. PLoS Biol. 12, e1001889 (2014).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457–D462 (2016).
Thaben, P. F. & Westermark, P. O. Detecting rhythms in time series with RAIN. J. Biol. Rhythms 29, 391–400 (2014).
Cuhel, R. L., Ortner, P. B. & Lean, D. R. S. Night synthesis of protein by algae. Limnol. Oceanogr. 29, 731–744 (1984).
Coesel, S. N. et al. Diel transcriptional oscillations of light-sensitive regulatory elements in open-ocean eukaryotic plankton communities. Proc. Natl Acad. Sci. USA 118, e2011038118 (2021).
Bolay, P., Muro-Pastor, M. I., Florencio, F. J. & Klähn, S. The distinctive regulation of cyanobacterial glutamine synthetase. Life (Basel) 8, 52 (2018).
Karl, D. M., Church, M. J., Dore, J. E., Letelier, R. M. & Mahaffey, C. Predictable and efficient carbon sequestration in the North Pacific Ocean supported by symbiotic nitrogen fixation. Proc. Natl Acad. Sci. USA 109, 1842–1849 (2012).
Berman, T. & Bronk, D. A. Dissolved organic nitrogen: a dynamic participant in aquatic ecosystems. Aquat. Microb. Ecol. 31, 279–305 (2003).
Lee, C. & Bada, J. L. Amino acids in equatorial Pacific Ocean water. Earth Planet. Sci. Lett. 26, 61–68 (1975).
Bada, J. L. & Lee, C. Decomposition and alteration of organic compounds dissolved in seawater. Mar. Chem. 5, 523–534 (1977).
Poretsky, R. S., Sun, S., Mou, X. & Moran, M. A. Transporter genes expressed by coastal bacterioplankton in response to dissolved organic carbon. Environ. Microbiol. 12, 616–627 (2010).
Berthelot, H. et al. NanoSIMS single cell analyses reveal the contrasting nitrogen sources for small phytoplankton. ISME J. 13, 651–662 (2019).
Moore, L. R., Post, A. F., Rocap, G. & Chisholm, S. W. Utilization of different nitrogen sources by the marine cyanobacteria Prochlorococcus and Synechococcus. Limnol. Oceanogr. 47, 989–996 (2002).
Hu, S. K., Connell, P. E., Mesrop, L. Y. & Caron, D. A. A hard day’s night: diel shifts in microbial eukaryotic activity in the North Pacific Subtropical Gyre. Front. Mar. Sci. https://doi.org/10.3389/fmars.2018.00351 (2018).
Hannides, C. C. S., Popp, B. N., Choy, C. A. & Drazen, J. C. Midwater zooplankton and suspended particle dynamics in the North Pacific Subtropical Gyre: a stable isotope perspective. Limnol. Oceanogr. 58, 1931–1946 (2013).
Becker, K. W. et al. Combined pigment and metatranscriptomic analysis reveals highly synchronized diel patterns of phenotypic light response across domains in the open oligotrophic ocean.ISME J. 15, 520–533 (2021).
Mruwat, N. et al. A single-cell polony method reveals low levels of infected Prochlorococcus in oligotrophic waters despite high cyanophage abundances. ISME J. 15, 41–54 (2021).
Chesson, P. L. & Warner, R. R. Environmental variability promotes coexistence in lottery competitive systems. Am. Nat. 117, 923–943 (1981).
Shmida, A. & Ellner, S. Coexistence of plant species with similar niches. Vegetatio 58, 29–55 (1984).
Ellner, S. P., Snyder, R. E. & Adler, P. B. How to quantify the temporal storage effect using simulations instead of math. Ecol. Lett. 19, 1333–1342 (2016).
Adler, P. B., Fajardo, A., Kleinhesselink, A. R. & Kraft, N. J. B. Trait-based tests of coexistence mechanisms. Ecol. Lett. 16, 1294–1306 (2013).
Adler, P. B., HilleRisLambers, J., Kyriakidis, P. C., Guan, Q. & Levine, J. M. Climate variability has a stabilizing effect on the coexistence of prairie grasses. Proc. Natl Acad. Sci. USA 103, 12793–12798 (2006).
Cáceres, C. E. Temporal variation, dormancy, and coexistence: a field test of the storage effect. Proc. Natl Acad. Sci. USA 94, 9171–9175 (1997).
Padisák, J. Identification of relevant time-scales in non-equilibrium community dynamics: conclusions from phytoplankton surveys. N. Z. J. Ecol. 18, 169–176 (1994).
Anderies, J. M. & Beisner, B. E. Fluctuating environments and phytoplankton community structure: a stochastic model. Am. Nat.155, 556–569 (2000).
