Abstract
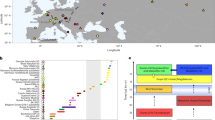
Steppe-pastoralist-related ancestry reached Central Europe by at least 2500 bc, whereas Iranian farmer-related ancestry was present in Aegean Europe by at least 1900 bc. However, the spread of these ancestries into the western Mediterranean, where they have contributed to many populations that live today, remains poorly understood. Here, we generated genome-wide ancient-DNA data from the Balearic Islands, Sicily and Sardinia, increasing the number of individuals with reported data from 5 to 66. The oldest individual from the Balearic Islands (~2400 bc) carried ancestry from steppe pastoralists that probably derived from west-to-east migration from Iberia, although two later Balearic individuals had less ancestry from steppe pastoralists. In Sicily, steppe pastoralist ancestry arrived by ~2200 bc, in part from Iberia; Iranian-related ancestry arrived by the mid-second millennium bc, contemporary to its previously documented spread to the Aegean; and there was large-scale population replacement after the Bronze Age. In Sardinia, nearly all ancestry derived from the island’s early farmers until the first millennium bc, with the exception of an outlier from the third millennium bc, who had primarily North African ancestry and who—along with an approximately contemporary Iberian—documents widespread Africa-to-Europe gene flow in the Chalcolithic. Major immigration into Sardinia began in the first millennium bc and, at present, no more than 56–62% of Sardinian ancestry is from its first farmers. This value is lower than previous estimates, highlighting that Sardinia, similar to every other region in Europe, has been a stage for major movement and mixtures of people.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All raw data are available at the European Nucleotide Archive under accession number PRJEB35980 and at https://reich.hms.harvard.edu/datasets.
Code availability
All custom code used in this study is provided at https://github.com/DReichLab/ADNA-Tools.
Change history
15 April 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41559-020-1197-3
References
Allentoft, M. E. et al. Population genomics of Bronze Age Eurasia. Nature 522, 167–172 (2015).
Haak, W. et al. Massive migration from the steppe was a source for Indo-European languages in Europe. Nature 522, 207–211 (2015).
Kristiansen, K. Re-theorising mobility and the formation of culture and language among the corded ware culture in Europe. Antiquity 91, 334–347 (2017).
Cassidy, L. M. et al. Neolithic and Bronze Age migration to Ireland and establishment of the insular Atlantic genome. Proc. Natl Acad. Sci. USA 113, 368–373 (2016).
Olalde, I. et al. The Beaker phenomenon and the genomic transformation of northwest Europe. Nature 555, 190–196 (2018).
Martiniano, R. et al. The population genomics of archaeological transition in west Iberia: investigation of ancient substructure using imputation and haplotype-based methods. PLoS Genet. 13, e1006852 (2017).
Lazaridis, I. et al. Genetic origins of the Minoans and Mycenaeans. Nature 548, 214–218 (2017).
Alcover, J. A. The first Mallorcans: prehistoric colonization in the western Mediterranean. J. World Prehist. 21, 19–84 (2008).
Ramis, D. in The Cambridge Prehistory of the Bronze and Iron Age Mediterranean (eds Knapp, A. B. & van Dommelen, P.) 40–56 (Cambridge Univ. Press, 2014).
Ramis, D. Animal exploitation in the early prehistory of the Balearic Islands. J. Isl. Coast. Archaeol. 13, 269–282 (2018).
Plantalamor, L. & Van Strydonck, M. La Cronologia de la Prehistòria de Menorca: (Noves datacions de 14C). Treballs del Museu de Menorca Vol. 20 (Museu de Menorca, 1997).
Lull, V., Mico, R., Rihuete, C. I. & Risch, R. Los cambios sociales en las Islas Baleares a lo largo del II milenio. Cypsela 15, 123–148 (2004).
Holt, E. in Forging Identities. The Mobility of Culture in Bronze Age Europe Vol. 1 (eds Suchowska-Ducke, P., Reiter, S. S. & Vandkilde, H.) 193–202 (British Archaeological Reports, 2015).
Ugas, G. L’alba Dei Nuraghi (Fabula, 2005).
Sestieri, A. M. B. in The Oxford Handbook of European Bronze Age (eds Harding, A. & Fokkens, H.) 653–667 (Oxford Univ. Press, 2013).
Sarno, S. et al. Ancient and recent admixture layers in Sicily and Southern Italy trace multiple migration routes along the Mediterranean. Sci. Rep. 7, 1984 (2017).
Dabney, J. et al. Complete mitochondrial genome sequence of a Middle Pleistocene cave bear reconstructed from ultrashort DNA fragments. Proc. Natl Acad. Sci. USA 110, 15758–15763 (2013).
