Abstract
Across diverse taxa, selfing species have evolved independently from outcrossing species thousands of times. The transition from outcrossing to selfing decreases the effective population size, effective recombination rate and heterozygosity within a species. These changes lead to a reduction in genetic diversity, and therefore adaptive potential, by intensifying the effects of random genetic drift and linked selection. Within the nematode genus Caenorhabditis, selfing has evolved at least three times, and all three species, including the model organism Caenorhabditis elegans, show substantially reduced genetic diversity relative to outcrossing species. Selfing and outcrossing Caenorhabditis species are often found in the same niches, but we still do not know how selfing species with limited genetic diversity can adapt to these environments. Here, we examine the whole-genome sequences from 609 wild C. elegans strains isolated worldwide and show that genetic variation is concentrated in punctuated hyper-divergent regions that cover 20% of the C. elegans reference genome. These regions are enriched in environmental response genes that mediate sensory perception, pathogen response and xenobiotic stress response. Population genomic evidence suggests that genetic diversity in these regions has been maintained by long-term balancing selection. Using long-read genome assemblies for 15 wild strains, we show that hyper-divergent haplotypes contain unique sets of genes and show levels of divergence comparable to levels found between Caenorhabditis species that diverged millions of years ago. These results provide an example of how species can avoid the evolutionary dead end associated with selfing.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The raw short-read sequencing reads for the strains used in this project are available from the NCBI Sequence Read Archive (project PRJNA549503). The raw PacBio long-read data, along with the de novo assemblies and gene predictions, are available from the NCBI Sequence Read Archive (project PRJNA692613). Strain information and short-read genomic variation data are available from the CeNDR (www.elegansvariation.org)68.
Code availability
All datasets and code for generating the figures and tables are available from GitHub (https://github.com/AndersenLab/Ce-328pop-div).
References
Barrett, S. C. H. The evolution of plant sexual diversity. Nat. Rev. Genet. 3, 274–284 (2002).
Cutter, A. D. Reproductive transitions in plants and animals: selfing syndrome, sexual selection and speciation. New Phytol. 224, 1080–1094 (2019).
Pollak, E. On the theory of partially inbreeding finite populations. I. Partial selfing. Genetics 117, 353–360 (1987).
Kaplan, N. L., Hudson, R. R. & Langley, C. H. The ‘hitchhiking effect’ revisited. Genetics 123, 887–899 (1989).
Charlesworth, D. & Charlesworth, B. Quantitative genetics in plants: the effect of the breeding system on genetic variability. Evolution 49, 911–920 (1995).
Baker, H. G. Self-compatibility and establishment after ‘long-distance’ dispersal. Evolution 9, 347–349 (1955).
Baker, H. G. Support for Baker’s law—as a rule. Evolution 21, 853–856 (1967).
Charlesworth, D. & Wright, S. I. Breeding systems and genome evolution. Curr. Opin. Genet. Dev. 11, 685–690 (2001).
Stebbins, G. L. Self fertilization and population variability in the higher plants. Am. Nat. 91, 337–354 (1957).
Andersen, E. C. et al. Chromosome-scale selective sweeps shape Caenorhabditis elegans genomic diversity. Nat. Genet. 44, 285–290 (2012).
Cutter, A. D., Baird, S. E. & Charlesworth, D. High nucleotide polymorphism and rapid decay of linkage disequilibrium in wild populations of Caenorhabditis remanei. Genetics 174, 901–913 (2006).
Dey, A., Chan, C. K. W., Thomas, C. G. & Cutter, A. D. Molecular hyperdiversity defines populations of the nematode Caenorhabditis brenneri. Proc. Natl Acad. Sci. USA 110, 11056–11060 (2013).
Kiontke, K. et al. Caenorhabditis phylogeny predicts convergence of hermaphroditism and extensive intron loss. Proc. Natl Acad. Sci. USA 101, 9003–9008 (2004).
