Abstract
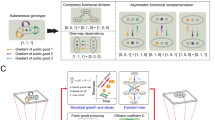
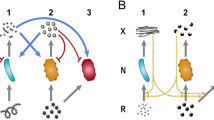
Microbes commonly deploy a risky strategy to acquire nutrients from their environment, involving the production of costly public goods that can be exploited by neighbouring individuals. Why engage in such a strategy when an exploitation-free alternative is readily available whereby public goods are kept private? We address this by examining metabolism of Saccharomyces cerevisiae in its native form and by creating a new three-strain synthetic community deploying different strategies of sucrose metabolism. Public-metabolizers digest resources externally, private-metabolizers internalize resources before digestion, and cheats avoid the metabolic costs of digestion but exploit external products generated by competitors. A combination of mathematical modelling and ecological experiments reveal that private-metabolizers invade and take over an otherwise stable community of public-metabolizers and cheats. However, owing to the reduced growth rate of private-metabolizers and population bottlenecks that are frequently associated with microbial communities, privatizing public goods can become unsustainable, leading to population decline.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The research data supporting this publication can be found at https://doi.org/10.24378/exe.1383.
References
Richards, T. A. & Talbot, N. J. Horizontal gene transfer in osmotrophs: playing with public goods. Nat. Rev. Microbiol. 11, 720–727 (2013).
Drescher, K., Nadell, C. D., Stone, H. A., Wingreen, N. S. & Bassler, B. L. Solutions to the public goods dilemma in bacterial biofilms. Curr. Biol. 24, 50–55 (2014).
Lindsay, R. J., Kershaw, M. J., Pawlowska, B. J., Talbot, N. J. & Gudelj, I. Harbouring public good mutants within a pathogen population can increase both fitness and virulence. eLife 5, e18678 (2016).
Bachmann, H., Molenaar, D., Kleerebezem, M. & van Hylckama Vlieg, J. E. High local substrate availability stabilizes a cooperative trait. ISME J. 5, 929–932 (2010).
Griffin, A. S., West, S. A. & Buckling, A. Cooperation and competition in pathogenic bacteria. Nature 430, 1024–1027 (2004).
Chang, Q. et al. A unique invertase is important for sugar absorption of an obligate biotrophic pathogen during infection. New Phytol. 215, 1548–1561 (2017).
Lincoln, L. & More, S. S. Bacterial invertases: occurrence, production, biochemical characterization, and significance of transfructosylation. J. Basic Microbiol. 57, 803–813 (2017).
Parrent, J. L., James, T. Y., Vasaitis, R. & Taylor, A. F. Friend or foe? Evolutionary history of glycoside hydrolase family 32 genes encoding for sucrolytic activity in fungi and its implications for plant–fungal symbioses. BMC Evol. Biol. 9, 148 (2009).
Voegele, R. T., Wirsel, S., Möll, U., Lechner, M. & Mendgen, K. Cloning and characterization of a novel invertase from the obligate biotroph Uromyces fabae and analysis of expression patterns of host and pathogen invertases in the course of infection. Mol. Plant Microbe Interact. 19, 625–634 (2006).
Koschwanez, J. H., Foster, K. R. & Murray, A. W. Improved use of a public good selects for the evolution of undifferentiated multicellularity. eLife 2, e00367 (2013).
Dai, L., Vorselen, D., Korolev, K. S. & Gore, J. Generic indicators for loss of resilience before a tipping point leading to population collapse. Science 336, 1175–1177 (2012).
Völker, C. & Wolf-Gladrow, D. A. Physical limits on iron uptake mediated by siderophores or surface reductases. Mar. Chem. 65, 227–244 (1999).
Greig, D. & Travisano, M. The Prisoner’s Dilemma and polymorphism in yeast SUC genes. Proc. Biol. Sci. 271(Suppl. 3), S25–S26 (2004).
Gore, J., Youk, H. & van Oudenaarden, A. Snowdrift game dynamics and facultative cheating in yeast. Nature 459, 253–256 (2009).
Harrison, F., Browning, L. E., Vos, M. & Buckling, A. Cooperation and virulence in acute Pseudomonas aeruginosa infections. BMC Biol. 4, 21 (2006).
Kümmerli, R., Schiessl, K. T., Waldvogel, T., McNeill, K. & Ackermann, M. Habitat structure and the evolution of diffusible siderophores in bacteria. Ecol. Lett. 17, 1536–1544 (2014).
