Abstract
Mushroom-forming fungi (Agaricomycetes) have the greatest morphological diversity and complexity of any group of fungi. They have radiated into most niches and fulfil diverse roles in the ecosystem, including wood decomposers, pathogens or mycorrhizal mutualists. Despite the importance of mushroom-forming fungi, large-scale patterns of their evolutionary history are poorly known, in part due to the lack of a comprehensive and dated molecular phylogeny. Here, using multigene and genome-based data, we assemble a 5,284-species phylogenetic tree and infer ages and broad patterns of speciation/extinction and morphological innovation in mushroom-forming fungi. Agaricomycetes started a rapid class-wide radiation in the Jurassic, coinciding with the spread of (sub)tropical coniferous forests and a warming climate. A possible mass extinction, several clade-specific adaptive radiations and morphological diversification of fruiting bodies followed during the Cretaceous and the Paleogene, convergently giving rise to the classic toadstool morphology, with a cap, stalk and gills (pileate-stipitate morphology). This morphology is associated with increased rates of lineage diversification, suggesting it represents a key innovation in the evolution of mushroom-forming fungi. The increase in mushroom diversity started during the Mesozoic-Cenozoic radiation event, an era of humid climate when terrestrial communities dominated by gymnosperms and reptiles were also expanding.
Similar content being viewed by others
Main
Explosive diversification events, with intermittent periods of relatively little change, have generated uneven patterns of species richness across the tree of life. Although rapid radiations have been inferred in many clades1,2,3,4,5, few examples exist for fungi, in part because of their limited fossil record, and the lack of comprehensive phylogenies. Advances in modelling the evolutionary process now allow such inferences to be made from phylogenetic trees. Studies of diversification of mushroom-forming fungi have focused on individual clades6,7,8,9, yielding hypotheses on ecological opportunities6, the evolution of mutualistic lifestyle7,10 and fruiting body morphologies6,11 as drivers of adaptive evolution in fungi. However, these inferences, based on specific clades, remained untested across larger phylogenetic scales. Fruiting bodies provide support and protection for developing spores, which probably represents a driving force for increasingly more efficient morphologies12. Ancestral fruiting bodies of the Agaricomycotina were probably crust-like, ‘resupinate’ forms13,14, which then evolved into increasingly more complex forms, including derived ‘pileate-stipitate’ types, which are differentiated into a cap, stipe and hymenophore (spore-bearing surface). Pileate-stipitate forms have arisen repeatedly from simpler morphologies (for example, resupinate or coral-like) during evolution12,13,15 and dominate extant agaricomycete diversity (>21,000 described species16), but how this diversity arose, what explains the dominance of pileate-stipitate species in the class and whether fruiting body morphology impacts diversification rates (for example, as a key innovation) are not known.
Results and discussion
We investigated broad patterns of diversification in Agaricomycetes using a phylogenetic comparative approach and generated a highly comprehensive phylogeny of the group, comprising 5,284 species (Fig. 1). Our dataset represents around 26% of described Agaricomycetes, a notably high sampling density for any fungal group. Our inferences of phylogenetic trees combine a robust genome-based backbone phylogeny of 104 species (650 genes, 141,951 amino acid characters, see Supplementary Figs. 1–3, Supplementary Table 1 and Supplementary Notes 1 and 2) with sequence data from up to three nuclear loci for 5,284 taxa (large subunit (LSU) rRNA, rpb2, ef-1a, 5,737 characters). New sequence data were generated for 1,222 species (Supplementary Note 2 and Supplementary Data 1), including nine new genomes of phylogenetically key taxa (Supplementary Note 3). The phylogenetic tree topology is largely congruent with previously published clade-specific phylogenies and resolves 21 orders of Agaricomycetes plus Dacrymycetes and Tremellomycetes as outgroups. Most of the nodes received strong bootstrap support, but we also identified nodes that require more scrutiny and probably denser taxon sampling to be resolved with certainty (for a detailed genome-based overview of agaricomycete phylogeny, see Supplementary Note 1). We inferred ten time-calibrated phylogenies for the 5,284 taxa dataset using a two-stage Bayesian approach and all taxonomically informative, non-conflicting fossil calibration points available in the Agaricomycetes (eight fossils, determined by a fossil cross-validation procedure, see Supplementary Fig. 4, Supplementary Table 2 and Supplementary Note 4). Inferred node heights of the order-level clades range from 183 to 71 Myr, which places the origin of most order-level clades in the Jurassic, with the Cantharellales (mean age: 184 Myr, 144–261 Myr across ten chronograms), Agaricales (mean: 173 Myr, 160–182 Myr), Hymenochaetales (mean: 167 Myr, 130–180 Myr) and Boletales (mean: 142 Myr, 133–153 Myr) being the oldest. The Agaricomycetidae, the group uniting the Agaricales and Boletales was estimated at ~185 Myr (174–192 Myr, Fig. 1 and Supplementary Note 4). Because these molecular clock estimates are older than previous phylogenomics-based figures17,18,19, we performed sensitivity analyses to address the underlying causes, which suggest that the differences are attributable to the higher taxon sampling density in both our genome-based and 5,284-species tree and not to differences in software or choice of priors in fossil-based molecular clock calibrations. Various taxon and fossil sampling schemes, tree (5,284-taxon versus the genome-based phylogeny) and software choice had little effect on inferred ages (Supplementary Data 2 and Supplementary Note 4), providing a robust set of age estimates for Agaricomycetes.