Damgaard, P. B. et al. Improving access to endogenous DNA in ancient bones and teeth. Sci. Rep. 5, 11184 (2015).
Korlević, P. et al. Reducing microbial and human contamination in DNA extractions from ancient bones and teeth. Biotechniques 59, 87–93 (2015).
Rohland, N., Glocke, I., Aximu-Petri, A. & Meyer, M. Extraction of highly degraded DNA from ancient bones, teeth and sediments for high-throughput sequencing. Nat. Protoc. 13, 2447–2461 (2018).
Rohland, N., Harney, E., Mallick, S., Nordenfelt, S. & Reich, D. Partial uracil-DNA-glycosylase treatment for screening of ancient DNA. Proc. R. Soc. B 370, 20130624 (2015).
Gansauge, M.-T. et al. Single-stranded DNA library preparation from highly degraded DNA using T4 DNA ligase. Nucleic Acids Res. 45, e79 (2017).
Fu, Q. et al. A revised timescale for human evolution based on ancient mitochondrial genomes. Curr. Biol. 23, 553–559 (2013).
Fu, Q. et al. An early modern human from Romania with a recent Neanderthal ancestor. Nature 524, 216–219 (2015).
Olalde, I. et al. The genomic history of the Iberian Peninsula over the past 8000 years. Science 363, 1230–1234 (2019).
Keller, A. et al. New insights into the Tyrolean Iceman’s origin and phenotype as inferred by whole-genome sequencing. Nat. Commun. 3, 698 (2012).
Lazaridis, I. et al. Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature 513, 409–413 (2014).
Gamba, C. et al. Genome flux and stasis in a five millennium transect of European prehistory. Nat. Commun. 5, 5257 (2014).
Olalde, I. et al. Derived immune and ancestral pigmentation alleles in a 7,000-year-old Mesolithic European. Nature 507, 225–228 (2014).
Raghavan, M. et al. Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans. Nature 505, 87–91 (2014).
Skoglund, P. et al. Genomic diversity and admixture differs for Stone-Age Scandinavian foragers and farmers. Science 344, 747–750 (2014).
Günther, T. et al. Ancient genomes link early farmers from Atapuerca in Spain to modern-day Basques. Proc. Natl Acad. Sci. USA 112, 11917–11922 (2015).
Jones, E. R. et al. Upper Palaeolithic genomes reveal deep roots of modern Eurasians. Nat. Commun. 6, 8912 (2015).
Mathieson, I. et al. Genome-wide patterns of selection in 230 ancient Eurasians. Nature 528, 499–503 (2015).
Olalde, I. et al. A common genetic origin for early farmers from Mediterranean Cardial and Central European LBK cultures. Mol. Biol. Evol. 32, 3132–3142 (2015).
Broushaki, F. et al. Early Neolithic genomes from the eastern Fertile Crescent. Science 353, 499–503 (2016).
Fu, Q. et al. The genetic history of Ice Age Europe. Nature 534, 200–205 (2016).
Hofmanová, Z. et al. Early farmers from across Europe directly descended from Neolithic Aegeans. Proc. Natl Acad. Sci. USA 113, 6886–6891 (2016).
Kılınç, G. M. et al. The demographic development of the first farmers in Anatolia. Curr. Biol. 26, 2659–2666 (2016).
Lazaridis, I. et al. Genomic insights into the origin of farming in the ancient Near East. Nature 536, 419–424 (2016).
Martiniano, R. et al. Genomic signals of migration and continuity in Britain before the Anglo-Saxons. Nat. Commun. 7, 10326 (2016).
Schiffels, S. et al. Iron age and Anglo-Saxon genomes from East England reveal British migration history. Nat. Commun. 7, 10408 (2016).
González-Fortes, G. et al. Paleogenomic evidence for multi-generational mixing between Neolithic farmers and Mesolithic hunter-gatherers in the Lower Danube basin. Curr. Biol. 27, 1801–1810 (2017).
Haber, M. et al. Continuity and admixture in the last five millennia of Levantine history from ancient Canaanite and present-day Lebanese genome sequences. Am. J. Hum. Genet. 101, 274–282 (2017).
Jones, E. R. et al. The Neolithic transition in the Baltic was not driven by admixture with early European farmers. Curr. Biol. 27, 576–582 (2017).
Lipson, M. et al. Parallel palaeogenomic transects reveal complex genetic history of early European farmers. Nature 551, 368–372 (2017).
Saag, L. et al. Extensive farming in Estonia started through a sex-biased migration from the steppe. Curr. Biol. 27, 2185–2193 (2017).
