Abstract
In the absence of antibiotics, it is essential that antibiotic resistance has a fitness cost for microorganisms if suspending antibiotics treatment is to be a useful strategy for reducing antibiotic resistance. However, the cost of antibiotic resistance within the complex ecosystem of the mammalian gut is not well understood. Here, using mice, we show that the same antibiotic resistance mutation can reduce fitness in one host, while being neutral or even increasing fitness in other hosts. Such antagonistic pleiotropy is shaped by the microbiota because resistance in germ-free mice is consistently costly across all hosts, and the host-specific effect on antibiotic resistance is reduced in hosts with similar microbiotas. Using an eco-evolutionary model of competition for resources, we identify a general mechanism that underlies between-host variation and predicts that the dynamics of compensatory evolution of resistant bacteria should be host specific, a prediction that was supported by experimental evolution in vivo. The microbiome of each human is close to unique, and our results suggest that the short-term cost of resistances and their long-term within-host evolution are also highly personalized, a finding that may contribute to the observed variable outcome of withdrawing antibiotics to reduce resistance levels.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The fitness and sequencing data that support the findings of this study are available from the Dryad Digital Repository (https://doi.org/10.5061/dryad.j0zpc869r).
Code availability
The code of the simulations to produce the data of Figs. 2 and 3 are available from the Dryad Digital Repository (https://doi.org/10.5061/dryad.j0zpc869r).
References
Antimicrobial Resistance: Global Report on Surveillance (WHO, 2014).
Gullberg, E. et al. Selection of resistant bacteria at very low antibiotic concentrations. PLoS Pathog. 7, e1002158 (2011).
MacLean, R. C. & Vogwill, T. Limits to compensatory adaptation and the persistence of antibiotic resistance in pathogenic bacteria. Evol. Med. Publ. Health 1, 4–12 (2015).
Bhullar, K. et al. Antibiotic resistance is prevalent in an isolated cave microbiome. PLoS ONE 7, e34953 (2012).
Forsberg, K. J. et al. The shared antibiotic resistome of soil bacteria and human pathogens. Science 337, 1107–1111 (2012).
Hu, Y. et al. Metagenome-wide analysis of antibiotic resistance genes in a large cohort of human gut microbiota. Nat. Commun. 4, 2151 (2013).
Andersson, D. I. & Hughes, D. Antibiotic resistance and its cost: is it possible to reverse resistance? Nat. Rev. Microbiol. 8, 260–271 (2010).
Durão, P., Balbontín, R. & Gordo, I. Evolutionary mechanisms shaping the maintenance of antibiotic resistance. Trends Microbiol. 26, 677–691 (2018).
Trindade, S. et al. Positive epistasis drives the acquisition of multidrug resistance. PLoS Genet 5, e1000578 (2009).
Miskinyte, M. & Gordo, I. Increased survival of antibiotic-resistant Escherichia coli inside macrophages. Antimicrob. Agents Chemother. 57, 189–195 (2013).
Durão, P., Gülereşi, D., Proença, J. & Gordo, I. Enhanced survival of rifampin- and streptomycin-resistant Escherichia coli inside macrophages. Antimicrob. Agents Chemother. 60, 4324–4332 (2016).
Reynolds, M. G. Compensatory evolution in rifampin-resistant Escherichia coli. Genetics 156, 1471–1481 (2000).
Enne, V. I., Bennett, P. M., Livermore, D. M. & Hall, L. M. C. Enhancement of host fitness by the sul2-coding plasmid p9123 in the absence of selective pressure. J. Antimicrob. Chemother. 53, 958–963 (2004).
Gagneux, S. et al. The competitive cost of antibiotic resistance in Mycobacterium tuberculosis. Science 312, 1944–1946 (2006).
Melnyk, A. H., Wong, A. & Kassen, R. The fitness costs of antibiotic resistance mutations. Evol. Appl. 8, 273–283 (2015).
Seppälä, H. et al. The effect of changes in the consumption of macrolide antibiotics on erythromycin resistance in group A streptococci in Finland. N. Engl. J. Med. 337, 441–446 (1997).
Enne, V. I., Livermore, D. M., Stephens, P. & Hall, L. M. Persistence of sulphonamide resistance in Escherichia coli in the UK despite national prescribing restriction. Lancet 357, 1325–1328 (2001).
Bean, D. C., Livermore, D. M., Papa, I. & Hall, L. M. C. Resistance among Escherichia coli to sulphonamides and other antimicrobials now little used in man. J. Antimicrob. Chemother. 56, 962–964 (2005).