Sivasundar, A. & Hey, J. Population genetics of Caenorhabditis elegans: the paradox of low polymorphism in a widespread species. Genetics 163, 147–157 (2003).
Barrière, A. & Félix, M.-A. High local genetic diversity and low outcrossing rate in Caenorhabditis elegans natural populations. Curr. Biol. 15, 1176–1184 (2005).
Félix, M.-A. & Duveau, F. Population dynamics and habitat sharing of natural populations of Caenorhabditis elegans and C. briggsae. BMC Biol. 10, 59 (2012).
Schulenburg, H. & Félix, M.-A. The natural biotic environment of Caenorhabditis elegans. Genetics 206, 55–86 (2017).
Crombie, T. A. et al. Deep sampling of Hawaiian Caenorhabditis elegans reveals high genetic diversity and admixture with global populations. eLife 8, e50465 (2019).
Andrés, A. M. et al. Targets of balancing selection in the human genome. Mol. Biol. Evol. 26, 2755–2764 (2009).
Amambua-Ngwa, A. et al. Population genomic scan for candidate signatures of balancing selection to guide antigen characterization in malaria parasites. PLoS Genet. 8, e1002992 (2012).
Siewert, K. M. & Voight, B. F. Detecting long-term balancing selection using allele frequency correlation. Mol. Biol. Evol. 34, 2996–3005 (2017).
Wu, Q. et al. Long-term balancing selection contributes to adaptation in Arabidopsis and its relatives. Genome Biol. 18, 217 (2017).
Koenig, D. et al. Long-term balancing selection drives evolution of immunity genes in Capsella. eLife 8, e43606 (2019).
Langley, C. H. et al. Genomic variation in natural populations of Drosophila melanogaster. Genetics 192, 533–598 (2012).
Leffler, E. M. et al. Multiple instances of ancient balancing selection shared between humans and chimpanzees. Science 339, 1578–1582 (2013).
Charlesworth, D. Balancing selection and its effects on sequences in nearby genome regions. PLoS Genet. 2, e64 (2006).
Nordborg, M., Charlesworth, B. & Charlesworth, D. Increased levels of polymorphism surrounding selectively maintained sites in highly selling species. Proc. R. Soc. Lond. Ser. B Biol. Sci. 263, 1033–1039 (1996).
Wiuf, C., Zhao, K., Innan, H. & Nordborg, M. The probability and chromosomal extent of trans-specific polymorphism. Genetics 168, 2363–2372 (2004).
Seidel, H. S., Rockman, M. V. & Kruglyak, L. Widespread genetic incompatibility in C. elegans maintained by balancing selection. Science 319, 589–594 (2008).
Greene, J. S. et al. Balancing selection shapes density-dependent foraging behaviour. Nature 539, 254–258 (2016).
Van Sluijs, L. et al. Balancing selection shapes the intracellular pathogen response in natural Caenorhabditis elegans populations. Preprint at bioRxiv https://doi.org/10.1101/579151 (2019).
Thompson, O. A. et al. Remarkably divergent regions punctuate the genome assembly of the Caenorhabditis elegans Hawaiian strain CB4856. Genetics 200, 975–989 (2015).
Kim, C. et al. Long-read sequencing reveals intra-species tolerance of substantial structural variations and new subtelomere formation in C. elegans. Genome Res. 29, 1023–1035 (2019).
Richaud, A., Zhang, G., Lee, D., Lee, J. & Félix, M.-A. The local coexistence pattern of selfing genotypes in Caenorhabditis elegans natural metapopulations. Genetics 208, 807–821 (2018).
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Rockman, M. V. & Kruglyak, L. Recombinational landscape and population genomics of Caenorhabditis elegans. PLoS Genet. 5, e1000419 (2009).
Rockman, M. V., Skrovanek, S. S. & Kruglyak, L. Selection at linked sites shapes heritable phenotypic variation in C. elegans. Science 330, 372–376 (2010).