Niehus, R., Picot, A., Oliveira, N. M., Mitri, S. & Foster, K. R. The evolution of siderophore production as a competitive trait. Evolution 71, 1443–1455 (2017).
Lee, W., Van Baalen, M. & Jansen, V. A. An evolutionary mechanism for diversity in siderophore‐producing bacteria. Ecol. Lett. 15, 119–125 (2012).
Riley, M. A. & Wertz, J. E. Bacteriocins: evolution, ecology, and application. Annu Rev. Microbiol. 56, 117–137 (2002).
Celiker, H. & Gore, J. Competition between species can stabilize public-goods cooperation within a species. Mol. Syst. Biol. 8, 621 (2012).
Verstrepen, K. J. et al. Glucose and sucrose: hazardous fast-food for industrial yeast? Trends Biotechnol. 22, 531–537 (2004).
Lindsay, R. J., Pawlowska, B. J. & Gudelj, I. When increasing population density can promote the evolution of metabolic cooperation. ISME J. 12, 849–859 (2018).
Ross-Gillespie, A., Gardner, A., West, S. A. & Griffin, A. S. Frequency dependence and cooperation: theory and a test with bacteria. Am. Nat. 170, 331–342 (2007).
Ross-Gillespie, A., Gardner, A., Buckling, A., West, S. A. & Griffin, A. S. Density dependence and cooperation: theory and a test with bacteria. Evolution 63, 2315–2325 (2009).
Kümmerli, R., Griffin, A. S., West, S. A., Buckling, A. & Harrison, F. Viscous medium promotes cooperation in the pathogenic bacterium Pseudomonas aeruginosa. Proc. Biol. Sci. 276, 3531–3538 (2009).
Allison, S. D. Cheaters, diffusion and nutrients constrain decomposition by microbial enzymes in spatially structured environments. Ecol. Lett. 8, 626–635 (2005).
Julou, T. et al. Cell–cell contacts confine public goods diffusion inside Pseudomonas aeruginosa clonal microcolonies. Proc. Natl Acad. Sci. USA 110, 12577–12582 (2013).
MacLean, R. C., Fuentes-Hernandez, A., Greig, D., Hurst, L. D. & Gudelj, I. A mixture of ‘cheats’ and ‘co-operators’ can enable maximal group benefit. PLoS Biol. 8, e1000486 (2010).
Driscoll, W. W., Pepper, J. W., Pierson, L. S. & Pierson, E. A. Spontaneous Gac mutants of Pseudomonas biological control strains: cheaters or mutualists? Appl. Environ. Microbiol. 77, 7227–7235 (2011).
Rainey, P. B. & Rainey, K. Evolution of cooperation and conflict in experimental bacterial populations. Nature 425, 72 (2003).
Sanchez, A. & Gore, J. Feedback between population and evolutionary dynamics determines the fate of social microbial populations. PLoS Biol. 11, e1001547 (2013).
Rakoff-Nahoum, S., Foster, K. R. & Comstock, L. E. The evolution of cooperation within the gut microbiota. Nature 533, 255–259 (2016).
Galeote, V. et al. FSY1, a horizontally transferred gene in the Saccharomyces cerevisiae EC1118 wine yeast strain, encodes a high-affinity fructose/H+ symporter. Microbiology 156, 3754–3761 (2010).
Brown, C. J., Todd, K. M. & Rosenzweig, R. F. Multiple duplications of yeast hexose transport genes in response to selection in a glucose-limited environment. Mol. Biol. Evol. 15, 931–942 (1998).
Wahl, R., Wippel, K., Goos, S., Kämper, J. & Sauer, N. A novel high-affinity sucrose transporter is required for virulence of the plant pathogen Ustilago maydis. PLoS Biol. 8, e1000303 (2010).
Cuskin, F. et al. Human gut Bacteroidetes can utilize yeast mannan through a selfish mechanism. Nature 517, 165–169 (2015).
Martens, E. C., Koropatkin, N. M., Smith, T. J. & Gordon, J. I. Complex glycan catabolism by the human gut microbiota: the Bacteroidetes Sus-like paradigm. J. Biol. Chem. 284, 24673–24677 (2009).
Reintjes, G., Arnosti, C., Fuchs, B. M. & Amann, R. An alternative polysaccharide uptake mechanism of marine bacteria. ISME J. 11, 1640 (2017).
Dandekar, A. A., Chugani, S. & Greenberg, E. P. Bacterial quorum sensing and metabolic incentives to cooperate. Science 338, 264–266 (2012).
