Abstract
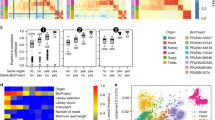
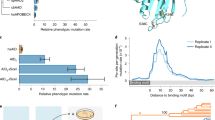
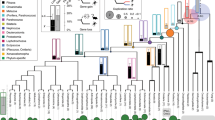
The level of conservation of ancient metazoan gene order (synteny) is remarkable. Despite this, the functionality of the vast majority of such regions in metazoan genomes remains elusive. Utilizing recently published single-cell expression data from several anciently diverging metazoan species, we reveal the level of correspondence between cell types and genomic synteny, identifying genomic regions conferring ancient cell type identity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
Single-cell datasets used in this study are available via gene expression omnibus accessions GSE95723 and GSE111068 and https://shiny.mdc-berlin.de/psca/.
Code availability
The analysis code is available at https://github.com/nijibabulu/metazoan_synteny.
References
Engström, P. G., Ho Sui, S. J., Drivenes, Ø., Becker, T. S. & Lenhard, B. Genome Res. 17, 1898–1908 (2007).
Irimia, M. et al. Genome Res. 22, 2356–2367 (2012).
Simakov, O. et al. Nature 493, 526–531 (2013).
Plass, M. et al. Science 360, eaaq1723 (2018).
Sebé-Pedrós, A. et al. Nat. Ecol. Evol. 2, 1176–1188 (2018).
Sebé-Pedrós, A. et al. Cell 173, 1520–1534.e20 (2018).
Emms, D. M. & Kelly, S. Genome Biol. 16, 157 (2015).
Kharchenko, P. V., Silberstein, L. & Scadden, D. T. Nat. Methods 11, 740–742 (2014).
Khan, A. et al. Nucleic Acids Res. 46, D260–D266 (2018).
Ghazanfar, S., Bisogni, A. J., Ormerod, J. T., Lin, D. M. & Yang, J. Y. H. BMC Syst. Biol. 10, 127 (2016).
Kersey, P. J. et al. Nucleic Acids Res. 46, D802–D808 (2018).
Ryan, J. F. et al. Science 342, 1242592 (2013).
Srivastava, M. et al. Nature 454, 955–960 (2008).
Putnam, N. H. et al. Science 317, 86–94 (2007).
Fredman, D., Schwaiger, M., Rentzsch, F. & Technau, U. Nematostella vectensis transcriptome and gene models v2.0. Figshare https://doi.org/10.6084/m9.figshare.807696.v1 (2013).
Brandl, H. et al. Nucleic Acids Res. 44, D764–D773 (2016).
Tang, S., Lomsadze, A. & Borodovsky, M. Nucleic Acids Res. 43, e78 (2015).
Lefort, V., Desper, R. & Gascuel, O. Mol. Biol. Evol. 32, 2798–2800 (2015).
Alexander, R. A. Bull. Psychon. Soc. 28, 335–336 (1990).
McLeay, R. C. & Bailey, T. L. BMC Bioinformatics 11, 165 (2010).
Yanai, I. et al. Bioinformatics 21, 650–659 (2005).
Acknowledgements
Computation was done at the Life Science Compute Cluster at the University of Vienna. This work was funded by grants from the Austrian Science Fund to U.T. (P27353) and O.S. (P32190).
Author information
Authors and Affiliations
Contributions
O.S. and B.Z. designed the analyses. B.Z. performed the synteny and expression analyses. B.Z., O.S. and N.S.M.R. analysed the data. B.Z., U.T. and O.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8.
Supplementary Data 1
Synteny blocks.
Supplementary Data 2
Expression data for 32 shared microsyntenic blocks.
Supplementary Data 3
High-resolution vector images for Supplementary Figs. 3–7.
Rights and permissions
About this article
Cite this article
Zimmermann, B., Robert, N.S.M., Technau, U. et al. Ancient animal genome architecture reflects cell type identities. Nat Ecol Evol 3, 1289–1293 (2019). https://doi.org/10.1038/s41559-019-0946-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-019-0946-7
This article is cited by
-
Pan-evolutionary and regulatory genome architecture delineated by an integrated macro- and microsynteny approach
Nature Protocols (2024)
-
Emergence of distinct syntenic density regimes is associated with early metazoan genomic transitions
BMC Genomics (2022)
-
Emergence of novel cephalopod gene regulation and expression through large-scale genome reorganization
Nature Communications (2022)
-
Animal cell type diversity
Nature Ecology & Evolution (2019)