Abstract
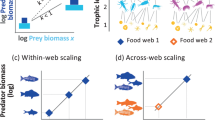
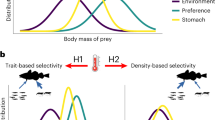
Predator–prey interactions in natural ecosystems generate complex food webs that have a simple universal body-size architecture where predators are systematically larger than their prey. Food-web theory shows that the highest predator–prey body-mass ratios found in natural food webs may be especially important because they create weak interactions with slow dynamics that stabilize communities against perturbations and maintain ecosystem functioning. Identifying these vital interactions in real communities typically requires arduous identification of interactions in complex food webs. Here, we overcome this obstacle by developing predator-trait models to predict average body-mass ratios based on a database comprising 290 food webs from freshwater, marine and terrestrial ecosystems across all continents. We analysed how species traits constrain body-size architecture by changing the slope of the predator–prey body-mass scaling. Across ecosystems, we found high body-mass ratios for predator groups with specific trait combinations including (1) small vertebrates and (2) large swimming or flying predators. Including the metabolic and movement types of predators increased the accuracy of predicting which species are engaged in high body-mass ratio interactions. We demonstrate that species traits explain striking patterns in the body-size architecture of natural food webs that underpin the stability and functioning of ecosystems, paving the way for community-level management of the most complex natural ecosystems.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data supporting the findings of this study (GATEWAy 1.0) are available at the iDiv data repository41.
Code availability
The R code of the statistical analyses is available as a Supplement.
References
Brose, U. et al. Consumer-resource body-size relationships in natural food webs. Ecology 87, 2411–2417 (2006).
Barnes, C., Maxwell, D., Reuman, D. C. & Jennings, S. Global patterns in predator–prey size relationships reveal size dependency of trophic transfer efficiency. Ecology 91, 222–232 (2010).
Nakazawa, T., Ohba, S. & Ushio, M. Predator–prey body size relationships when predators can consume prey larger than themselves. Biol. Lett. 9, 20121193 (2013).
Woodward, G. et al. Body size in ecological networks. Trends Ecol. Evol. 20, 402–409 (2005).
Petchey, O. L., Beckerman, A. P., Riede, J. O. & Warren, P. H. Size, foraging, and food web structure. Proc. Natl Acad. Sci. USA 105, 4191–4196 (2008).
Eklöf, A. et al. The dimensionality of ecological networks. Ecol. Lett. 16, 577–583 (2013).
Rall, B., Kalinkat, G., Ott, D., Vucic-Pestic, O. & Brose, U. Taxonomic versus allometric constraints on non-linear interaction strengths. Oikos 120, 483–492 (2011).
Emmerson, M. C. & Raffaelli, D. Predator–prey body size, interaction strength and the stability of a real food web. J. Anim. Ecol. 73, 399–409 (2004).
Reuman, D. C. & Cohen, J. E. Estimating relative energy fluxes using the food web, species abundance, and body size. Adv. Ecol. Res. 36, 137–182 (2005).
Schneider, F. D., Scheu, S. & Brose, U. Body mass constraints on feeding rates determine the consequences of predator loss. Ecol. Lett. 15, 436–443 (2012).
Brose, U. et al. Foraging theory predicts predator–prey energy fluxes. J. Anim. Ecol. 77, 1072–1078 (2008).
McCann, K., Hastings, A. & Huxel, G. R. Weak trophic interactions and the balance of nature. Nature 395, 794–798 (1998).
Brose, U., Williams, R. J. & Martinez, N. D. Allometric scaling enhances stability in complex food webs. Ecol. Lett. 9, 1228–1236 (2006).
Rooney, N., McCann, K., Gellner, G. & Moore, J. C. Structural asymmetry and the stability of diverse food webs. Nature 442, 265–269 (2006).
Otto, S. B., Rall, B. C. & Brose, U. Allometric degree distributions facilitate food-web stability. Nature 450, 1226–1229 (2007).
Blanchard, J. L., Law, R., Castle, M. D. & Jennings, S. Coupled energy pathways and the resilience of size-structured food webs. Theor. Ecol. 4, 289–300 (2011).
Schneider, F. D., Brose, U., Rall, B. C. & Guill, C. Animal diversity and ecosystem functioning in dynamic food webs. Nat. Commun. 7, 12718 (2016).
Wang, S. & Brose, U. Biodiversity and ecosystem functioning in food webs: the vertical diversity hypothesis. Ecol. Lett. 21, 9–20 (2018).
Binzer, A., Guill, C., Rall, B. C. & Brose, U. Interactive effects of warming, eutrophication and size structure: impacts on biodiversity and food-web structure. Glob. Change Biol. 22, 220–227 (2016).
Rall, B. C., Guill, C. & Brose, U. Food-web connectance and predator interference dampen the paradox of enrichment. Oikos 117, 202–213 (2008).
Brose, U. et al. Predicting the consequences of species loss using size-structured biodiversity approaches. Biol. Rev. Camb. Philos. Soc. 92, 684–697 (2017).
Cohen, J. E., Pimm, S. L., Yodzis, P. & Saldaña, J. Body sizes of animal predators and animal prey in food webs. J. Anim. Ecol. 62, 67–78 (1993).
Riede, J. O. et al. Stepping in Elton’s footprints: a general scaling model for body masses and trophic levels across ecosystems. Ecol. Lett. 14, 169–178 (2011).
Carbone, C., Codron, D., Scofield, C., Clauss, M. & Bielby, J. Geometric factors influencing the diet of vertebrate predators in marine and terrestrial environments. Ecol. Lett. 17, 1553–1559 (2014).