Gottesman, B. S., Carmeli, Y., Shitrit, P. & Chowers, M. Impact of quinolone restriction on resistance patterns of Escherichia coli isolated from urine by culture in a community setting. Clin. Infect. Dis. 49, 869–875 (2009).
Trindade, S., Sousa, A. & Gordo, I. Antibiotic resistance and stress in the light of Fisher’s model. Evolution 66, 3815–3824 (2012).
Hall, A. R., Angst, D. C., Schiessl, K. T. & Ackermann, M. Costs of antibiotic resistance—separating trait effects and selective effects. Evol. Appl. 8, 261–272 (2015).
Durão, P., Trindade, S., Sousa, A. & Gordo, I. Multiple resistance at no cost: rifampicin and streptomycin a dangerous liaison in the spread of antibiotic resistance. Mol. Biol. Evol. 32, 2675–2680 (2015).
Rodríguez-Verdugo, A., Gaut, B. S. & Tenaillon, O. Evolution of Escherichia coli rifampicin resistance in an antibiotic-free environment during thermal stress. BMC Evol. Biol. 13, 50 (2013).
Silva, R. F. et al. Pervasive sign epistasis between conjugative plasmids and drug-resistance chromosomal mutations. PLoS Genet. 7, e1002181 (2011).
Knopp, M. & Andersson, D. I. Predictable phenotypes of antibiotic resistance mutations. mBio 9, e00770-18 (2018).
Roux, D. et al. Fitness cost of antibiotic susceptibility during bacterial infection. Sci. Transl. Med. 7, 297ra114 (2015).
Luo, N. et al. Enhanced in vivo fitness of fluoroquinolone-resistant Campylobacter jejuni in the absence of antibiotic selection pressure. Proc. Natl Acad. Sci. USA 102, 541–546 (2005).
Koch, G. et al. Evolution of resistance to a last-resort antibiotic in Staphylococcus aureus via bacterial competition. Cell 158, 1060–1071 (2014).
López-Rojas, R. et al. Impaired virulence and in vivo fitness of colistin-resistant Acinetobacter baumannii. J. Infect. Dis. 203, 545–548 (2011).
Björkholm, B. et al. Mutation frequency and biological cost of antibiotic resistance in Helicobacter pylori. Proc. Natl Acad. Sci. USA 98, 14607–14612 (2001).
Warner, D. M., Folster, J. P., Shafer, W. M. & Jerse, A. E. Regulation of the MtrC-MtrD-MtrE efflux-pump system modulates the in vivo fitness of Neisseria gonorrhoeae. J. Infect. Dis. 196, 1804–1812 (2007).
Björkman, J., Hughes, D. & Andersson, D. I. Virulence of antibiotic-resistant Salmonella typhimurium. Proc. Natl Acad. Sci. USA 95, 3949–3953 (1998).
Gumpert, H. et al. Transfer and persistence of a multi-drug resistance plasmid in situ of the infant gut microbiota in the absence of antibiotic treatment. Front. Microbiol. 8, 1852 (2017).
Porse, A. et al. Genome dynamics of Escherichia coli during antibiotic treatment: transfer, loss, and persistence of genetic elements in situ of the infant gut. Front. Cell. Infect. Microbiol. 7, 126 (2017).
Barreto, Â. et al. Detection of antibiotic resistant E. coli and Enterococcus spp. in stool of healthy growing children in Portugal. J. Basic Microbiol. 49, 503–512 (2009).
Hong, S. et al. Genetic characterization of atypical Shigella flexneri isolated in Korea. J. Microbiol. Biotechnol. 20, 1457–1462 (2010).
Rahmani, F., Fooladi, A. A. I., Marashi, S. M. A. & Nourani, M. R. Drug resistance in Vibrio cholerae strains isolated from clinical specimens. Acta Microbiol. Immunol. Hung. 59, 77–84 (2012).
Barroso-Batista, J. et al. The first steps of adaptation of Escherichia coli to the gut are dominated by soft sweeps. PLOS Genet. 10, e1004182 (2014).
Stebbins, R. B., Graessle, O. E. & Robinson, H. J. Studies on the absorption and excretion of streptomycin in animals. Proc. Soc. Exp. Biol. Med. 60, 68–73 (1945).
Ng, K. M. et al. Recovery of the gut microbiota after antibiotics depends on host diet, community context, and environmental reservoirs. Cell Host Microbe 26, 650–665 (2019).