Wagg, C. et al. Functional trait dissimilarity drives both species complementarity and competitive disparity. Funct. Ecol. 31, 2320–2329 (2017).
Bligh, E.G. & Dyer, W. J. A rapid method of total lipid extraction and purification. Can. J. Biochem. Physiol. 37, 911–917 (1959).
Boysen, A. K., Heal, K. R., Carlson, L. T. & Ingalls, A. E. Best-matched internal standard normalization in liquid chromatography–mass spectrometry metabolomics applied to environmental samples. Anal. Chem. 90, 1363–1369 (2018).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Fountoulakis, M. & Lahm, H. W. Hydrolysis and amino acid composition analysis of proteins. J. Chromatogr. A 826, 109–134 (1998).
Popendorf, K. J., Fredricks, H. F. & Van Mooy, B. A. S. Molecular ion-independent quantification of polar glycerolipid classes in marine plankton using triple quadrupole MS. Lipids 48, 185–195 (2013).
Collins, J. R., Edwards, B. R., Fredricks, H. F. & Van Mooy, B. A. S. LOBSTAHS: an adduct-based lipidomics strategy for discovery and identification of oxidative stress biomarkers. Anal. Chem. 88, 7154–7162 (2016).
Hummel, J. et al. Ultra performance liquid chromatography and high resolution mass spectrometry for the analysis of plant lipids. Front. Plant Sci. 2, 54 (2011).
Smith, C. A., Want, E. J., O’Maille, G., Abagyan, R. & Siuzdak, G. XCMS: processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal. Chem. 78, 779–787 (2006).
Kuhl, C., Tautenhahn, R., Böttcher, C., Larson, T. R. & Neumann, S. CAMERA: an integrated strategy for compound spectra extraction and annotation of liquid chromatography/mass spectrometry data sets. Anal. Chem. 84, 283–289 (2012).
Biller, S. J. et al. Prochlorococcus extracellular vesicles: molecular composition and adsorption to diverse microbes.Environ. Microbiol. https://doi.org/10.1111/1462-2920.15834 (2021).
Aylward, F. O. et al. Diel cycling and long-term persistence of viruses in the ocean’s euphotic zone. Proc. Natl Acad. Sci. USA 114, 11446–11451 (2017).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Masella, A. P., Bartram, A. K., Truszkowski, J. M., Brown, D. G. & Neufeld, J. D. PANDAseq: paired-end assembler for illumina sequences. BMC Bioinformatics 13, 31 (2012).
Joshi, N. & Fass, J. Sickle: A sliding-window, adaptive, quality-based trimming tool for FastQ files. Version 1.33. GitHub https://github.com/najoshi/sickle (2015).
Kopylova, E., Noé, L. & Touzet, H. SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data. Bioinformatics 28, 3211–3217 (2012).
Kiełbasa, S. M., Wan, R., Sato, K., Horton, P. & Frith, M. C. Adaptive seeds tame genomic sequence comparison. Genome Res. 21, 487–493 (2011).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010).
Alexander, H. et al. Functional group-specific traits drive phytoplankton dynamics in the oligotrophic ocean. Proc. Natl Acad. Sci. USA 112, E5972–E5979 (2015).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Meinicke, P. UProC: tools for ultra-fast protein domain classification. Bioinformatics 31, 1382–1388 (2015).
Mende, D. R., Boeuf, D. & DeLong, E. F. Persistent core populations shape the microbiome throughout the water column in the North Pacific Subtropical Gyre. Front. Microbiol. 10, 2273 (2019).
White, A. E. et al. Phenology of particle size distributions and primary productivity in the North Pacific subtropical gyre (Station ALOHA). J. Geophys. Res. Oceans 120, 7381–7399 (2015).
Borchers, H. W. pracma: Practical numerical math functions. R package version 2 https://cran.r-project.org/web/packages/pracma/index.html (2019).
Maechler, M., Rousseeuw, P., Struyf, A., Hubert, M. & Hornik, K. cluster: Cluster analysis basics and extensions. R package version 1.56 (2012).
Wehrens, R. & Buydens, L. M. C. Self- and super-organizing maps in R: the Kohonen package. J. Stat. Softw. 21, 1–19 (2007).
Hennig, C. fpc: Flexible procedures for clustering. R package version 2.2-9 (2010).
Muratore, D. Code for complex marine microbial communities partition metabolism of scarce resources over the diel cycle. Zenodo https://doi.org/10.5281/zenodo.3817416 (2020).