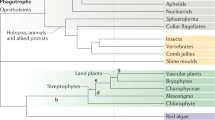
One of the 245 analysed maximum-likelihood trees was randomly chosen and visualized. Trees were inferred from nrLSU, rpb2, ef1-a sequences with a phylogenomic backbone constraint of deep nodes. Branches are coloured by net diversification (speciation minus extinction) rate inferred in Bayesian Analysis of Macroevolutionary Mixtures (BAMM). Warmer colours denote a higher rate of diversification. Significant shifts in diversification rate are shown by triangles at nodes. Only shifts present on >50% of ten trees, with a Bayesian posterior probability >0.5 and a posterior odds ratio >5 are shown. See Supplementary Data 6 for detailed discussion of shifts. Reconstructed probabilities of ancestral plant hosts for order-level clades are shown as pie charts partitioned by the inferred ancestral probability for gymnosperm (green) and angiosperm host (black). Pie charts are given for the most recent common ancestors of each order plus backbone nodes within the Agaricales—for small orders see Supplementary Data 3. Inner and outer bars around the tree denote extant substrate preference (black, angiosperm; green, gymnosperm; grey, generalist) and the placement of species used for inferring the 650-gene phylogenomic backbone phylogeny. Geological time scale is indicated with grey/white concentric rings.
We analysed global rates of speciation and extinction in a macroevolutionary framework using BAMM20. We accounted for incomplete and unequal sampling by using clade-specific sampling fractions and by ensuring that each genus was sampled in proportion to the number of its described species. Analyses of diversification rates in ten time-calibrated phylogenies provided evidence for rate variation through time and across lineages. Diversification rates increased abruptly at ~180 Myr ago in the early Jurassic (Fig. 2), coinciding with the breakup of Pangea, the onset of a warm, humid climate and the spread of (sub)tropical gymnosperm forests. Consistent with this, ancestral state reconstructions suggest that the last common ancestors of most orders were likely associated with gymnosperms as hosts (Figs. 1 and 3a, Supplementary Data 3 and Supplementary Note 5), followed by multiple shifts to and diversification in association with angiosperms as hosts during the Cretaceous, similar to patterns observed in a radiation of herbivorous beetles21. This agrees with observations of fungus-plant co-evolution at smaller scales10,22, frequent host switches during fungal evolution23 and might reflect the dependence of fungi on plants as nutrient sources or mutualistic partners.
a–d, Lineages through time plot, a, showing the log number of lineages through time; b, speciation rate; c, extinction rate and d, net diversification rate through geologic time estimated using BAMM on ten chronograms comprising 5,284-species. 95% confidence intervals of rate estimates are shown by shaded areas. The Jurassic period is shaded in green. Carb., Carboniferous; Perm., Permian and Tri., Triassic.