Robertson, S. J. et al. Comparison of co-housing and littermate methods for microbiota standardization in mouse models. Cell Rep. 27, 1910–1919 (2019).
Barroso-Batista, J., Demengeot, J. & Gordo, I. Adaptive immunity increases the pace and predictability of evolutionary change in commensal gut bacteria. Nat. Commun. 6, 8945 (2015).
Coyte, K. Z., Schluter, J. & Foster, K. R. The ecology of the microbiome: networks, competition, and stability. Science 350, 663–666 (2015).
Posfai, A., Taillefumier, T. & Wingreen, N. S. Metabolic trade-offs promote diversity in a model ecosystem. Phys. Rev. Lett. 118, 028103 (2017).
Ley, R. E. et al. Obesity alters gut microbial ecology. Proc. Natl Acad. Sci. USA 102, 11070–11075 (2005).
Ubeda, C. et al. Familial transmission rather than defective innate immunity shapes the distinct intestinal microbiota of TLR-deficient mice. J. Exp. Med. 209, 1445–1456 (2012).
Brandis, G., Wrande, M., Liljas, L. & Hughes, D. Fitness-compensatory mutations in rifampicin-resistant RNA polymerase. Mol. Microbiol. 85, 142–151 (2012).
Maisnier-Patin, S., Berg, O. G., Liljas, L. & Andersson, D. I. Compensatory adaptation to the deleterious effect of antibiotic resistance in Salmonella typhimurium. Mol. Microbiol. 46, 355–366 (2002).
Moura de Sousa, J., Balbontín, R., Durão, P. & Gordo, I. Multidrug-resistant bacteria compensate for the epistasis between resistances. PLoS Biol. 15, e2001741 (2017).
Lourenço, M. et al. A mutational hotspot and strong selection contribute to the order of mutations selected for during Escherichia coli adaptation to the gut. PLOS Genet. 12, e1006420 (2016).
Frazão, N., Sousa, A., Lässig, M. & Gordo, I. Horizontal gene transfer overrides mutation in Escherichia coli colonizing the mammalian gut. Proc. Natl Acad. Sci. USA 116, 17906–17915 (2019).
Ghalayini, M. et al. Evolution of a dominant natural isolate of Escherichia coli in the human gut over the course of a year suggests a neutral evolution with reduced effective population size. Appl. Environ. Microbiol. 84, e02377-17 (2018).
Jakobsson, H. E. et al. Short-term antibiotic treatment has differing long-term impacts on the human throat and gut microbiome. PLoS ONE 5, e9836 (2010).
Jernberg, C., Löfmark, S., Edlund, C. & Jansson, J. K. Long-term ecological impacts of antibiotic administration on the human intestinal microbiota. ISME J. 1, 56–66 (2007).
Qi, Q., Preston, G. M. & MacLean, R. C. Linking system-wide impacts of RNA polymerase mutations to the fitness cost of rifampin resistance in Pseudomonas aeruginosa. mBio 5, e01562–14 (2014).
Barnard, A. M. L., Simpson, N. J. L., Lilley, K. S. & Salmond, G. P. C. Mutations in rpsL that confer streptomycin resistance show pleiotropic effects on virulence and the production of a carbapenem antibiotic in Erwinia carotovora. Microbiology 156, 1030–1039 (2010).
Robinson, L. J., Cameron, A. D. S. & Stavrinides, J. Spontaneous and on point: do spontaneous mutations used for laboratory experiments cause pleiotropic effects that might confound bacterial infection and evolution assays? FEMS Microbiol. Lett. 362, fnv177 (2015).
Ruusala, T., Andersson, D., Ehrenberg, M. & Kurland, C. G. Hyper-accurate ribosomes inhibit growth. EMBO J. 3, 2575–2580 (1984).
Libby, R. T., Nelson, J. L., Calvo, J. M. & Gallant, J. A. Transcriptional proofreading in Escherichia coli. EMBO J. 8, 3153–3158 (1989).
Blank, A., Gallant, J. A., Burgess, R. R. & Loeb, L. A. An RNA polymerase mutant with reduced accuracy of chain elongation. Biochemistry 25, 5920–5928 (1986).
Strathern, J. N., Jin, D. J., Court, D. L. & Kashlev, M. Isolation and characterization of transcription fidelity mutants. Biochim. Biophys. Acta 1819, 694–699 (2012).