Schuenemann, V. J. et al. Ancient Egyptian mummy genomes suggest an increase of sub-Saharan African ancestry in post-Roman periods. Nat. Commun. 8, 15694 (2017).
Unterländer, M. et al. Ancestry and demography and descendants of Iron Age nomads of the Eurasian steppe. Nat. Commun. 8, 14615 (2017).
Amorim, C. E. G. et al. Understanding 6th-century barbarian social organization and migration through paleogenomics. Nat. Commun. 9, 3547 (2018).
Damgaard, P. et al. The first horse herders and the impact of Early Bronze Age steppe expansions into Asia. Science 360, eaar7711 (2018).
Fernandes, D. M. et al. A genomic Neolithic time transect of hunter-farmer admixture in central Poland. Sci. Rep. 8, 14879 (2018).
Fregel, R. et al. Ancient genomes from North Africa evidence prehistoric migrations to the Maghreb from both the Levant and Europe. Proc. Natl Acad. Sci. USA 115, 6774–6779 (2018).
Mathieson, I. et al. The genomic history of southeastern Europe. Nature 555, 197–203 (2018).
Mittnik, A. et al. The genetic prehistory of the Baltic Sea region. Nat. Commun. 9, 442 (2018).
Valdiosera, C. et al. Four millennia of Iberian biomolecular prehistory illustrate the impact of prehistoric migrations at the far end of Eurasia. Proc. Natl Acad. Sci. USA 115, 3428–3433 (2018).
van de Loosdrecht, M. et al. Pleistocene North African genomes link Near Eastern and sub-Saharan African human populations. Science 360, 548–552 (2018).
Veeramah, K. R. et al. Population genomic analysis of elongated skulls reveals extensive female-biased immigration in Early Medieval Bavaria. Proc. Natl Acad. Sci. USA 115, 3494–3499 (2018).
Zalloua, P. et al. Ancient DNA of Phoenician remains indicates discontinuity in the settlement history of Ibiza. Sci. Rep. 8, 17567 (2018).
Feldman, M. et al. Late Pleistocene human genome suggests a local origin for the first farmers of central Anatolia. Nat. Commun. 10, 1218 (2019).
González-Fortes, G. et al. A western route of prehistoric human migration from Africa into the Iberian Peninsula. Proc. R. Soc. B 286, 20182288 (2019).
Narasimhan, V. M. et al. The formation of human populations in South and Central Asia. Science 365, eaat7487 (2019).
Patterson, N. et al. Ancient admixture in human history. Genetics 192, 1065–1093 (2012).
Pickrell, J. K. et al. The genetic prehistory of southern Africa. Nat. Commun. 3, 1143 (2012).
Qin, P. & Stoneking, M. Denisovan ancestry in east Eurasian and native American populations. Mol. Biol. Evol. 32, 2665–2674 (2015).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Ramis, D., Alcover, J. A., Coll, J. & Trias, M. The chronology of the first settlement of the Balearic Islands. J. Mediterr. Archaeol. 15, 3–24 (2002).
Picornell, A., Gómez-Barbeito, L., Tomàs, C., Castro, J. A. & Ramon, M. M. Mitochondrial DNA HVRI variation in Balearic populations. Am. J. Phys. Anthropol. 128, 119–130 (2005).
Adams, S. M. et al. The genetic legacy of religious diversity and intolerance: paternal lineages of Christians, Jews, and Muslims in the Iberian Peninsula. Am. J. Hum. Genet. 83, 725–736 (2008).
Solé-Morata, N. et al. Analysis of the R1b-DF27 haplogroup shows that a large fraction of Iberian Y-chromosome lineages originated recently in situ. Sci. Rep. 7, 7341 (2017).
Raveane, A. et al. Population structure of modern-day Italians reveals patterns of ancient and archaic ancestries in southern Europe. Sci. Adv. 5, eaaw3492 (2019).
Di Gaetano, C. et al. Differential Greek and northern African migrations to Sicily are supported by genetic evidence from the Y chromosome. Eur. J. Hum. Genet. 17, 91–99 (2009).
Sarno, S. et al. An ancient Mediterranean melting pot: investigating the uniparental genetic structure and population history of Sicily and southern Italy. PLoS ONE 9, e96074 (2014).
Holt, E. M. Economy and Environment in Complex Societies: A Case Study from Bronze Age Sardinia (Univ. Michigan, 2013).
Magoon, G. R. et al. Generation of high-resolution a priori Y-chromosome phylogenies using ‘next-generation’ sequencing data. Preprint at bioRxiv https://doi.org/10.1101/000802 (2013).
