Abstract
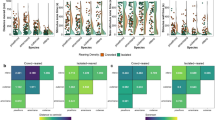
Genetic interactions such as epistasis are widespread in nature and can shape evolutionary dynamics. Epistasis occurs due to nonlinearity in biological systems, which can arise via cellular processes that convert genotype to phenotype and via selective processes that connect phenotype to fitness. Few studies in nature have connected genotype to phenotype to fitness for multiple potentially interacting genetic variants. Thus, the causes of epistasis in the wild remain poorly understood. Here, we show that epistasis for fitness is an emergent and predictable property of nonlinear selective processes. We do so by measuring the genetic basis of cryptic colouration and survival in a field experiment with stick insects. We find that colouration shows a largely additive genetic basis but with some effects of epistasis that enhance differentiation between colour morphs. In terms of fitness, different combinations of loci affecting colouration confer high survival in one host-plant treatment. Specifically, nonlinear correlational selection for specific combinations of colour traits in this treatment drives the emergence of pairwise and higher-order epistasis for fitness at loci underlying colour. In turn, this results in a rugged fitness landscape for genotypes. In contrast, fitness epistasis was dampened in another treatment, where selection was weaker. Patterns of epistasis that are shaped by ecologically based selection could be common and central to understanding fitness landscapes, the dynamics of evolution and potentially other complex systems.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
DNA sequences have been deposited in the NCBI SRA (BioProject PRJNA656892). Other data, including colour measurements and results from the experiment, have been archived in Dryad Digital Repository (https://doi.org/10.5061/dryad.2z34tmpjr). Correspondence and requests for materials should be sent to P.N.
Code availability
Core scripts have been archived in Dryad Digital Repository (https://doi.org/10.5061/dryad.2z34tmpjr).
References
Barrett, R. D. H. & Hoekstra, H. E. Molecular spandrels: tests of adaptation at the genetic level. Nat. Rev. Genet. 12, 767–780 (2011).
Martin, A. & Orgogozo, V. The loci of repeated evolution: a catalog of genetic hotspots of phenotypic variation. Evolution 67, 1235–1250 (2013).
Barrett, R. D. H., Rogers, S. M. & Schluter, D. Natural selection on a major armor gene in threespine stickleback. Science 322, 255–257 (2008).
Barrett, R. D. H. et al. Linking a mutation to survival in wild mice. Science 363, 499–504 (2019).
Gratten, J. et al. A localized negative genetic correlation constrains microevolution of coat color in wild sheep. Science 319, 318–320 (2008).
Lamichhaney, S. et al. A beak size locus in Darwin’s finches facilitated character displacement during a drought. Science 352, 470–474 (2016).
Coberly, L. C. & Rausher, M. D. Pleiotropic effects of an allele producing white flowers in Ipomoea purpurea. Evolution 62, 1076–1085 (2008).
Korves, T. M., others. Fitness effects associated with the major flowering time gene FRIGIDA in Arabidopsis thaliana in the field. Am. Nat. 169, 141–157 (2007).
Rockman, M. V. The QTN program and the alleles that matter for evolution: all that’s gold does not glitter. Evolution 66, 1–17 (2012).
de Visser, J. C. F. T. & Elena, S. F. The causes of epistasis. Proc. R. Soc. B 278, 3617–3624 (2011).
Arnegard, M. E. et al. Genetics of ecological divergence during speciation. Nature 511, 307–311 (2014).
Storz, J. F. Causes of molecular convergence and parallelism in protein evolution. Nat. Rev. Genet. 17, 239–250 (2016).
Kryazhimskiy, S., Rice, D. P., Jerison, E. R. & Desai, M. M. Global epistasis makes adaptation predictable despite sequence-level stochasticity. Science 344, 1519–1522 (2014).
Marques, D. A. et al. Experimental evidence for rapid genomic adaptation to a new niche in an adaptive radiation. Nat. Ecol. Evol. 2, 1128–1138 (2018).