Li, J. et al. Antibiotic treatment drives the diversification of the human gut resistome. Genom. Proteom. Bioinform. 17, 39–51 (2019).
Sousa, A. et al. Recurrent reverse evolution maintains polymorphism after strong bottlenecks in commensal gut bacteria. Mol. Biol. Evol. 34, 2879–2892 (2017).
Muinck, E. Jde et al. Context-dependent competition in a model gut bacterial community. PLoS ONE 8, e67210 (2013).
Barroso-Batista, J. et al. Specific eco-evolutionary contexts in the mouse gut reveal Escherichia coli metabolic versatility. Curr. Biol. 30, 1049–1062.e7 (2020).
Tramontano, M. et al. Nutritional preferences of human gut bacteria reveal their metabolic idiosyncrasies. Nat. Microbiol. 3, 514–522 (2018).
Görke, B. & Stülke, J. Carbon catabolite repression in bacteria: many ways to make the most out of nutrients. Nat. Rev. Microbiol. 6, 613–624 (2008).
Kovárová-Kovar, K. & Egli, T. Growth kinetics of suspended microbial cells: from single-substrate-controlled growth to mixed-substrate kinetics. Microbiol. Mol. Biol. Rev. 62, 646–666 (1998).
Chang, D.-E. et al. Carbon nutrition of Escherichia coli in the mouse intestine. Proc. Natl Acad. Sci. USA 101, 7427–7432 (2004).
Belenguer, A. et al. Two routes of metabolic cross-feeding between Bifidobacterium adolescentis and butyrate-producing anaerobes from the human gut. Appl. Environ. Microbiol. 72, 3593–3599 (2006).
Samuel, B. S. & Gordon, J. I. A humanized gnotobiotic mouse model of host-archaeal-bacterial mutualism. Proc. Natl Acad. Sci. USA. 103, 10011–10016 (2006).
Goldford, J. E. et al. Emergent simplicity in microbial community assembly. Science 361, 469–474 (2018).
Filippo, C. D. et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc. Natl Acad. Sci. USA 107, 14691–14696 (2010).
Franzosa, E. A. et al. Identifying personal microbiomes using metagenomic codes. Proc. Natl Acad. Sci. USA 112, E2930–E2938 (2015).
Thompson, J. A., Oliveira, R. A., Djukovic, A., Ubeda, C. & Xavier, K. B. Manipulation of the quorum sensing signal AI-2 affects the antibiotic-treated gut microbiota. Cell Rep. 10, 1861–1871 (2015).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Mandal, S. et al. Analysis of composition of microbiomes: a novel method for studying microbial composition. Microb. Ecol. Health Dis. 26, 27663 (2015).
Lozupone, C. & Knight, R. UniFrac: a new phylogenetic method for comparing microbial communities. Appl. Environ. Microbiol. 71, 8228–8235 (2005).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072 (2006).
Soetaert, K., Petzoldt, T. & Setzer, R. W. Solving differential equations in R: package deSolve. J. Stat. Softw. 33, 1–25 (2010).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer-Verlag, 2016); https://ggplot2.tidyverse.org
Oksanen, J. et al. vegan: Community Ecology Package. R package version 2.5-6 (2019); https://CRAN.R-project.org/package=vegan
Acknowledgements
We thank the personnel of the IGC Rodent Facility, Genomic Facility and the Bioinformatics Unit for their assistance and A. Konrad for reading of the paper. L.L.C., P.D. and M.A. were supported by Fundação para a Ciência e Tecnologia (FCT) fellowships PD/BD/106003/2014, SFRH/BPD/118474/2016 and PD/BD/138735/2018, respectively. Research was supported by project JPIAMR/0001/2016-ERA NET and ONEIDA project (LISBOA-01-0145-FEDER-016417) cofunded by FEEI—Fundos Europeus Estruturais e de Investimento from Programa Operacional Regional Lisboa 2020, and by national funds from FCT.
Author information
Authors and Affiliations
Contributions
L.L.C. and P.D. performed the experiments. M.A. performed the modelling. All of the authors analysed the results. I.G. designed and coordinated the study. All of the authors contributed to writing the manuscript and provided final approval for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 An indirect method for streptomycin detection in fecal samples.
