Abstract
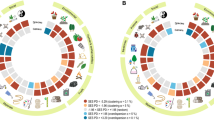
The divergent nature of evolution suggests that securing the human benefits that are directly provided by biodiversity may require counting on disparate lineages of the Tree of Life. However, quantitative evidence supporting this claim is still tenuous. Here, we draw on a global review of plant-use records demonstrating that maximum levels of phylogenetic diversity capture significantly greater numbers of plant-use records than random selection of taxa. Our study establishes an empirical foundation that links evolutionary history to human wellbeing, and it will serve as a discussion baseline to promote better-grounded accounts of the services that are directly provided by biodiversity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The data that support the findings of this study are available at https://doi.org/10.6084/m9.figshare.13625546.v1.
Code availability
All the code used in this research is available as functions that were either implemented in published R packages or provided as supplementary material in a previous open-access study.
References
Faith, D. P. et al. Evosystem services: an evolutionary perspective on the links between biodiversity and human well-being. Curr. Opin. Environ. Sust. 2, 66–74 (2010).
Cámara-Leret, R. et al. Fundamental species traits explain provisioning services of tropical American palms. Nat. Plants 3, 16220 (2017).
Oka, C., Aiba, M. & Nakashizuka, T. Phylogenetic clustering in beneficial attributes of tree species directly linked to provisioning, regulating and cultural ecosystem services. Ecol. Indic. 96, 477–495 (2019).
Faith, D. P. Conservation evaluation and phylogenetic diversity. Biol. Conserv. 61, 1–10 (1992).
Vane-Wright, R. I., Humphries, C. J. & Williams, P. H. What to protect?—Systematics and the agony of choice. Biol. Conserv. 55, 235–254 (1991).
Crozier, R. H. Genetic diversity and the agony of choice. Biol. Conserv. 61, 11–15 (1992).
Tucker, C. M. et al. Assessing the utility of conserving evolutionary history. Biol. Rev. 94, 1740–1760 (2019).
Owen, N. R., Gumbs, R., Gray, C. L. & Faith, D. P. Global conservation of phylogenetic diversity captures more than just functional diversity. Nat. Commun. 10, 859 (2019).
Mazel, F. et al. Prioritizing phylogenetic diversity captures functional diversity unreliably. Nat. Commun. 9, 2888 (2018).
Mazel, F. et al. Reply to: ‘Global conservation of phylogenetic diversity captures more than just functional diversity’. Nat. Commun. 10, 858 (2019).
Forest, F. et al. Preserving the evolutionary potential of floras in biodiversity hotspots. Nature 445, 757–760 (2007).
Cook, F. E. M. Economic Botany Data Collection Standard (International Working Group on Taxonomic Databases for Plant Sciences, Royal Botanic Gardens, UK, 1995).
Smith, S. A. & Brown, J. W. Constructing a broadly inclusive seed plant phylogeny. Am. J. Bot. 105, 302–314 (2018).
Jin, Y. & Qian, H. V. PhyloMaker: an R package that can generate very large phylogenies for vascular plants. Ecography 42, 1353–1359 (2019).
Mabberley, D. J. Mabberley’s Plant-book: A Portable Dictionary of Plants, Their Classification and Uses 4th edn (Cambridge Univ. Press, 2017).
Cox, P. A. Will tribal knowledge survive the millennium? Science 287, 44–45 (2000).
Cámara-Leret, R., Paniagua-Zambrana, N., Balslev, H. & Macía, M. J. Ethnobotanical knowledge is vastly under-documented in northwestern South America. PLoS ONE 9, e85794 (2014).
Cámara-Leret, R. & Dennehy, Z. Information gaps in indigenous and local knowledge for science-policy assessments. Nat. Sustain. 2, 736–741 (2019).
Novotny, V. et al. Low host specificity of herbivorous insects in a tropical forest. Nature 416, 841–844 (2002).
Gilbert, G. S., Magarey, R., Suiter, K. & Webb, C. O. Evolutionary tools for phytosanitary risk analysis: phylogenetic signal as a predictor of host range of plant pests and pathogens. Evol. Appl. 5, 869–878 (2012).
Calatayud, J. et al. Geography and major host evolutionary transitions shape the resource use of plant parasites. Proc. Natl Acad. Sci. USA 113, 9840–9845 (2016).
Pecl, G. T. et al. Biodiversity redistribution under climate change: impacts on ecosystems and human well-being. Science 355, eai9214 (2017).
Lehmann, P. et al. Complex responses of global insect pests to climate warming. Front. Ecol. Environ. 18, 141–150 (2020).
de Lucena, R. F. P. et al. The ecological apparency hypothesis and the importance of useful plants in rural communities from Northeastern Brazil: an assessment based on use value. J. Environ. Manag. 96, 106–115 (2012).
Menendez-Baceta, G. et al. The importance of cultural factors in the distribution of medicinal plant knowledge: a case study in four Basque regions. J. Ethnopharmacol. 161, 116–127 (2015).
Webb, C. O., Ackerly, D. D., McPeek, M. A. & Donoghue, M. J. Phylogenies and community ecology. Annu. Rev. Ecol. Syst. 33, 475–505 (2002).