Natarajan, C. et al. Epistasis among adaptive mutations in deer mouse hemoglobin. Science 340, 1324–1327 (2013).
Dettman, J. R., Sirjusingh, C., Kohn, L. M. & Anderson, J. B. Incipient speciation by divergent adaptation and antagonistic epistasis in yeast. Nature 447, 585–588 (2007).
Orr, H. A. The population genetics of speciation— the evolution of hybrid incompatibilities. Genetics 139, 1805–1813 (1995).
Gavrilets, S. Evolution and speciation on holey adaptive landscapes. Trends Ecol. Evol. 12, 307–312 (1997).
Schwander, T., Libbrecht, R. & Keller, L. Supergenes and complex phenotypes. Curr. Biol. 24, R288–R294 (2014).
Wilfert, L. & Schmid-Hempel, P. The genetic architecture of susceptibility to parasites. BMC Evol. Biol. 8, 187 (2008).
Weinreich, D. M., Delaney, N. F., DePristo, M. A. & Hartl, D. L. Darwinian evolution can follow only very few mutational paths to fitter proteins. Science 312, 111–114 (2006).
Gavrilets, S. Fitness Landscapes and the Origin of Species (Princeton Univ. Press, 2004); https://doi.org/10.2307/j.ctv39x541
Wright, S. The roles of mutation, inbreeding, crossbreeding, and selection in evolution. Proc. Sixth Int. Congr. Genet. 1, 356–366 (1932).
Lehner, B. Molecular mechanisms of epistasis within and between genes. Trends Genet. 27, 323–331 (2011).
Whitlock, M. C., Phillips, P. C., Moore, F. B. & Tonsor, S. J. Multiple fitness peaks and epistasis. Annu. Rev. Ecol. Syst. 26, 601–629 (1995).
Whitlock, M. C. Founder effects and peak shifts without genetic drift: adaptive peak shifts occur easily when environments fluctuate slightly. Evolution 51, 1044–1048 (1997).
Kingsolver, J. G. et al. The strength of phenotypic selection in natural populations. Am. Nat. 157, 245–261 (2001).
Sinervo, B. & Svensson, E. Correlational selection and the evolution of genomic architecture. Heredity 89, 329–338 (2002).
Poelwijk, F. J., Kiviet, D. J., Weinreich, D. M. & Tans, S. J. Empirical fitness landscapes reveal accessible evolutionary paths. Nature 445, 383–386 (2007).
Plucain, J. et al. Epistasis and allele specificity in the emergence of a stable polymorphism in Escherichia coli. Science 343, 1366–1369 (2014).
Kirkpatrick, M. How and why chromosome inversions evolve. PLoS Biol. 8, e1000501 (2010).
Sandoval, C. P. Differential visual predation on morphs of Timema cristinae (Phasmatodeae:Timemidae) and its consequences for host range. Biol. J. Linn. Soc. 52, 341–356 (1994).
Sandoval, C. P. The effects of the relative geographic scales of gene flow and selection on morph frequencies in the walking‐stick Timema cristinae. Evolution 48, 1866–1879 (1994).
Sandoval, C. P. & Nosil, P. Counteracting selective regimes and host preference evolution in ecotypes of two species of walking-sticks. Evolution 59, 2405–2413 (2005).
Comeault, A. A. et al. Selection on a genetic polymorphism counteracts ecological speciation in a stick insect. Curr. Biol. 25, 1975–1981 (2015).
Nosil, P. et al. Natural selection and the predictability of evolution in Timema stick insects. Science 359, 765–770 (2018).
Villoutreix, R. et al. Large-scale mutation in the evolution of a gene complex for cryptic coloration. Science 369, 460–466 (2020).
Lindtke, D. et al. Long-term balancing selection on chromosomal variants associated with crypsis in a stick insect. Mol. Ecol. 26, 6189–6205 (2017).
Endler, J. A. A framework for analysing colour pattern geometry: adjacent colours. Biol. J. Linn. Soc. 107, 233–253 (2012).