Jin, Z. et al. Conditional privatization of a public siderophore enables Pseudomonas aeruginosa to resist cheater invasion. Nat. Commun. 9, 1383 (2018).
Zhang, X. X. & Rainey, P. B. Exploring the sociobiology of pyoverdin‐producing Pseudomonas. Evolution 67, 3161–3174 (2013).
Pande, S. et al. Privatization of cooperative benefits stabilizes mutualistic cross-feeding interactions in spatially structured environments. ISME J. 10, 1413 (2016).
Inglis, R. F., Biernaskie, J. M., Gardner, A. & Kümmerli, R. Presence of a loner strain maintains cooperation and diversity in well-mixed bacterial communities. Proc. Biol. Sci. 283, https://doi.org/10.1098/rspb.2015.2682 (2016).
Schweizer, M. & Dickinson, J. R. The Metabolism and Molecular Physiology of Saccharomyces cerevisiae (CRC Press, 2004).
Zhou, L., Slamti, L., Nielsen-LeRoux, C., Lereclus, D. & Raymond, B. The social biology of quorum sensing in a naturalistic host pathogen system. Curr. Biol. 24, 2417–2422 (2014).
Travisano, M. & Velicer, G. J. Strategies of microbial cheater control. Trends Microbiol. 12, 72–78 (2004).
Brockhurst, M. A., Buckling, A. & Gardner, A. Cooperation peaks at intermediate disturbance. Curr. Biol. 17, 761–765 (2007).
Rouwenhorst, R. J., Van Der Baan, A. A., Scheffers, W. A. & Van Dijken, J. Production and localization of beta-fructosidase in asynchronous and synchronous chemostat cultures of yeasts. Appl. Environ. Microbiol. 57, 557–562 (1991).
Tammi, M., Ballou, L., Taylor, A. & Ballou, C. Effect of glycosylation on yeast invertase oligomer stability. J. Biol. Chem. 262, 4395–4401 (1987).
Carlson, M. & Botstein, D. Two differentially regulated mRNAs with different 5′ ends encode secreted and intracellular forms of yeast invertase. Cell 28, 145–154 (1982).
Williams, R. S., Trumbly, R. J., MacColl, R., Trimble, R. B. & Maley, F. Comparative properties of amplified external and internal invertase from the yeast SUC2 gene. J. Biol. Chem. 260, 13334–13341 (1985).
Gascón, S., Neumann, N. P. & Lampen, J. O. Comparative study of the properties of the purified internal and external invertases from yeast. J. Biol. Chem. 243, 1573–1577 (1968).
Kaiser, C. A. & Botstein, D. Secretion-defective mutations in the signal sequence for Saccharomyces cerevisiae invertase. Mol. Cell. Biol. 6, 2382–2391 (1986).
Stambuk, B. U., Batista, A. S. & De Araujo, P. S. Kinetics of active sucrose transport in Saccharomyces cerevisiae. J. Biosci. Bioeng. 89, 212–214 (2000).
Stambuk, B. U., da Silva, M. A., Panek, A. D. & de Araujo, P. S. Active α-glucoside transport in Saccharomyces cerevisiae. FEMS Microbiol. Lett. 170, 105–110 (1999).
Vidgren, V., Ruohonen, L. & Londesborough, J. Characterization and functional analysis of the MAL and MPH loci for maltose utilization in some ale and lager yeast strains. Appl. Environ. Microbiol. 71, 7846–7857 (2005).
Lion, S. Theoretical approaches in evolutionary ecology: environmental feedback as a unifying perspective. Am. Nat. 191, 21–44 (2017).
McPeek, M. A. The ecological dynamics of natural selection: traits and the coevolution of community structure. Am. Nat. 189, E91–E117 (2017).
Otterstedt, K. et al. Switching the mode of metabolism in the yeast Saccharomyces cerevisiae. EMBO Rep. 5, 532–537 (2004).
Sauer, N. & Stolz, J. SUC1 and SUC2: two sucrose transporters from Arabidopsis thaliana; expression and characterization in baker’s yeast and identification of the histidine‐tagged protein. Plant J. 6, 67–77 (1994).
Weise, A. et al. A new subfamily of sucrose transporters, SUT4, with low affinity/high capacity localized in enucleate sieve elements of plants. Plant Cell 12, 1345–1355 (2000).