Cutter, A. D. & Payseur, B. A. Genomic signatures of selection at linked sites: unifying the disparity among species. Nat. Rev. Genet. 14, 262–274 (2013).
Gimond, C. et al. Outbreeding depression with low genetic variation in selfing Caenorhabditis nematodes. Evolution 67, 3087–3101 (2013).
Cutter, A. D., Morran, L. T. & Phillips, P. C. Males, outcrossing, and sexual selection in Caenorhabditis nematodes. Genetics 213, 27–57 (2019).
Barrett, R. D. H. & Schluter, D. Adaptation from standing genetic variation. Trends Ecol. Evol. 23, 38–44 (2008).
Schulenburg, H., Hoeppner, M. P., Weiner, J. 3rd & Bornberg-Bauer, E. Specificity of the innate immune system and diversity of C-type lectin domain (CTLD) proteins in the nematode Caenorhabditis elegans. Immunobiology 213, 237–250 (2008).
Reddy, K. C. et al. An intracellular pathogen response pathway promotes proteostasis in C. elegans. Curr. Biol. 27, 3544–3553.e5 (2017).
Bakowski, M. A. et al. Ubiquitin-mediated response to microsporidia and virus infection in C. elegans. PLoS Pathog. 10, e1004200 (2014).
Chang, H. C., Paek, J. & Kim, D. H. Natural polymorphisms in C. elegans HECW-1 E3 ligase affect pathogen avoidance behaviour. Nature 480, 525–529 (2011).
Troemel, E. R., Félix, M.-A., Whiteman, N. K., Barrière, A. & Ausubel, F. M. Microsporidia are natural intracellular parasites of the nematode Caenorhabditis elegans. PLoS Biol. 6, 2736–2752 (2008).
Félix, M.-A. et al. Natural and experimental infection of Caenorhabditis nematodes by novel viruses related to nodaviruses. PLoS Biol. 9, e1000586 (2011).
Chen, K., Franz, C. J., Jiang, H., Jiang, Y. & Wang, D. An evolutionarily conserved transcriptional response to viral infection in Caenorhabditis nematodes. BMC Genom. 18, 303 (2017).
Balla, K. M., Andersen, E. C., Kruglyak, L. & Troemel, E. R. A wild C. elegans strain has enhanced epithelial immunity to a natural microsporidian parasite. PLoS Pathog. 11, e1004583 (2015).
Ashe, A. et al. A deletion polymorphism in the Caenorhabditis elegans RIG-I homolog disables viral RNA dicing and antiviral immunity. eLife 2, e00994 (2013).
Martin, N., Singh, J. & Aballay, A. Natural genetic variation in the Caenorhabditis elegans response to Pseudomonas aeruginosa. G3 7, 1137–1147 (2017).
Thomas, C. G. et al. Full-genome evolutionary histories of selfing, splitting, and selection in Caenorhabditis. Genome Res. 25, 667–678 (2015).
Kiontke, K. C. et al. A phylogeny and molecular barcodes for Caenorhabditis, with numerous new species from rotting fruits. BMC Evol. Biol. 11, 339 (2011).
Busch, J. W. & Delph, L. F. Evolution: selfing takes species down Stebbins’s blind alley. Curr. Biol. 27, R61–R63 (2017).
Ferrari, C. et al. Ephemeral-habitat colonization and neotropical species richness of Caenorhabditis nematodes. BMC Ecol. 17, 43 (2017).
Greene, J. S., Dobosiewicz, M., Butcher, R. A., McGrath, P. T. & Bargmann, C. I.Regulatory changes in two chemoreceptor genes contribute to a Caenorhabditis elegans QTL for foraging behavior. eLife 5, e21454 (2016).
Lee, D. et al. Selection and gene flow shape niche-associated variation in pheromone response. Nat. Ecol. Evol. 3, 1455–1463 (2019).
Webster, A. K. et al. Population selection and sequencing of Caenorhabditis elegans wild isolates identifies a region on chromosome III affecting starvation resistance. G3 9, 3477–3488 (2019).