Endler, J. A. On the measurement and classification of colour in studies of animal colour patterns. Biol. J. Linn. Soc. 41, 315–352 (1990).
Hurvich, L. M. Color Vision (Sinauer Associates, 1981).
Gompert, Z. et al. Experimental evidence for ecological selection on genome variation in the wild. Ecol. Lett. 17, 369–379 (2014).
Zhou, X., Carbonetto, P. & Stephens, M. Polygenic modeling with Bayesian sparse linear mixed models. PLoS Genet. 9, e1003264 (2013).
Crawford, L., Zeng, P., Mukherjee, S. & Zhou, X. Detecting epistasis with the marginal epistasis test in genetic mapping studies of quantitative traits. PLoS Genet. 13, e1006869 (2017).
Comeault, A. A., Ferreira, C., Dennis, S., Soria-Carrasco, V. & Nosil, P. Color phenotypes are under similar genetic control in two distantly related species of Timema stick insect. Evolution 70, 1283–1296 (2016).
Nosil, P. & Crespi, B. J. Experimental evidence that predation promotes divergence in adaptive radiation. Proc. Natl Acad. Sci. USA 103, 9090–9095 (2006).
Rennison, D. J., Heilbron, K., Barrett, R. D. H. & Schluter, D. Discriminating selection on lateral plate phenotype and its underlying gene, ectodysplasin, in threespine stickleback. Am. Nat. 185, 150–156 (2015).
Wright, S. The shifting balance theory and macroevolution. Annu. Rev. Genet. 16, 1–19 (1982).
Coyne, J. A., Barton, N. H. & Turelli, M. Perspective: a critique of Sewall Wright’s shifting balance theory of evolution. Evolution 51, 643–671 (1997).
Wade, M. J. & Goodnight, C. J. Perspective: the theories of Fisher and Wright in the context of metapopulations: when nature does many small experiments. Evolution 52, 1537–1553 (1998).
Reimchen, T. E. Predator-induced cyclical changes in lateral plate frequencies of Gasterosteus. Behaviour 132, 1079–1094 (1995).
Coyne, J. A. & Orr, H. A. Speciation (Sinauer Associates, 2004).
Sackman, A. M. & Rokyta, D. R. Additive phenotypes underlie epistasis of fitness effects. Genetics 208, 339–348 (2018).
Knief, U. et al. Epistatic mutations under divergent selection govern phenotypic variation in the crow hybrid zone. Nat. Ecol. Evol. 3, 570–576 (2019).
Hench, K., Vargas, M., Höppner, M. P., McMillan, W. O. & Puebla, O. Inter-chromosomal coupling between vision and pigmentation genes during genomic divergence. Nat. Ecol. Evol. 3, 657–667 (2019).
Lewontin, R. C. The Genetic Basis of Evolutionary Change (Columbia Univ. Press, 1974).
Scheffer, M. Critical Transitions in Nature and Society (Princeton Univ. Press, 2009).
Scheffer, M. et al. Anticipating critical transitions. Science 338, 344–348 (2012).
Parchman, T. L. et al. Genome-wide association genetics of an adaptive trait in lodgepole pine. Mol. Ecol. 21, 2991–3005 (2012).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
Soria-Carrasco, V. et al. Stick insect genomes reveal natural selection’s role in parallel speciation. Science 344, 738–742 (2014).
Guan, Y. & Stephens, M. Bayesian variable selection regression for genome-wide association studies and other large-scale problems. Ann. Appl. Stat. 5, 1780–1815 (2011).
Nosil, P. Reproductive isolation caused by visual predation on migrants between divergent environments. Proc. R. Soc. B 271, 1521–1528 (2004).
Nosil, P. et al. Genomic consequences of multiple speciation processes in a stick insect. Proc. R. Soc. B 279, 5058–5065 (2012).
Sandoval, C. P. Persistence of a walking-stick population (Phasmatoptera: Timematodea) after a wildfire. Southwest. Nat. 45, 123–127 (2000).
Plummer, M. rjags: Bayesian graphical models using MCMC. R package version 4-8 (2018).