Global Information on Scoping for the Thematic Assessment of Sustainable Use of Wild Species (Intergovernmental Science-Policy Platform on Biodiversity and Ecosystem Services, 2018); https://ipbes.net/sustainable-use-wild-species-assessment
Karki, M., Senaratna Sellamuttu, S., Okayasu, S. & Suzuki, W. (eds) Regional Assessment Report on Biodiversity and Ecosystem Services for Asia and the Pacific (Secretariat of the IPBES, 2018).
Pardo-de-Santayana, M. & Macía, M. The benefits of traditional knowledge. Nature 518, 487–488 (2015).
Díaz, S. et al. Assessing nature’s contributions to people. Science 359, 270–272 (2018).
Antonelli, A. et al. State of the World’s Plants and Fungi 2020 (Royal Botanic Gardens, Kew, 2020).
Ulian, T. et al. Unlocking plant resources to support food security and promote sustainable agriculture. Plants People Planet 2, 421–445 (2020).
Plants of the World Online (Royal Botanic Gardens, Kew, 2021); http://www.plantsoftheworldonline.org/
Zanne, A. E. et al. Three keys to the radiation of angiosperms into freezing environments. Nature 506, 89–92 (2014).
The Plant List, version 1.1 (The Plant List, 2013); http://www.theplantlist.org/
Rangel, T. F. et al. Phylogenetic uncertainty revisited: implications for ecological analyses. Evolution 69, 1301–1312 (2015).
Federhen, S. The NCBI taxonomy database. Nucleic Acids Res. 40, D136–D143 (2012).
Hörandl, E. & Stuessy, T. F. Paraphyletic groups as natural units of biological classification. Taxon 59, 1641–1653 (2010).
Revell, L. J. phytools: an R package for phylogenetic comparative biology (and other things). Methods Ecol. Evol. 3, 217–223 (2012).
Rodrigues, A. S. L. & Gaston, K. J. Maximising phylogenetic diversity in the selection of networks of conservation areas. Biol. Conserv. 105, 103–111 (2002).
Bordewich, M., Rodrigo, A. G. & Semple, C. Selecting taxa to save or sequence: desirable criteria and a greedy solution. Syst. Biol. 57, 825–834 (2008).
Pielou, E. C. The measurement of diversity in different types of biological collections. J. Theor. Biol. 13, 131–144 (1966).
Kembel, S. W. Disentangling niche and neutral influences on community assembly: assessing the performance of community phylogenetic structure tests. Ecol. Lett. 12, 949–960 (2009).
R Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2020).
Kembel, S. W. et al. Picante: R tools for integrating phylogenies and ecology. Bioinformatics 26, 1463–1464 (2010).
Brummitt, R. K. World Geographical Scheme for Recording Plant Distributions 2nd edn (International Working Group on Taxonomic Databases for Plant Sciences, 2001).
Baselga, A. Partitioning the turnover and nestedness components of beta diversity. Glob. Ecol. Biogeogr. 19, 134–143 (2010).
Acknowledgements
We thank the Scientific Computation Center of Andalusia (CICA) for the computing services they provided and H. Lima for assistance in downloading plant distributional information from the web. This work was supported by the Regional Government of the Community of Madrid and the University of Alcalá through the project ‘Plant evolutionary history and human wellbeing in a changing world; assessing theoretical foundations using empirical evidence and new phylogenetic tools’, which was granted to R.M.-V. (CM/JIN/2019-005). R.M.-V. was supported by the TALENTO programme of the Regional Government of the Community of Madrid (2018-T2/AMB-10332). M.Á.R. was supported by the Ministry of Science and Innovation of Spain (grant CGL2017-86926-P).
Author information
Authors and Affiliations
Contributions
R.M.-V. conceived the ideas, led the assemblage of the plant-use dataset with the help of M.P.S. and D.J.M., conducted the analyses and led the writing. C.R. led the assemblage of the continental datasets. M.Á.R. helped to design the structure of the draft. All the authors read, edited and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Peer review information Nature Ecology & Evolution thanks Rainer Bussmann and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Relationship between the phylogenetic structure of plant-use categories and relative gains per category under the PDmax strategy.
The dotted lines represent the regression models between the phylogenetic structure of plant-use categories (SES scores of PD averaged across 100 phylogenetic hypotheses) and SES scores of the relative gains per category across different sample sizes (S = 20, 40, 60 and 80% of the total pool). All regressions were significant for a nominal alpha of 0.1%.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10 and Tables 1, 2 and 5.
Supplementary Tables 3 and 4
Supplementary Table 3. List of genera included in the study. Supplementary Table 4. Most derived consensus clades (MDCCs) for the phylogenetically uncertain taxa (PUTs) of the analysis.
Rights and permissions
About this article
Cite this article
Molina-Venegas, R., Rodríguez, M.Á., Pardo-de-Santayana, M. et al. Maximum levels of global phylogenetic diversity efficiently capture plant services for humankind. Nat Ecol Evol 5, 583–588 (2021). https://doi.org/10.1038/s41559-021-01414-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-021-01414-2
This article is cited by
-
Global conservation status of the jawed vertebrate Tree of Life
Nature Communications (2024)
-
Human activities and species biological traits drive the long-term persistence of old trees in human-dominated landscapes
Nature Plants (2023)
-
Global hotspots of traded phylogenetic and functional diversity
Nature (2023)
-
Spatially Well Structured Mangroves Fish Communities of the Persian Gulf; a Functional Perspective
Wetlands (2023)
-
The likely extinction of hundreds of palm species threatens their contributions to people and ecosystems
Nature Ecology & Evolution (2022)