Acknowledgements
We thank T. Clemente and E. Wood-Charlson for facilitating data collection. We also thank the captains and crew of the R/V Kilo Moana and research staff at the School of Ocean and Earth Science and Technology and S. Haley and K. Frischkorn for sample collection of eukaryotic transcriptomes. This work was supported by grants from the Simons Foundation as part of the SCOPE collaboration: no. 329108 to E.V.A., E.F.D., D.M.K., A.E.W., J.P.Z., A.E.I., B.A.S.V.M., S.T.D. and J.S.W.; no. 721244 to E.V.A.; no. 721223 to E.F.D.; no. 721252 to D.M.K.; no. 721256 to A.E.W.; no. 724220 to J.P.Z.; no. 723787 to A.E.I.; no. 721229 to B.A.S.V.M.; no. 721225 to S.T.D.; and no. 721231 to J.S.W. A.K.B. was supported by a National Science Foundation Graduate Research Fellowship. K.W.B. was further supported by the Postdoctoral Scholarship Program at the Woods Hole Oceanographic Institution & US Geological Survey and J.R.C. was supported by the Simons Collaboration on Computational Biogeochemical Modeling of Marine Ecosystems (Simons Foundation grant no. 549894).
Author information
Authors and Affiliations
Contributions
A.K.B., F.O.A., A.V., A.E.W. and S.T.D. contributed to data collection. A.K.B., M.J.H., K.W.B., F.O.A., J.M.E., D.R.M., A.E.I., B.A.S.V.M. and S.T.D. contributed to sample processing and data preparation. S.T.W. served as chief scientist for the research expedition. D.M., A.K.B., M.J.H., K.W.B., J.R.C., S.N.C., D.R.M., S.T.D. and J.S.W. developed the data analysis methods. D.M., A.K.B. and S.N.C. wrote the code. D.M., S.N.C., S.J.B., S.P., R.A.R.-G., A.E.I. and J.S.W. contributed to analysis design. D.M., A.K.B., M.J.H., J.R.C., B.A.S.V.M., A.E.I., S.T.D. and J.S.W. analysed the data. E.F.D., A.E.I., B.A.S.V.M., S.T.D. and J.S.W. designed the research, with contributions from all authors. D.M. and J.S.W. led the writing of the manuscript. A.K.B., M.J.H., K.W.B., D.R.M. and S.J.B. contributed to writing the manuscript. D.M., A.K.B., M.J.H., K.W.B., J.R.C., S.N.C., F.O.A., A.V., D.R.M., S.T.W., S.J.B., E.V.A., E.F.D., D.M.K., A.E.W., J.P.Z., A.E.I., B.A.S.V.M., S.T.D. and J.S.W. edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Ecology & Evolution thanks Daniel Garza and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Comparison of cluster metrics for archetype clustering.
Comparison of cluster metrics for archetype clustering. Top panels show dynamics in clustering Calinski-Harabasz index and average silhouette width for increasing number of clusters comparing 3 clustering algorithms – self-organizing maps (SOM), hierarchical clustering (HC), and clustering about perimedoids (PAM). SOM was chosen for further clustering based on the advantage in C-H index and average silhouette width (A). The number of clusters was then selected based on the plateau in average silhouette width between four and five clusters. We then used ordered dissimilarity images (ODIs) to compare the four-cluster and five-cluster results (B). For additional information, silhouette profiles were constructed for all clusters in both clusterings. The 4 cluster clustering was chosen on the heuristic basis of higher maximum silhouette widths for all clusters in the 4 cluster SOM and fewer negative silhouette widths in all clusters (using negative silhouette width as a proxy for misclassification). Summary statistics for silhouette profiles are provided in Supplementary Data Item 6. Briefly, the SOM using 4 clusters had 235/6273 (3.7%) of silhouette widths less than 0 and maximum per-cluster silhouette widths between 0.322–0.456, while the SOM using 5 clusters had 353/6273 (5.6%) of silhouette widths less than 0 and maximum per-cluster silhouette widths between 0.265–0.419, indicating fewer misclassifications in the 4 cluster SOM and greater within-cluster similarity.
Extended Data Fig. 2 Time series of hydrolysable nitrogenous bases (top) and amino acids (bottom).
Details on quantification are available in Methods. Samples were measured in technical triplicates, with amino acids being separately measured in a biological duplicate. Time series shows mean total concentration with one standard deviation. Both THNB and THAA were detrended and RAIN analysis was conducted for 24-hour diel periodicity using replicates (p < 1e-6,p = 0.00034, respectively). These data were measured and incorporated into our analysis after the initial SOM clustering was conducted, so the detrended and z-score transformed time series were fit to the existing SOM model, and both THNB and THAA were assigned to the night cluster.