Wang, C.-C. et al. Ancient human genome-wide data from a 3000-year interval in the Caucasus corresponds with eco-geographic regions. Nat. Commun. 10, 590 (2019).
Matisoo-Smith, E. et al. Ancient mitogenomes of Phoenicians from Sardinia and Lebanon: a story of settlement, integration, and female mobility. PLoS ONE 13, e0190169 (2018).
Sikora, M. et al. Population genomic analysis of ancient and modern genomes yields new insights into the genetic ancestry of the Tyrolean Iceman and the genetic structure of Europe. PLoS Genet. 10, e1004353 (2014).
Chiang, C. W. K. et al. Genomic history of the Sardinian population. Nat. Genet. 50, 1426–1434 (2018).
Moorjani, P. et al. The history of African gene flow into southern Europeans, Levantines, and Jews. PLoS Genet. 7, e1001373 (2011).
Loh, P.-R. et al. Inferring admixture histories of human populations using linkage disequilibrium. Genetics 193, 1233–1254 (2013).
Hellenthal, G. et al. A genetic atlas of human admixture history. Science 343, 747–751 (2014).
Olivieri, A. et al. Mitogenome diversity in Sardinians: a genetic window onto an Island’s past. Mol. Biol. Evol. 34, 1230–1239 (2017).
Morelli, L. et al. A comparison of Y-chromosome variation in Sardinia and Anatolia is more consistent with cultural rather than demic diffusion of agriculture. PLoS ONE 5, e10419 (2010).
Marcus, J. H. et al. Genetic history from the Middle Neolithic to present on the Mediterranean island of Sardinia. Nat. Commun. https://doi.org/10.1038/s41467-020-14523-6 (2020).
Sangmeister, E. Die datierung des rickstroms der Glockenbecker und ihre auswirkung auf die chronologie der Kupferzeit in Portugal. Palaeohistoria 12, 395–407 (1966).
Holloway, R. The Archaeology of Ancient Sicily (Routledge, 2000).
D’Agata, A. L. Interactions between Aegean groups and local communities in Sicily in the Bronze Age: the evidence from pottery. Stud. Micenei ed Egeo-Anatolici 42, 61–83 (2000).
Shelton, K. in The Oxford Handbook of the Bronze Age Aegean (ed. Kline, E.) 139–148 (Oxford Univ. Press, 2012).
Alberti, G. Issues in the absolute chronology of the Early-Middle Bronze Age transition in Sicily and southern Italy: a Bayesian radiocarbon view. J. Quat. Sci. 28, 630–640 (2013).
Heyd, V. in The Oxford Handbook of the European Bronze Age (ed. Harding, A.) 47–67 (Oxford Univ. Press, 2013).
Sabatini, S. Late Bronze Age oxhide and oxhide-like ingots from areas other than the Mediterranean: problems and challenges. Oxf. J. Archaeol. 35, 29–45 (2016).
Aubet, M. E. & Turton, M. The Phoenicians and the West: Politics, Colonies and Trade (Cambridge Univ. Press, 1997).
Pinhasi, R. et al. Optimal ancient DNA Yields from the inner ear part of the human petrous bone. PLoS ONE 10, e0129102 (2015).
Pinhasi, R., Fernandes, D. M., Sirak, K. & Cheronet, O. Isolating the human cochlea to generate bone powder for ancient DNA analysis. Nat. Protoc. 14, 1194–1205 (2019).
Briggs, A. W. et al. Removal of deaminated cytosines and detection of in vivo methylation in ancient DNA. Nucleic Acids Res. 38, e87 (2010).
Maricic, T., Whitten, M. & Pääbo, S. Multiplexed DNA sequence capture of mitochondrial genomes using PCR products. PLoS ONE 5, e14004 (2010).
Behar, D. M. et al. A ‘Copernican’ reassessment of the human mitochondrial DNA tree from its root. Am. J. Hum. Genet. 90, 675–684 (2012).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Weissensteiner, H. et al. HaploGrep 2: mitochondrial haplogroup classification in the era of high-throughput sequencing. Nucleic Acids Res. 44, W58–W63 (2016).
Korneliussen, T. S., Albrechtsen, A. & Nielsen, R. ANGSD: analysis of next generation sequencing data. BMC Bioinform. 15, 356 (2014).
Kennett, D. J. et al. Archaeogenomic evidence reveals prehistoric matrilineal dynasty. Nat. Commun. 8, 14115 (2017).