Ghosh, R., Andersen, E. C., Shapiro, J. A., Gerke, J. P. & Kruglyak, L. Natural variation in a chloride channel subunit confers avermectin resistance in C. elegans. Science 335, 574–578 (2012).
Ben-David, E., Burga, A. & Kruglyak, L. A maternal-effect selfish genetic element in Caenorhabditis elegans. Science 356, 1051–1055 (2017).
Liu, Y. et al. Pan-genome of wild and cultivated soybeans. Cell 182, 162–176.e13 (2020).
Cutter, A. D., Wasmuth, J. D. & Washington, N. L. Patterns of molecular evolution in Caenorhabditis preclude ancient origins of selfing. Genetics 178, 2093–2104 (2008).
Brandvain, Y., Slotte, T., Hazzouri, K. M., Wright, S. I. & Coop, G. Genomic identification of founding haplotypes reveals the history of the selfing species Capsella rubella. PLoS Genet. 9, e1003754 (2013).
Todesco, M. et al. Massive haplotypes underlie ecotypic differentiation in sunflowers. Nature 584, 602–607 (2020).
Burgarella, C. et al. Adaptive introgression: an untapped evolutionary mechanism for crop adaptation. Front. Plant Sci. 10, 4 (2019).
Kanzaki, N. et al. Biology and genome of a newly discovered sibling species of Caenorhabditis elegans. Nat. Commun. 9, 3216 (2018).
Andersen, E. C., Bloom, J. S., Gerke, J. P. & Kruglyak, L. A variant in the neuropeptide receptor npr-1 is a major determinant of Caenorhabditis elegans growth and physiology. PLoS Genet. 10, e1004156 (2014).
Cook, D. E., Zdraljevic, S., Roberts, J. P. & Andersen, E. C. CeNDR, the Caenorhabditis elegans Natural Diversity Resource. Nucleic Acids Res. 45, D650–D657 (2017).
Cook, D. E. et al. The genetic basis of natural variation in Caenorhabditis elegans telomere length. Genetics 204, 371–383 (2016).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Lee, R. Y. N. et al. WormBase 2017: molting into a new stage. Nucleic Acids Res. 46, D869–D874 (2018).
Poplin, R. et al. Scaling accurate genetic variant discovery to tens of thousands of samples. Preprint at bioRxiv https://doi.org/10.1101/201178 (2018).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 6, 80–92 (2012).
Ortiz, E. M. vcf2phylip v2.0: convert a VCF matrix into several matrix formats for phylogenetic analysis. GitHub https://github.com/edgardomortiz/vcf2phylip (2019).
Schliep, K. P. phangorn: phylogenetic analysis in R. Bioinformatics 27, 592–593 (2011).
Yu, G., Smith, D. K., Zhu, H., Guan, Y. & Lam, T. T.-Y. ggtree: an R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 8, 28–36 (2017).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Browning, B. L. & Browning, S. R. Detecting identity by descent and estimating genotype error rates in sequence data. Am. J. Hum. Genet. 93, 840–851 (2013).
Miles, A., Ralph, P., Rae, S. & Pisupati, R. cggh/scikit-allel: v1.2.1. Zenodo https://doi.org/10.5281/zenodo.3238280 (2019).
Siewert, K. M. & Voight, B. F.BetaScan2: standardized statistics to detect balancing selection utilizing substitution data. Genome Biol. Evol. 12, 3873–3877 (2020).
Siewert, K. BetaScan GitHub https://github.com/ksiewert/BetaScan (2017).
Zhang, C., Dong, S.-S., Xu, J.-Y., He, W.-M. & Yang, T.-L. PopLDdecay: a fast and effective tool for linkage disequilibrium decay analysis based on variant call format files. Bioinformatics 35, 1786–1788 (2019).
Ruan, J. & Li, H. Fast and accurate long-read assembly with wtdbg2. Nat. Methods 17, 155–158 (2020).