Lande, R. & Arnold, S. J. The measurement of selection on correlated characters. Evolution 37, 1210–1226 (1983).
Janzen, F. J. & Stern, H. S. Logistic regression for empirical studies of multivariate selection. Evolution 52, 1564–1571 (1998).
Zeugner, S. & Feldkircher, M. Bayesian model averaging employing fixed and flexible priors: the BMS package for R. J. Stat. Softw. 68, 1–37 (2015).
Zellner, A. in Bayesian Inference and Decision Techniques: Essays in Honor of Bruno de Finetti (eds Goel, P. & Zellner, A.) 233–243 (1986).
Csardi, G. & Nepusz, T. The igraph software package for complex network research. InterJ. Complex Syst. 1695, 1–9 (2006).
Weinberger, E. Correlated and uncorrelated fitness landscapes and how to tell the difference. Biol. Cybern. 63, 325–336 (1990).
Vassilev, V. K., Fogarty, T. C. & Miller, J. F. Information characteristics and the structure of landscapes. Evol. Comput. 8, 31–60 (2000).
Kouyos, R. D. et al. Exploring the complexity of the HIV-1 fitness landscape. PLoS Genet. 8, e1002551–e1002551 (2012).
Malan, K. M. & Engelbrecht, A. P. A survey of techniques for characterising fitness landscapes and some possible ways forward. Inf. Sci. 241, 148–163 (2013).
Kondrashov, D. A. & Kondrashov, F. A. Topological features of rugged fitness landscapes in sequence space. Trends Genet. 31, 24–33 (2015).
Poursoltan, S. & Neumann, F. in Evolutionary Constrained Optimization (eds Datta, R. & Deb, K.) 29–50 (Springer, 2015); https://doi.org/10.1007/978-81-322-2184-5_2
Paten, B. et al. Cactus: algorithms for genome multiple sequence alignment. Genome Res. 21, 1512–1528 (2011).
Hickey, G., Paten, B., Earl, D., Zerbino, D. & Haussler, D. HAL: a hierarchical format for storing and analyzing multiple genome alignments. Bioinformatics 29, 1341–1342 (2013).
Endler, J. A. & Mielke, P. W. Comparing entire colour patterns as birds see them. Biol. J. Linn. Soc. 86, 405–431 (2005).
Acknowledgements
We thank T. Reimchen, M. Joron, L.-M. Chevin and D. Ayala for discussion and comments on previous versions of the manuscript, M. Muschick for help with photography and reflectance measurements and T. Oakley for laboratory space. The support and resources from the Center for High Performance Computing at the University of Utah are gratefully acknowledged, as well as access to the High Performance Computing Facilities, particularly to the Iceberg and ShARC HPC clusters, from the Corporate Information and Computing Services at the University of Sheffield. The work was funded by a grant from the European Research Council (grant no. EE-Dynamics 770826, https://erc.europa.eu/) and a grant from the National Science Foundation of the United States (grant no. NSF DEB 1638768). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
All authors conceived the project and contributed to writing. R.V., C.F.C., T.L.P. and P.N. collected data. Z.G. led data analysis, aided by all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods, Tables 1–3, Figs. 1–10 and references.
Rights and permissions
About this article
Cite this article
Nosil, P., Villoutreix, R., de Carvalho, C.F. et al. Ecology shapes epistasis in a genotype–phenotype–fitness map for stick insect colour. Nat Ecol Evol 4, 1673–1684 (2020). https://doi.org/10.1038/s41559-020-01305-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-020-01305-y
This article is cited by
-
Climatic similarity and genomic background shape the extent of parallel adaptation in Timema stick insects
Nature Ecology & Evolution (2022)
-
Correlational selection in the age of genomics
Nature Ecology & Evolution (2021)
-
Disparate genetic variants associated with distinct components of cowpea resistance to the seed beetle Callosobruchus maculatus
Theoretical and Applied Genetics (2021)
-
Increasing our ability to predict contemporary evolution
Nature Communications (2020)