Abstract
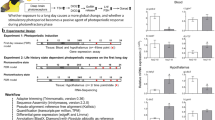
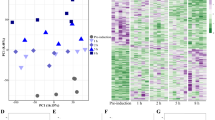
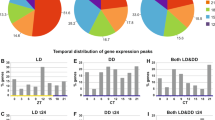
To cope with seasonal environmental changes, animals adapt their physiology and behaviour in response to photoperiod. However, the molecular mechanisms underlying these adaptive changes are not completely understood. Here, using genome-wide expression analysis, we show that an uncharacterized long noncoding RNA (lncRNA), LDAIR, is strongly regulated by photoperiod in Japanese medaka fish (Oryzias latipes). Numerous transcripts and signalling pathways are activated during the transition from short- to long-day conditions; however, LDAIR is one of the first genes to be induced and its expression shows a robust daily rhythm under long-day conditions. Transcriptome analysis of LDAIR knockout fish reveals that the LDAIR locus regulates a gene neighbourhood, including corticotropin releasing hormone receptor 2, which is involved in the stress response. Behavioural analysis of LDAIR knockout fish demonstrates that LDAIR affects self-protective behaviours under long-day conditions. Therefore, we propose that photoperiodic regulation of corticotropin releasing hormone receptor 2 by LDAIR modulates adaptive behaviours to seasonal environmental changes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The microarray and RNA-Seq data are available at the NCBI Gene Expression Omnibus (accession no. GSE119905). All other data are available from the authors upon request.
References
Romero, L. M. Seasonal changes in plasma glucocorticoid concentrations in free-living vertebrates. Gen. Comp. Endocrinol. 128, 1–24 (2002).
Rowan, W. Relation of light to bird migration and developmental changes. Nature 115, 494–495 (1925).
Follett, B. K. & Sharp, P. J. Circadian rhythmicity in photoperiodically induced gonadotrophin release and gonadal growth in the quail. Nature 223, 968–971 (1969).
Egami, N. Effect of artificial photoperiodicity on time of oviposition in the fish, Oryzias latipes. Annot. Zool. Jpn 27, 57–62 (1954).
Awaji, M. & Hanyu, I. Temperature-photoperiod conditions necessary to begin the spawning season in wild type medaka. Nippon Suisan Gakk. 55, 747 (1989).
Kasahara, M. et al. The medaka draft genome and insights into vertebrate genome evolution. Nature 447, 714–719 (2007).
Ansai, S. & Kinoshita, M.Targeted mutagenesis using CRISPR/Cas system in medaka. Biol. Open 3, 362–371 (2014).
Shimmura, T. et al. Dynamic plasticity in phototransduction regulates seasonal changes in color perception. Nat. Commun. 8, 412 (2017).
Ichikawa, K. et al. Centromere evolution and CpG methylation during vertebrate speciation. Nat. Commun. 8, 1833 (2017).
Bale, T. L. et al. Mice deficient for corticotropin-releasing hormone receptor-2 display anxiety-like behaviour and are hypersensitive to stress. Nat. Genet. 24, 410–414 (2000).
Kishimoto, T. et al. Deletion of Crhr2 reveals an anxiolytic role for corticotropin-releasing hormone receptor-2. Nat. Genet. 24, 415–419 (2000).
Heo, J. B. & Sung, S. Vernalization-mediated epigenetic silencing by a long intronic noncoding RNA. Science 331, 76–79 (2011).
Salameh, A. et al. PRUNE2 is a human prostate cancer suppressor regulated by the intronic long noncoding RNA PCA3. Proc. Natl Acad. Sci. USA 112, 8403–8408 (2015).
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. S. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
Hezroni, H. et al. Principles of long noncoding RNA evolution derived from direct comparison of transcriptomes in 17 species. Cell Rep. 11, 1110–1122 (2015).
Engreitz, J. M. et al. Local regulation of gene expression by lncRNA promoters, transcription and splicing. Nature 539, 452–455 (2016).
Joung, J. et al. Genome-scale activation screen identifies a lncRNA locus regulating a gene neighbourhood. Nature 548, 343–346 (2017).
Werner, M. S. et al. Chromatin-enriched lncRNAs can act as cell-type specific activators of proximal gene transcription. Nat. Struct. Mol. Biol. 24, 596–603 (2017).
Moore, I. T. & Jessop, T. S. Stress, reproduction, and adrenocortical modulation in amphibians and reptiles. Horm. Behav. 43, 39–47 (2003).
Wingfield, J. C. et al. Ecological bases of hormone–behavior interactions: the ‘emergency life history stage'. Am. Zool. 38, 191–206 (1998).
Balm, P. H. M. Stress Physiology in Animals (Blackwell, 1999).
Kysil, E. V. et al. Comparative analyses of zebrafish anxiety-like behavior using conflict-based novelty tests. Zebrafish 14, 197–208 (2017).
Goritz, C., Mauch, D. H. & Pfrieger, F. W. Multiple mechanisms mediate cholesterol-induced synaptogenesis in a CNS neuron. Mol. Cell. Neurosci. 29, 190–201 (2005).
Fester, L. et al. Cholesterol-promoted synaptogenesis requires the conversion of cholesterol to estradiol in the hippocampus. Hippocampus 19, 692–705 (2009).
Posse de Chaves, E. I., Rusiñol, A. E., Vance, D. E., Campenot, R. B. & Vance, J. E. Role of lipoproteins in the delivery of lipids to axons during axonal regeneration. J. Biol. Chem. 272, 30766–30773 (1997).