Kolmogorov, M., Yuan, J., Lin, Y. & Pevzner, P. A. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 37, 540–546 (2019).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963 (2014).
Laetsch, D. R. & Blaxter, M. L. BlobTools: interrogation of genome assemblies. F1000Res. 6, 1287 (2017).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinform. 10, 421 (2009).
Pundir, S., Martin, M. J. & O’Donovan, C. in Protein Bioinformatics: From Protein Modifications and Networks to Proteomics (eds Wu, C. H. et al.) 41–55 (Springer, 2017).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2015).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Pedersen, B. S. & Quinlan, A. R. Mosdepth: quick coverage calculation for genomes and exomes. Bioinformatics 34, 867–868 (2018).
C. elegans Sequencing Consortium. Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282, 2012–2018 (1998).
Delcher, A. L., Salzberg, S. L. & Phillippy, A. M. Using MUMmer to identify similar regions in large sequence sets. Curr. Protoc. Bioinform. 10, 10.3 (2003).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Holdorf, A. D. et al. WormCat: an online tool for annotation and visualization of Caenorhabditis elegans genome-scale data. Genetics 214, 279–294 (2019).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Carlson, M. org.Ce.eg.db: Genome wide annotation for Worm. R package version 3.8.2 https://bioconductor.org/packages/release/data/annotation/html/org.Ce.eg.db.html (2019).
Bao, W., Kojima, K. K. & Kohany, O. Repbase Update, a database of repetitive elements in eukaryotic genomes. Mob. DNA 6, 11 (2015).
Stanke, M., Diekhans, M., Baertsch, R. & Haussler, D. Using native and syntenically mapped cDNA alignments to improve de novo gene finding. Bioinformatics 24, 637–644 (2008).
Emms, D. M. & Kelly, S. OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biol. 16, 157 (2015).
Slater, G. S. C. & Birney, E. Automated generation of heuristics for biological sequence comparison. BMC Bioinform. 6, 31 (2005).
Finn, R. D. et al. The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 44, D279–D285 (2016).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2016).
Bradley, R. K. et al. Fast statistical alignment. PLoS Comput. Biol. 5, e1000392 (2009).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., von Haeseler, A. & Jermiin, L. S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Zheng, X. et al. A high-performance computing toolset for relatedness and principal component analysis of SNP data. Bioinformatics 28, 3326–3328 (2012).
Stein, L. D. et al. The genome sequence of Caenorhabditis briggsae: a platform for comparative genomics. PLoS Biol. 1, E45 (2003).
Yin, D. et al. Rapid genome shrinkage in a self-fertile nematode reveals sperm competition proteins. Science 359, 55–61 (2018).
Stevens, L. et al. The genome of Caenorhabditis bovis. Curr. Biol. 30, 1023–1031.e4 (2020).
Acknowledgements
We thank members of the Andersen laboratory for providing comments on this manuscript. We especially thank M. Ailion, J. David, R. Luallen, N. Pujol and citizen scientists for contributing wild C. elegans strains to CeNDR. We also thank the Duke University School of Medicine for use of the Sequencing and Genomic Technologies Shared Resource, which provided Pacific Biosciences long-read sequencing. This work was funded by an NSF CAREER award (1751035) and a Human Frontier Science Program Award (RGP0001/2019) (to E.C.A.). This work was also funded by National Institutes of Health (NIH) grant ES029930 (to E.C.A., M.V.R. and L.R.B.). S.Z. received funding from The Cellular and Molecular Basis of Disease training programme (T32GM008061) and the Rappaport Award for Research Excellence through the IBiS graduate programme. A.K.W. is supported by the National Science Foundation Graduate Research Fellowship. Long-read sequencing of three isolates was funded by the NIH (R01 GM117408 to L.R.B.) and a T32 training grant for the University Program in Genetics and Genomics (GM007754). M.V.R. is supported by NIH grant GM121828. M.G.S. was supported by an NWO Domain Applied and Engineering Sciences Veni grant (17282).