Extended Data Fig. 3 Sampling distribution for taxon synchrony hypothesis test.
As described in Methods section “Assessing Average Peak Time Rank Difference,” for each KEGG orthologue with diel transcript abundance from at least 4 taxa, the average of all pairwise differences in peak time was calculated (points and boxplot to the left on each panel). To assess the significance of the calculated difference in peak time, empirical peak time distributions were calculated by randomly selecting groups of diel transcripts using a fixed number of taxa. We calculated the average of pairwise differences for these randomly selected transcripts and repeated the process 10,000 times for each hypothesis test conducted. The aggregate of all testing distributions is shown as a histogram on the right-hand side of each figure. A one-sided test of difference was conducted to identify those KEGG orthologues with a lower average peak time difference than could be expected by randomly selecting groups of diel orthologues of that size from the data (synchronous). Multiple testing corrections were conducted using the adaptive Benjamini-Hochberg method for a false discovery rate (FDR) of 0.1. Orthologues deemed significantly synchronous after this procedure are marked in orange. The dashed line shows the corresponding portion of the tail in the distribution where the highest average peak time difference was still deemed significant. The transcripts marked in main text Fig. 5a are also labeled.
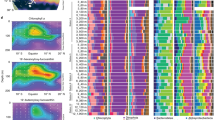
Extended Data Fig. 4 Diel asynchrony amongst ammonia-assimilation genes.
All taxa were examined for the presence/absence of transcripts for the amt ammonia transporter, and all KEGG orthologues from the nitrogen metabolism/arginine biosynthesis pathways associated with reactions involving ammonia, glutamine, and glutamate (as well as the glnK PII regulatory gene). Transcripts that were present in the data but not identified to have diel changes in abundance are shown in gray. Colored blocks indicate taxa that did have diel transcript abundance for that transcript, and color indicates the associated peak time. All peak times were represented at least once in this set of diel transcripts associated with ammonia-assimilation genes. Note that taxa in Extended Data Figs. 4 and 5 are not equivalent; for this figure taxa are selected based on presence of ammonia-assimilation genes, irrespective of diel status.
Extended Data Fig. 5 Diel synchrony amongst photosynthesis-related genes.
All taxa were examined for the presence/absence of transcripts for photosystem subunits as defined by the KEGG photosynthesis pathway (photosystem I and photosystem II D1, D2, cp43, cp47, and cytochrome b559 shown). Transcripts that were present in the data but not identified to have diel changes in abundance are shown in gray. Colored blocks indicate taxa that did have diel transcript abundance for that transcript, and color indicates the associated peak time. Peak times for diel transcripts amongst photosynthesis-related genes were only found in the daytime (0600, 1000, and 1400). Note that taxa in Extended Data Figs. 4 and 5 are not equivalent; for this figure taxa are selected based on presence of photosynthesis-related genes, irrespective of diel status.
Supplementary information
Supplementary Information
Supplementary Discussion and Tables 1–3.
Supplementary Data 1
Summary of KEGG pathway enrichment analysis results for eukaryotic (a,d), prokaryotic (non-photoautotroph) (b,e) and prokaryotic (photoautotroph) (c,f). Tables a–c indicate the enrichment analysis results for the top 20 KEGG pathways that had the most diel orthologues for each group in each cluster. Tables d–f show all pathways with a significant enrichment in each cluster using a Benjamini–Hochberg FDR of 10%. Values in columns are the number of KEGG orthologue IDs (KEGG orthologue) in the defined pathway in the indicated cluster, the number of KEGG orthologues in the defined pathway assigned to different clusters, the number of KEGG orthologues in other pathways assigned to the indicated cluster and the number of KEGG orthologues from other pathways assigned to other clusters. These are the four quantities used to calculate the test statistic for Fisher’s exact test, for which the P value and the Benjamini–Hochberg-adjusted P value are reported.
Supplementary Data 2
Files containing the results from the RAIN analysis for all analysed transcripts, lipids and metabolites separated by data source. Diel YES/NO indicates a Benjamini–Hochberg-adjusted P value at 0.05. The extended data file for transcripts also contains transcript KEGG annotation and putative assigned pathway. Because pathways for diel KEGG orthologues were manually curated, some pathways are left as indeterminate. For lipid and metabolite data, columns indicate the lipid/metabolite name, Benjamini–Hochberg-adjusted P value and YES/NO for whether the signal reaches the significance threshold under multiple testing.