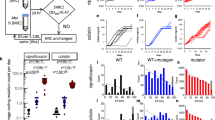
a, Calibration curve to measure the concentration of streptomycin that provides a fitness advantage. Effect of different concentrations of streptomycin on fitness (+2SE) from pairwise competitions between E. coli MG1655 resistant to streptomycin (RpsLK43T or RpsLK43R single mutants or double mutant RpsLK43TRpoBH526Y, MIC above detection level > 8192 μg/ml) against a susceptible strain (MIC ≈ 2 μg/ml) in fecal medium supplemented with LB (n = 3 replicates per resistant mutant). Fecal samples of SPF mice not treated with antibiotic were resuspended (8 mL of PBS per gram of fecal content) and then filtered to remove the bacteria. To enrich the fecal medium with nutrients, 2 ml of LB was added per 8 mL of filtered medium. Increasing concentrations of antibiotic were added to this mixed fecal medium and pairwise competitions of 1:1 between resistant and susceptible strains were performed for 24 h at 37 °C. Fitness of the StrR mutants were measured as a ratio of Ln(Mut/WT) per generation. The method does not allow a reliable detection of streptomycin at concentrations below ≈2 µg/ml of streptomycin, as at these concentrations the mutants still show a fitness cost. b, The amount of streptomycin detected on the fecal samples of the single housed SPF mice (Fig. 1b), which were collected 4 h after the gavage of E. coli (20 h before the first time point in Fig. 1b). A sample of mice which receives continuous antibiotic treatment is shown as a red bar. Samples were filtered in 1 ml PBS and 250 μl LB added and competitions between the resistant (rpsLK43R) and susceptible strains were performed in triplicate. No antibiotic was detected in any sample using this assay (except the positive control).
Extended Data Fig. 2 Fitness effects of double and single resistances are individualized in the presence of a complex microbiome.
The fitness costs of the double mutant StrRRifR (DM) were measured against the single StrR (Mut) and against the single RifR in both in 3 single housed three specific-pathogen-free (SPF) mice carrying a complex microbiota (right panels, n = 3) and in germ-free devoid of microbiota (left panels, n = 3). Each mouse is represented by a different symbol (circles, squares and diamonds) and SPF mice have additional colors to highlight fitness effects differences (red and green represent cost or no cost of the AR mutation on that mouse, respectively). High variance of AR fitness effects across mice is observed when competing the double against the single resistances in the presence of a microbiota. This variance is observed even when both strains are resistant to streptomycin (StrRRifR vs StrR). To inquire if fitness effects were transitive, the fitness effects estimated from these competitions were compared to the values estimated from the competitions of the single mutants against the susceptible strain (Supplementary Table 6).
Extended Data Fig. 3 Effect of streptomycin perturbation on the microbiota composition shown as the relative frequencies of bacteria at the phylum level.
Microbiota composition was assayed through 16S amplicon sequencing, before and after antibiotic treatment, a, in individual caged mice and b, in co-housed mice. Each bar graph represents the composition in each mouse. In (b) the mice that were co-housed are shown by the same color (6 mice in green, 6 mice in blue and 6 mice in gray).
Extended Data Fig. 4 Co-housed animals share a more similar microbiota.
Principal coordinate Analysis (PCoA) from Unweighted UniFrac diversity measure of the microbiotas after antibiotic treatment (Day 1 of the competition in Figs. 1b and 1c). Co-Housed animals (green circles (6 mice where the fitness effect of StrR was measured), blue circles (6 mice where the fitness effect of RifR was measured), and grey circles (5 mice where the fitness effect of the double resistant was measured)) cluster accordingly to the housing and show reduced variance when compared with the singly housed animals (white circles). Variance analysis was conducted via the pairwise distances from the centroids of each group by using permutational ANOVA on the dispersion test (betadisper). The ellipses represent the standard deviation of point scores with a 95% confidence limit for each group.
Extended Data Fig. 5 Fitness effects and level of epistasis in vitro differ from the in vivo ones.
a, Fitness effects of each resistance mutation - StrR (rpsLK43T, in green), RifR (rpoBH526Y, in blue), StrRRifR (rpsLK43TrpoBH526Y, in grey) – in two commonly used in vitro environments: rich Luria Broth (LB) media and minimal media supplemented with 0.4% glucose (MM)- and in germ free (GF). Fitness effects in GF (shown in full bars darker color) from Fig. 1d (n = 3 mice, 5-day competition). Fitness effects in each in vitro media (LB, shown in full bars light color, and MM, shown in slashed bars, n = 3, 1 day-competition) with the same strains used in the GF mice. Error bars represent 2SE. b, Level of epistasis (ε) was measured assuming an additive model and its error calculated by error propagation1. Epistasis in LB and MM is shown in full grey bars and slashed bars, respectively. No epistasis is detected in GF (see Supplementary Table 6). The p-values of a 2-sided t-test are shown.