Author information
Authors and Affiliations
Contributions
D.L., S.Z. and E.C.A. conceived of and designed the study. D.L., S.Z., L.S. and E.C.A. analysed the data and wrote the manuscript. Y.W., R.E.T. and D.E.C. performed whole-genome sequencing and isotype characterization for 609 wild C. elegans strains. R.E.T. performed long-read sequencing for 11 C. elegans wild isolates. R.C., A.K.W. and L.R.B. performed long-read sequencing for three C. elegans wild isolates. M.G.S., C.B., M.V.R. and M.-A.F. contributed wild isolates to the C. elegans strain collection. M.G.S., C.B., M.V.R., M.-A.F. and T.A.C. edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Ecology & Evolution thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Chromosome-scale selective sweeps across wild C. elegans isotypes.
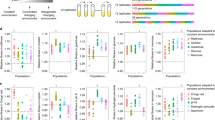
a, The genome-wide distribution of the most frequent haplotype (red) among 324 wild isotypes with known geographic origin is shown. Grey genomic regions represent other haplotypes, and white represents unclassified haplotypes. Each row is one of the 324 isotypes, grouped by the geographic origin. The genomic position in Mb is plotted on the x-axis, and each tick mark represents 5 Mb of the chromosome. b, Beeswarm plots of the proportion of the most frequent haplotype for each chromosome from (a) for 324 isotypes with known geographic origins are shown. Wild isotypes are grouped by geographic origin. Each point corresponds to one of the 324 isotypes, and geographic origins are shown on the y-axis.
Extended Data Fig. 2 Patterns of molecular diversity across the C. elegans genome.
The chromosomal patterns a, Watterson’s theta (θ) and b, nucleotide diversity (pi) for non-overlapping 1 kb windows are shown. Each dot corresponds to the calculated value for a particular window. The genomic position in Mb is plotted on the x-axis. Diversity statistic values are shown on the y-axis. Smoothed lines (blue) are LOESS fits. c, Tukey box plots of genetic diversity statistics from (a) are shown with outlier data points plotted. Genetic diversity statistics for each sliding window are grouped by the chromosomal region defined previously36. Genetic diversity statistic values are shown on the y-axis. The horizontal line in the middle of the box is the median, and the box denotes the 25th to 75th quantiles of the data. The vertical line represents the 1.5x interquartile range.
Extended Data Fig. 3 Optimization of parameters for the characterization of hyper-divergent regions.
a,b, The total detected hyper-divergent regions in Mb (x-axis) and the percent overlap of long-read and short-read hyper-divergent classification (y-axis) are shown (Methods). Each point corresponds to one of the combination of threshold parameters for the variant count and coverage fraction of 1 kb bin to be classified as hyper-divergent. Each point is coloured by the variant count threshold (a) or the coverage fraction threshold (b). c, The relationship between the total size of hyper-divergent regions detected by the optimized short-read or long-read based approach is shown. Each point corresponds to one of the 15 long-read sequenced isotypes. Total sizes of hyper-divergent regions detected by the short-read based approach are shown on the x-axis, and total sizes of hyper-divergent regions detected by the long-read based approach are shown on the y-axis. d, The overlap between hyper-divergent regions defined by the optimized short-read based approach and long-read based approach is shown. Each point corresponds to one of the 15 long-read sequenced isotypes. Total sizes of hyper-divergent regions detected by either short-read or long-read based approach are shown on the x-axis, and the percentages of hyper-divergent regions detected by both approaches are shown on the y-axis.
Extended Data Fig. 4 Summary statistics for hyper-divergent regions across six chromosomes.
a, Bar plots for the comparisons of variant (SNV/indel) density (top) and coverage fraction (bottom) between hyper-divergent regions (red) and the rest of the regions (blue) in each chromosomal region are shown. Note that no hyper-divergent region was found on the tips of chromosome I. b, Fold differences between hyper-divergent regions and the rest of the regions from (a) are shown.