Minami, Y., Ode, K. L. & Ueda, H. R. Mammalian circadian clock: the roles of transcriptional repression and delay. Handb. Exp. Pharmacol 217, 359–377 (2013).
Mracek, P. et al. Regulation of per and cry genes reveals a central role for the D-box enhancer in light-dependent gene expression. PLoS ONE 7, e51278 (2012).
Seasholtz, A. F., Valverde, R. A. & Denver, R. J. Corticotropin-releasing hormone-binding protein: biochemistry and function from fishes to mammals. J. Endocrinol. 175, 89–97 (2002).
Timpl, P. et al. Impaired stress response and reduced anxiety in mice lacking a functional corticotropin-releasing hormone receptor 1. Nat. Genet. 19, 162–166 (1998).
Smith, G. W. et al. Corticotropin releasing factor receptor 1-deficient mice display decreased anxiety, impaired stress response, and aberrant neuroendocrine development. Neuron 20, 1093–1102 (1998).
Wang, K. C. & Chang, H. Y. Molecular mechanisms of long noncoding RNAs. Mol. Cell 43, 904–914 (2011).
Guttman, M. & Rinn, J. L. Modular regulatory principles of large non-coding RNAs. Nature 482, 339–346 (2012).
Yang, G., Lu, X. & Yuan, L. LncRNA: A link between RNA and cancer. Biochim. Biophys. Acta 1839, 1097–1109 (2014).
Sun, Q., Hao, Q. & Prasanth, K. V. Nuclear long noncoding RNAs: key regulators of gene expression. Trends Genet. 34, 142–157 (2018).
Ding, J. et al. A long noncoding RNA regulates photoperiod-sensitive male sterility, an essential component of hybrid rice. Proc. Natl Acad. Sci. USA 109, 2654–2659 (2012).
Wang, Y. et al. Arabidopsis noncoding RNA mediates control of photomorphogenesis by red light. Proc. Natl Acad. Sci. USA 111, 10359–10364 (2014).
Wang, Y., Li, J., Deng, X.-W. & Zhu, D. Arabidopsis noncoding RNA modulates seedling greening during deetiolation. Sci. China Life Sci. 61, 199–203 (2018).
Henriques, R. et al. The antiphasic regulatory module comprising CDF5 and its antisense RNA FLORE links the circadian clock to photoperiodic flowering. New Phytol. 216, 854–867 (2017).
Sánchez-Retuerta, C., Suaréz-López, P. & Henriques, R. Under a new light: regulation of light-dependent pathways by non-coding RNAs. Front. Plant Sci. 9, 962 (2018).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Janich, P., Arpat, A. B., Castelo-Szekely, V., Lopes, M. & Gatfield, D. Ribosome profiling reveals the rhythmic liver translatome and circadian clock regulation by upstream open reading frames. Genome Res. 25, 1848–1859 (2015).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Kikuchi, Y., Hosono, K., Yamashita, J., Kawabata, Y. & Okubo, K. Glucocorticoid receptor exhibits sexually dimorphic expression in the medaka brain. Gen. Comp. Endocrinol. 223, 47–53 (2015).
Acknowledgements
We thank NBRP-Medaka (National Bio-Resource Project of MEXT, Japan) and the Data Integration and Analysis Facility for use of their facilities. We also thank M. Okubo, A. Akama, N. Baba and C. Kinoshita for technical assistance, and T.K. Tamai for comments on the manuscript. This work was supported in part by the JSPS KAKENHI Grant-in-Aid for Specially Promoted Research (grant no. 26000013), the Human Frontier Science Program (grant no. RGP0030/2015) and Grant-in-Aid for JSPS Fellows (grant no. 18J10936). The WPI-ITbM is supported by the World Premier International Research Center Initiative, MEXT, Japan.
Author information
Authors and Affiliations
Contributions
T.Y. conceived the research. T. Nakayama, T. Shimmura and T.Y. designed the research. T. Shimmura, T. Shimo, T. Senga and T.Y. performed the microarray analysis for the short-day to long-day transition experiments. T. Nakayama, K. Okimura and T.Y. performed the IPA. T. Nakayama and Y.F. performed the strand-specific RNA-Seq. T. Nakayama performed the Ribo-Seq, qPCR analysis, cortisol measurement and behavioural assays. T. Nakayama, T. Shimmura, Y.T., T. Shimo and M.N. generated and genotyped LDAIR knockout medaka. T. Nakayama and A.S. performed the microarray analysis for the experiment using the LDAIR knockout fish. T. Nishimura and M.T. provided the medaka microarrays. Y.K., K.N. and K. Okubo provided new methods and materials. T. Nakayama and T.Y. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6 and Supplementary Table 1
Supplementary Data
Supplementary Data 1: List of photoperiodically-regulated transcripts. Supplementary Data 2: List of differentially expressed transcripts between wild type (WT) and LDAIR knockout (KO) medaka.
Rights and permissions
About this article
Cite this article
Nakayama, T., Shimmura, T., Shinomiya, A. et al. Seasonal regulation of the lncRNA LDAIR modulates self-protective behaviours during the breeding season. Nat Ecol Evol 3, 845–852 (2019). https://doi.org/10.1038/s41559-019-0866-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-019-0866-6
This article is cited by
-
Epigenetics and seasonal timing in animals: a concise review
Journal of Comparative Physiology A (2023)
-
Long non-coding RNAs in the physiology of aquaculture animals: a perspective update
Reviews in Fish Biology and Fisheries (2022)