Lohse, J. C., Madsen, D. B., Culleton, B. J. & Kennett, D. J. Isotope paleoecology of episodic mid-to-late Holocene bison population expansions in the southern Plains, U.S.A. Quat. Sci. Rev. 102, 14–26 (2014).
van Klinken, G. J. Bone collagen quality indicators for palaeodietary and radiocarbon measurements. J. Archaeol. Sci. 26, 687–695 (1999).
Santos, G. M., Southon, J. R., Druffel-Rodriguez, K. C., Griffin, S. & Mazon, M. Magnesium perchlorate as an alternative water trap in AMS graphite sample preparation: a report on sample preparation at Kccams at the University of California, Irvine. Radiocarbon 46, 165–173 (2004).
Stuiver, M. & Polach, H. A. Discussion reporting of 14C data. Radiocarbon 19, 355–363 (1977).
Ramsey, C. B. & Lee, S. Recent and planned developments of the program OxCal. Radiocarbon 55, 720–730 (2013).
Reimer, P. J. et al. IntCal13 and Marine13 radiocarbon age calibration curves 0–50,000 years cal BP. Radiocarbon 55, 1869–1887 (2013).
van Oven, M. & Kayser, M. Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation. Hum. Mutat. 30, E386–E394 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Skoglund, P. et al. Origins and genetic legacy of Neolithic farmers and hunter-gatherers in Europe. Science 336, 466–469 (2012).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Acknowledgements
This manuscript is dedicated to the memory of S. Tusa of the Soprintendenza del Mare in Palermo, who would have been an author of this study had he not tragically died in the crash of Ethiopia Airlines flight 302 on 10 March 2019. We thank Z. Zhang for database support; the Soprintendenza BBCCAA Palermo and R. Schicchi (director of Museum of Castelbuono) for facilitating access to important skeletal materials. D.M.F. was supported by an Irish Research Council grant GOIPG/2013/36. Radiocarbon work was supported in part by the NSF Archaeometry program BCS-1460369 (to D.J.K. and B.J.C). C.L.-F. was supported by Obra Social La Caixa and by FEDER-MINECO (BFU2015-64699-P and PGC2018-095931-B-100). D.C. was supported by grant 20177PJ9XF MIUR PRIN 2017. D.Reich is an Investigator of the Howard Hughes Medical Institute, and his ancient-DNA laboratory work was supported by National Science Foundation HOMINID grant BCS-1032255, a National Institutes of Health grant GM100233, an Allen Discovery Center grant, and grant no. 61220 from the John Templeton Foundation.
Author information
Authors and Affiliations
Contributions
D.M.F., D.Reich and R.P. conceived the study. D.M.F., E.C., C.C., G.C., M.C., V.F., M.Lozano, E.M., M.Michel, R.M.M., D.Ramis, M.R.P., V.S., P.S., L.T., M.T.-N., C.L.-F., L.S., D.C., A.C., M.Lucci, G.G., F.C., G.S. and R.P. excavated, assembled and/or studied the osteological material. D.M.F., O.C., N.R., N.B., M.F., B.G., M.Lari, M.Micheletti, A.Modi, M.N., F.C., J.O., K.A.S., K.S., K.M., C.S., K.T.Ö. and S.V. performed laboratory work under the supervision of N.R., D.C. and R.P. J.C. provided computing resources. B.J.C. performed radiocarbon analysis under the supervision of D.J.K. D.M.F., I.O., R.B., S.M. and M.Mah performed bioinformatics and population genetics analysis with input from A.Mittnik, I.L., N.P. and D.Reich.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–4, Tables 1–27, notes and references, and legends for Supplementary Data 1–6.
Supplementary Data
Supplementary Data 1–6: six supplementary tables in Excel (a single file with six tabs).
Rights and permissions
About this article
Cite this article
Fernandes, D.M., Mittnik, A., Olalde, I. et al. The spread of steppe and Iranian-related ancestry in the islands of the western Mediterranean. Nat Ecol Evol 4, 334–345 (2020). https://doi.org/10.1038/s41559-020-1102-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-020-1102-0
This article is cited by
-
The Allen Ancient DNA Resource (AADR) a curated compendium of ancient human genomes
Scientific Data (2024)
-
Bioarchaeological and paleogenomic profiling of the unusual Neolithic burial from Grotta di Pietra Sant’Angelo (Calabria, Italy)
Scientific Reports (2023)
-
Socio-cultural practices may have affected sex differences in stature in Early Neolithic Europe
Nature Human Behaviour (2023)
-
Multiproxy bioarchaeological data reveals interplay between growth, diet and population dynamics across the transition to farming in the central Mediterranean
Scientific Reports (2023)
-
A genetic history of continuity and mobility in the Iron Age central Mediterranean
Nature Ecology & Evolution (2023)