Extended Data Fig. 5 Genomic signatures of balancing selection in non-divergent regions and hyper-divergent regions.
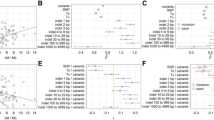
Tukey box plots of Tajima’s D (a) and standardized beta (b) are shown. Genomic bins (1 kb) (a) or variants (b) are grouped and coloured by their classification: (1) non-divergent bins (yellow), (2) hyper-divergent bins with high variant density (≥ 16 SNVs/indels, red), (3) hyper-divergent bins with low read depth (< 35%, blue). Hyper-divergent bins are grouped by their species-wide frequencies: rare (<1%), intermediate (≥ 1% and < 5%), or common (≥ 5%). The horizontal line in the middle of the box is the median, and the box denotes the 25th to 75th quantiles of the data. The vertical line represents the 1.5x interquartile range.
Extended Data Fig. 6 Gene ontology (GO) enrichment for hyper-divergent regions.
Gene ontology (GO) enrichment for the biological process category (a) and the molecular function category (b) for non-divergent chromosomal arms (square) and hyper-divergent regions (circle) are shown. Significantly enriched GO terms in control regions or hyper-divergent regions or both are shown on the y-axis. Bonferroni-corrected significance values for GO enrichment are shown on the x-axis. Sizes of squares and circles correspond to the fold enrichment of the annotation, and colours of square and circle correspond to the gene counts of the annotation. The blue line shows the Bonferroni-corrected significance threshold (corrected p-value = 0.05). Note, we did not detect any GO-term enrichment of genes in non-divergent chromosomal arms for the biological process category.
Extended Data Fig. 7 Species-wide SNP-based relatedness of divergent regions is in agreement with long-read sequencing results.
The inferred for the C. elegans species-wide relatedness for the hyper-divergent regions that span (a) II:3,667,179-3,701,405, (b) I:2,318,291-2,381,851, and (c) V:20,193,463-20,267,244 are shown. The x-axis represents the dissimilarity of the fraction of identity-by-state in the region. For a-c, the isotype names are coloured to match the haplotypes defined by long-read sequence data in Fig. 5 and Extended Data Figs. 8, 9, respectively. The branch colours correspond to the species-wide genetic groups identified by PCA in Fig. 1c.
Extended Data Fig. 8 Two hyper-divergent haplotypes at the peel-1 zeel-1 incompatibility locus.
a, The protein-coding gene contents of the two hyper-divergent haplotypes at the peel-1 zeel-1 incompatibility locus on the left arm of chromosome I (I:2,318,291-2,381,851 of the N2 reference genome). The tree was inferred using SNVs and coloured by inferred haplotypes. For each distinct haplotype, we chose a single isotype as a haplotype representative (orange haplotype: N2, blue haplotype: CB4856) and predicted protein-coding genes using both protein-based alignments and ab initio approaches. Protein-coding genes are shown as boxes; those genes that are conserved in all haplotypes are coloured based on their haplotype, and those genes that are not are coloured light grey. Dark grey boxes behind genes indicate coordinates of divergent regions. Genes with locus names in N2 are highlighted. b, Heatmaps showing amino acid identity for alleles of four genes (mcm-4, srbc-64, ugt-31, and sydn-1). The percentage identity was calculated using alignments of protein sequences from all 16 isotypes. Heatmaps are ordered by the SNV tree shown in (a). c, Maximum-likelihood gene trees of four genes (mcm-4, srbc-64, ugt-31, and sydn-1) inferred using amino acid alignments. Trees are plotted on the same scale (scale shown; scale is in substitutions per site). Strain names are coloured by their haplotype.