Extended Data Fig. 6 Pairwise correlation between the fitness effect of mutations and the Weighted UniFrac principal coordinates (PCoAs).
PCoAs from Weighted UniFrac diversity measure of the microbiotas after antibiotic treatment (Day 1 of the competition) grouped by mutation background a, StrR, b, RifR and c, StrRRifR). a-b, Both the fitness effects of StrR and RifR mutations correlate significantly with the PC2 (Pearson’s R and p-value shown).
Extended Data Fig. 7 Resistance mutations alter metabolism on specific carbon sources.
Growth curves were measured in minimal media supplemented either with 0.4% Glucose or 0.4% Mannose. a, Maximum growth rates in either of the sugars is shown for each of strains (susceptible in black, StrR in green, RifR in red and StrRRifR in gray). b, Carrying capacity measured as final OD600nm in both sugars. G and M stand for significant changes (P < 0.05, after correction for multiple testing) in glucose and mannose, respectively (N = 4, for each strain in each carbon source, Mann-Whitney test for comparison of medians between each resistance and the susceptible).
Extended Data Fig. 8 Selection is time-dependent and depends on the level of pleiotropy of the mutation.
a, Long-term dynamics of mutant x with low pleiotropy (left panel) and y with high pleiotropy (right panel) in each ecosystem (100 simulated microbiomes). b, Quantification of the selection coefficient (slope) from panel (a). Mutant x, (defined in Fig. 2), that represents a mutation conserving the trait ratio, shows similar fitness effect across the perturbed microbiomes, implying that pleiotropy is a necessary condition for the cost of resistance to be host-specific. Consistently mutant y shows dependence on the host microbiome. Over time the average cost of mutant y tends to the analytical prediction, which has a small value since for this mutation the presence of other species at equilibrium buffers its effect (Fig. 2).
Extended Data Fig. 9 Evidence for a time-dependent cost of AR in long-term competitions is consistent with model predictions.
The dynamics of Ln(Mut/WT) were followed for two weeks in 4 mice colonized with a mix of StrRRifR and RifR strains a, a mix of RifR and WT strains b, and a mix of StrRRifR and WT strains c, d. A long-term cost of resistance is observed (slopes (per day), after day 5 of the competitions: −0.2, p = 0.01; −0.37 p = 0.007; −0.37, p = 0.04; −0.27, p = 0.001; for (a), (b), (c) and (d), respectively, linear regression shown as black lines) even when no cost or even an increase in frequency of the resistance is detected in the short term (slopes (per day), during the initial days of the competitions: +0.3, p = 0.006; −0.1, p = 0.5; +0.43, p = 0.1; −0.26, p = 0.15; for (a), (b), (c) and (d), respectively, linear regression shown as red lines), as predicted by the model of resource competition. Fit of all the time points of the competition between StrRRifR and RifR strains to a quadratic model (indicating variable selection over time) is also shown in (a) (p = 0.0003, R2 = 0.93, quadratic model shown as green lines).
Extended Data Fig. 10 Loads of the resistant strains of E. coli colonizing the mice at weeks 3 and 6 of the evolution experiment.
The number of CFUs per feces gram are shown (pink measured at week 3 and red at week 6) for each mouse and each resistance background (x-axis). No significant correlation is found between the mean population size and the number of mutations accumulated (Spearman Correlation on the Log of the loads of E. coli with the number of mutations detected at week 6: Rs = 0.46, P = 0.056).
Supplementary information
Supplementary Information
Supplementary Figs. 1–4, Tables 1–7 and equations.
Rights and permissions
About this article
Cite this article
Leónidas Cardoso, L., Durão, P., Amicone, M. et al. Dysbiosis individualizes the fitness effect of antibiotic resistance in the mammalian gut. Nat Ecol Evol 4, 1268–1278 (2020). https://doi.org/10.1038/s41559-020-1235-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-020-1235-1
This article is cited by
-
Intrahost evolution of the gut microbiota
Nature Reviews Microbiology (2023)
-
Gut microbiota response to antibiotics is personalized and depends on baseline microbiota
Microbiome (2021)
-
Probiotics impact the antibiotic resistance gene reservoir along the human GI tract in a person-specific and antibiotic-dependent manner
Nature Microbiology (2021)