Reinders, A. & Ward, J. M. Functional characterization of the α‐glucoside transporter Sut1p from Schizosaccharomyces pombe, the first fungal homologue of plant sucrose transporters. Mol. Microbiol. 39, 445–455 (2001).
Elbing, K. et al. Role of hexose transport in control of glycolytic flux in Saccharomyces cerevisiae. Appl. Environ. Microbiol. 70, 5323–5330 (2004).
Bisson, L. F., Coons, D. M., Kruckeberg, A. L. & Lewis, D. A. Yeast sugar transporters. Crit. Rev. Biochem. Mol. Biol. 28, 259–308 (1993).
Özcan, S. & Johnston, M. Function and regulation of yeast hexose transporters. Microbiol. Mol. Biol. Rev. 63, 554–569 (1999).
Leventhal, G. E., Ackermann, M. & Schiessl, K. T. Why microbes secrete molecules to modify their environment: the case of iron-chelating siderophores. J. R. Soc. Interface 16, 20180674 (2019).
Andersen, S. B. et al. Privatisation rescues function following loss of cooperation. eLife 7, e38594 (2018).
Vetter, Y., Deming, J., Jumars, P. & Krieger-Brockett, B. A predictive model of bacterial foraging by means of freely released extracellular enzymes. Microb. Ecol. 36, 75–92 (1998).
Dean, R. et al. The top 10 fungal pathogens in molecular plant pathology. Mol. Plant Pathol. 13, 414–430 (2012).
Kothary, M. H. & Babu, U. S. Infective dose of foodborne pathogens in volunteers: a review. J. Food Safety 21, 49–68 (2001).
Schmid-Hempel, P. & Frank, S. A. Pathogenesis, virulence, and infective dose. PLoS Pathog. 3, e147 (2007).
Piskur, J., Rozpedowska, E., Polakova, S., Merico, A. & Compagno, C. How did Saccharomyces evolve to become a good brewer? Trends Genet. 22, 183–186 (2006).
Sikorski, R. S. & Hieter, P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122, 19–27 (1989).
Reding-Roman, C. et al. The unconstrained evolution of fast and efficient antibiotic-resistant bacterial genomes. Nat. Ecol. Evol. 1, 0050 (2017).
Lenski, R. E., Rose, M. R., Simpson, S. C. & Tadler, S. C. Long-term experimental evolution in Escherichia coli. I. Adaptation and divergence during 2,000 generations. Am. Nat. 138, 1315–1341 (1991).
Pfeiffer, T., Schuster, S. & Bonhoeffer, S. Cooperation and competition in the evolution of ATP-producing pathways. Science 292, 504 (2001).
Bauchop, T. & Elsden, S. R. The growth of micro-organisms in relation to their energy supply. J. Gen. Microbiol. 23, 457–469 (1960).
Postma, E., Verduyn, C., Scheffers, W. A. & Van Dijken, J. P. Enzymic analysis of the crabtree effect in glucose-limited chemostat cultures of Saccharomyces cerevisiae. Appl Environ. Microbiol 55, 468–477 (1989).
Weusthuis, R. A., Pronk, J. T., van den Broek, P. J. & van Dijken, J. P. Chemostat cultivation as a tool for studies on sugar transport in yeasts. Microbiol. Rev. 58, 616–630 (1994).
Badotti, F. et al. Switching the mode of sucrose utilization by Saccharomyces cerevisiae. Microb. Cell Fact. 7, 4 (2008).
Acknowledgements
We thank R. Beardmore, A. Jepson and P. Holder for comments and helpful discussions. R.J.L. and I.G. are funded by European Research Council Consolidator Grant No. 647292 MathModExp awarded to I.G., and B.J.P. was funded by an EPSRC Doctoral training studentship.
Author information
Authors and Affiliations
Contributions
R.J.L. and I.G. conceived the idea. R.J.L. and I.G designed the experiments. R.J.L. carried out the experiments. B.J.P. and I.G. developed the mathematical model. B.J.P. carried out numerical simulations. R.J.L. and I.G. analysed the results and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–14 and Table 1.
Rights and permissions
About this article
Cite this article
Lindsay, R.J., Pawlowska, B.J. & Gudelj, I. Privatization of public goods can cause population decline. Nat Ecol Evol 3, 1206–1216 (2019). https://doi.org/10.1038/s41559-019-0944-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-019-0944-9
This article is cited by
-
Universality of dissipative self-assembly from quantum dots to human cells
Nature Physics (2020)