Extended Data Fig. 9 Hyper-divergent haplotypes at a region on the right arm of chromosome V.
a, The protein-coding gene contents of the seven hyper-divergent haplotypes at a region on the right arm of chromosome V (V:20,193,463-20,267,244 of the N2 reference genome). The tree was inferred using SNVs and coloured by inferred haplotypes. For each distinct haplotype, we chose a single isotype as a haplotype representative (orange haplotype: N2, light blue haplotype: JU2526, red haplotype: EG4725, pink haplotype: ECA36, green haplotype: DL238, dark blue haplotype: QX1794, purple haplotype: NIC526) and predicted protein-coding genes using both protein-based alignments and ab initio approaches. JU2526 shares the reference haplotype at fbxa-113 and fbxb-59 (six hyper-divergent haplotypes at these loci) but is divergent at Y113G7B.15 (seven hyper-divergent haplotypes at this locus). Protein-coding genes are shown as boxes; those genes that are conserved in all haplotypes are coloured based on their haplotypes, and those genes that are not are coloured light grey. Dark grey boxes behind genes indicate coordinates of divergent regions. Genes with locus names in N2 are highlighted. Of the 25 genes that are not conserved in all haplotypes (light grey boxes), ten are alleles of the three reference haplotype (N2) loci coloured in light grey. The remaining 15 do not have a clear one-to-one relationship with a gene in the reference haplotype. Seven of these 15 have homology to F54E12.2 (present in the reference haplotype) and are likely the product of duplication and diversification. Six have homology to either M04C3.1, F19B2.5, or F54E12.2, all of which are genes with SNF2 family N-terminal domains and which exist elsewhere in the N2 reference genome. Of the remaining two genes, one has homology to Y113G7B.15, which is present in the reference haplotype, and the other has homology to W09C3.8, a gene on chromosome I in the reference genome. Functional annotations of all unconserved loci (including BLAST hits and Pfam domains identified by InterProScan) can be found in Supplementary Data 4. b, Heatmaps show amino acid identity for between alleles of five genes (srh-217, fbxb-113, fbxb-59, Y113G7B.15, and mdt-17). The percentage identity was calculated using alignments of proteins sequences from all 16 isotypes. Heatmaps are ordered by the SNV tree shown in (a). c, Maximum-likelihood gene trees of five genes (srh-217, fbxb-113, fbxb-59, Y113G7B.15, and mdt-17) inferred using amino acid alignments. Trees are plotted on the same scale (scale shown; scale is in substitutions per site). Strain names are coloured by their haplotype.
Extended Data Fig. 10 Hyper-divergent regions in C. briggsae.
The genome-wide distribution of hyper-divergent regions across 35 non-reference wild C. briggsae strains is shown. In the top panel, each row is one of the 35 strains, grouped by previously defined clades (tropical or others) ordered by the total amount of genome covered by hyper-divergent regions (black). In the bottom panel, brown bars indicate genomic positions in which more than 10% of strains are classified as hyper-divergent at the locus. The genomic position in Mb is plotted on the x-axis, and each tick represents 5 Mb of the chromosome.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8 and Tables 1–6.
Supplementary Tables
Supplementary Tables 1–6.
Supplementary Data
Supplementary Data 1–4.
Rights and permissions
About this article
Cite this article
Lee, D., Zdraljevic, S., Stevens, L. et al. Balancing selection maintains hyper-divergent haplotypes in Caenorhabditis elegans. Nat Ecol Evol 5, 794–807 (2021). https://doi.org/10.1038/s41559-021-01435-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-021-01435-x
This article is cited by
-
The genome and transcriptome of the snail Biomphalaria sudanica s.l.: immune gene diversification and highly polymorphic genomic regions in an important African vector of Schistosoma mansoni
BMC Genomics (2024)
-
Novel and improved Caenorhabditis briggsae gene models generated by community curation
BMC Genomics (2023)
-
The parasitic nematode Strongyloides ratti exists predominantly as populations of long-lived asexual lineages
Nature Communications (2023)
-
Higher-order epistasis shapes natural variation in germ stem cell niche activity
Nature Communications (2023)
-
Ancient diversity in host-parasite interaction genes in a model parasitic nematode
Nature Communications (2023)