Naisbit, R. E., Kehrli, P., Rohr, R. P. & Bersier, L.-F. Phylogenetic signal in predator–prey body-size relationships. Ecology 92, 2183–2189 (2011).
Costa-Pereira, R., Araújo, M. S., Olivier, R., Souza, F. L. & Rudolf, V. H. W. Prey limitation drives variation in allometric scaling of predator–prey interactions. Am. Nat. 192, E139–E149 (2018).
Hirt, M. R., Jetz, W., Rall, B. C. & Brose, U. A general scaling law reveals why the largest animals are not the fastest. Nat. Ecol. Evol. 1, 1116–1122 (2017).
Pawar, S., Dell, A. I. & Savage, V. M. Dimensionality of consumer search space drives trophic interaction strengths. Nature 486, 485–489 (2012).
Digel, C., Curtsdotter, A., Riede, J., Klarner, B. & Brose, U. Unravelling the complex structure of forest soil food webs: higher omnivory and more trophic levels. Oikos 123, 1157–1172 (2014).
Stan Development Team. RStan: the R interface to Stan. R package v.2.14.2 (2016).
Warton, D. I., Wright, I. J., Falster, D. S. & Westoby, M. Bivariate line-fitting methods for allometry. Biol. Rev. Camb. Philos. Soc. 81, 259–291 (2006).
Laigle, I. et al. Species traits as drivers of food web structure. Oikos 127, 316–326 (2018).
Tucker, M. A. & Rogers, T. L. Examining predator–prey body size, trophic level and body mass across marine and terrestrial mammals. Proc. Biol. Sci. 281, 20142103 (2014).
Ings, T. C. et al. Ecological networks: beyond food webs. J. Anim. Ecol. 78, 253–269 (2009).
Nakazawa, T., Ushio, M. & Kondoh, M. Scale dependence of predator–prey mass ratio: determinants and applications. Adv. Ecol. Res. 45, 269–302 (2011).
Wood, S. A., Russell, R., Hanson, D., Williams, R. J. & Dunne, J. A. Effects of spatial scale of sampling on food web structure. Ecol. Evol. 5, 3769–3782 (2015).
Dobashi, T., Iida, M. & Takemoto, K. Decomposing the effects of ocean environments on predator–prey body-size relationships in food webs. R. Soc. Open Sci. 5, 180707 (2018).
Gibert, J. P. & DeLong, J. P. Temperature alters food web body-size structure. Biol. Lett. 10, 20140473 (2014).
Lafferty, K. D., Dobson, A. P. & Kuris, A. M. Parasites dominate food web links. Proc. Natl Acad. Sci. USA 103, 11211–11216 (2006).
LaffertyMarcogliese, K. D. et al. Parasites in food webs: the ultimate missing links. Ecol. Lett. 11, 533–546 (2008).
Brose, U. et al. (2018) GlobAL daTabasE of traits and food Web Architecture (GATEWAy) v.1.0. (iDiv Data Repository, accessed 17 April 2019); https://doi.org/10.25829/iDiv.283-3-756
Acknowledgements
This study was supported by the German Centre for integrative Biodiversity Research (iDiv) Halle-Jena-Leipzig funded by the German Research Foundation (grant no. FZT 118). R.T. was supported by an Australian Research Council Future Fellowship (no. FT110100957). A.C.I. was supported by the Alexander von Humboldt Foundation (grant ID 1156434). C.V. acknowledges a researcher position and strategic project (no. UID/MAR/04292/2013), funded by the Portuguese Science Foundation. We thank L. Rohde, F. Schwarzmüller and A. Dell for help in organizing a prior version of the database.
Author information
Authors and Affiliations
Contributions
U.B. developed the study design. U.B., P.A., A.D.B., L.-F.B., T.B., J.C.-C., E.C., M.D., C.D., A.D., A.A.V.F., K.F., B.G., C.G., J.H., M.R.H., U.J., M.J., S.K., O.M., M.M.M., E.L., K.L.-D., P.L., Y.L., C.M., N.D.M., V.M., C.M., S.A.N., E.J.O., D.O., J.P., D. Perkins, D. Piechnik, I.P., D.R., B.C.R., B.R., R.R., A.S., E.H.S., N.S., M.S.A.T., R.M.T., F.V., C.V., S.W., J.M.W., R.J.W., E.W., G.W. and A.C.I. gathered, contributed or organized data. U.B. and B.R. carried out statistical analyses. M.R.H. created the figures. U.B. and A.C.I. wrote the first draft of the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8, Supplementary Tables 1–4, Supplementary Statistical Methods, Supplementary Metadata and Supplementary References
Supplementary Code
R code of the statistical analysis
Rights and permissions
About this article
Cite this article
Brose, U., Archambault, P., Barnes, A.D. et al. Predator traits determine food-web architecture across ecosystems. Nat Ecol Evol 3, 919–927 (2019). https://doi.org/10.1038/s41559-019-0899-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-019-0899-x
This article is cited by
-
Rainforest transformation reallocates energy from green to brown food webs
Nature (2024)
-
The Pelagic Species Trait Database, an open data resource to support trait-based ocean research
Scientific Data (2024)
-
Flexible foraging behaviour increases predator vulnerability to climate change
Nature Climate Change (2024)
-
Modeling time-varying phytoplankton subsidy reveals at-risk species in a Chilean intertidal ecosystem
Scientific Reports (2024)
-
Human pressures modulate climate-warming-induced changes in size spectra of stream fish communities
Nature Ecology & Evolution (2023)