Supplementary Data 3
Table detailing for each taxon studied how many unique KEGG orthologues were observed (number_kos_analyzed), how many of them were diel (number_diel_kos), the proportion of total observed KEGG orthologues for that taxon that were diel (proportion_kos_diel) and whether they came from the small size fraction or large size fraction transcriptomes (fraction). The term ‘Ind’ refers to indeterminate taxonomic classification.
Supplementary Data 4
Summary statistics of silhouette profiles comparing SOM clustering with 3,4 and 5 clusters. The rows represent the statistics for one of the clusters in each possible clustering (3, 4, 5). The columns provide an ID for that cluster in that clustering (clus4id), the maximum silhouette width of a signal in that cluster (max_width), the minimum silhouette width of a signal in that cluster (min_width), the total number of signals with negative silhouette widths as a proxy for misclassifications (tot_negative), the total number of signals in that cluster out of the total 6,273 (tot_sig), the mean silhouette width of that cluster (mean_sil), the median silhouette width for that cluster (med_sil), the s.d. of silhouette widths (sd_sil), the proportion of all signals with negative silhouette width in that cluster (tot_neg_normed) and which clustering the cluster belongs to (SOM_version).
Supplementary Data 5
Table showing the proportion of diel transcripts in each cluster and absolute number of detected KEGG orthologues for each taxon studied with diel signals. The columns refer to the taxon/molecule type (Taxon_or_Molecule_Type), the size fraction transcripts came from (Size_Fraction), the total number of diel signals belonging to that taxon (Total_Diel), then the total number and proportion of the total of signals for each taxon across the four SOM clusters.
Supplementary Data 6
Details for every diel signal from SOM clustering, including the signal, its cluster, the silhouette width for that signal (Methods) and the nearest neighbouring cluster for that signal. Files are divided by data source. The columns provide the cluster assignment (home_cluster), the identity of the transcript/lipid/metabolite (signal), the silhouette width of that signal (silhouette_width) and the neighbouring cluster as ascertained by the silhouette width calculation (neighbor_cluster).
Supplementary Data 7
NMDS ordination results including mean peak rank time calculation. In this calculation, peak rank time works as follows: a peak rank time of 1 indicates a 22:00 peak, 2 indicates 2:00, 3 indicates 6:00, 4 indicates 10:00, 5 indicates 14:00 and 6 indicates 18:00. The table includes coordinates from the initial NMDS ordination as well as coordinates rotated by π/16 to align midnight peak time with the top center of the plot. The columns are as follows: the original NMDS x coordinate of the signal (x); the original NMDS y coordinate of the signal (y); the full name of the signal (full_ids); the taxon of all transcripts with molecule names repeated for molecules (taxa); the KEGG orthologue number of all transcripts with ‘metabolite’ designating metabolites and ‘lipid’ designating lipids (KEGG orthologues); the calculated peak rank time of the signal as described above (time_rank); whether the signal is a lipid/metabolite/transcript (big_class); the labelling scheme used for Fig. 4 (new_tax); the KEGG pathway assignment for the transcripts (path); a column distinguishing molecules, bacterial heterotrophs, bacterial photoautotrophs and eukaryotes (tax_group); the assigned SOM cluster (cluster); the rotated NMDS x coordinate so that midnight peak time is at the top of the projection (x_rot); and the rotated NMDS y coordinate so that midnight peak time is at the top of the projection (y_rot).
Supplementary Data 8
Summary of mean peak rank time difference analysis. The columns include each KEGG orthologue that was included in the synchronous/asynchronous analysis (KEGG orthologue), the number of taxa that had diel transcription of that KEGG orthologue (n_tax), the average difference in peak times between each taxa with diel expression of that KEGG orthologue (diffs), the associated P value of the permutation test (pval) and the decision on whether the expression was significantly synchronous (reject) or not (fail) (rejects).
Rights and permissions
About this article
Cite this article
Muratore, D., Boysen, A.K., Harke, M.J. et al. Complex marine microbial communities partition metabolism of scarce resources over the diel cycle. Nat Ecol Evol 6, 218–229 (2022). https://doi.org/10.1038/s41559-021-01606-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-021-01606-w
This article is cited by
-
Disentangling top-down drivers of mortality underlying diel population dynamics of Prochlorococcus in the North Pacific Subtropical Gyre
Nature Communications (2024)
-
Gene expression dynamics of natural assemblages of heterotrophic flagellates during bacterivory
Microbiome (2023)
-
Marine phytoplankton downregulate core photosynthesis and carbon storage genes upon rapid mixed layer shallowing
The ISME Journal (2023)
-
Trait biases in microbial reference genomes
Scientific Data (2023)
-
Tracking nitrogen allocation to proteome biosynthesis in a marine microbial community
Nature Microbiology (2023)