a, The evolution of substrate association through geologic time (top) and the evolution of vascular plant diversity and O2/CO2 levels (bottom, adapted from Niklas41 and Berner42. Inferred ancestral probabilities of fruiting body types were summarized across the ten phylogenies, by splitting the time scale of each tree into 100 bins and summing probabilities across the tree. b, The evolution of fruiting body types through time and the radiation of the agaricoid morphology since the Jurassic. Plot created as in a. For details of character coding and ancestral state reconstructions and detailed results, see Supplementary Note 5. c, Schematic model of the evolution of pileate-stipitate morphologies. The posterior distribution of transition rates from non-pileate (left) towards pileate-stipitate forms (right) and vice versa are shown above the arrows. Rates were estimated by Markov chain Monte Carlo (MCMC) in BayesTraits. d, Posterior distribution of trait-dependent speciation and extinction rates estimated under the binary state speciation and extinction (BiSSE) model for non-pileate versus pileate-stipitate fruiting body morphologies. Carb., Carboniferous; Perm., Permian and Tri., Triassic.
Acceleration of speciation is observed across all larger orders (Supplementary Data 4), suggestive of an external underlying cause, such as climate change or nutrient availability, instead of clade-specific events. We find that the rise in speciation rates in the Jurassic is coupled with a slightly delayed increase in extinction rates ~150 Myr ago (Fig. 2). Although estimating extinction rates from phylogenies is notoriously difficult24,25,26, this signal is also detected in an independent analysis using an episodic speciation/extinction model27 under a variety of conditions (Supplementary Note 6, Supplementary Fig. 6 and Supplementary Tables 3 and 4). Together, these data suggest that fungi experienced severe extinction in a period that falls close to a mass extinction event at the Jurassic/Cretaceous boundary28, an era with a fluctuating edaphic conditions, differential extinction of several metazoan groups, and a relatively stable flora. On the other hand, we find no evidence for increased probability of extinction around the Cretaceous/Tertiary boundary, a period of severe mass extinction in plant and animal lineages. It has been hypothesized that the massive deforestation after the bolide impact has buffered fungi through the period29, which is supported by a peak in fungal spore and hyphae fossilization right after the cataclysm30 and would explain the lack of mass extinction signal in our data.
The latitudinal diversity gradient (LDG) hypothesis posits that species richness increases from the poles towards the tropics. To test if this general prediction is valid for Agaricomycetes, we analysed latitude-dependent diversification rates across all 4,429 species for which sufficient geographic data could be obtained (see Methods). Agaricomycetes display their highest species diversity and net diversification rates in the temperate zone (Supplementary Figs. 7 and 8, Supplementary Tables 5 and 6 and Supplementary Note 7), which contrasts with the diversity gradient from the poles towards the tropics in animals and plants31. Although such a pattern could arise as a result of more concentrated efforts of recording fungi in the temperate zone, we find that the zone with highest speciation rate (between 20 and 40° in the temperate zone) does not coincide with the zone of the highest sampling density (30–60°) in our data. This suggests that our inference of a temperate peak in diversification rates is largely independent from sampling density. A reversed LDG has already been implied for major fungal groups by several surveys of environmental sequences32,33,34 and phylogenetic analyses22,35, in particular for ectomycorrhizal fungi. Although such signals might also be sensitive to biased taxonomic efforts across ecoregions, these data suggest that Agaricomycetes might not conform to broad macroecological patterns that dominate in animal and plant lineages.
To obtain a more resolved picture on diversification histories, we analysed rate variation across clades. We identified 85 significant shifts in diversification rate (defined as shifts present in ≥50% of trees with marginal odds ratio ≥5) that included clades of diverse ages spread out across the Agaricomycetes (Fig. 1, Supplementary Data 5, Supplementary Figs. 9 and 10, Supplementary Tables 7 and 8 and Supplementary Note 8). The largest of these comprises the Agaricomycetidae, consistent with results of rate through time and lineages through time analyses, which revealed an acceleration of net diversification rates in the Jurassic (Fig. 2). Order-level shifts to higher diversification rates were inferred in the last common ancestor of the Auriculariales; that of the Cantharellales, the Hymenochaetales (excluding the Repetobasidiaceae), the Russulales and the Agaricales (excluding the hygrophoroid and pleurotoid clades (sensu Matheny15) (Fig. 1). Most of the shifts (67%) were found in the Agaricales and predominantly in the Paleogene, consistent with a dynamic evolutionary history of the order but also with its largest share of diversity in the class. We found accelerated diversification in the genera Coprinellus and Laccaria, as reported previously6,7, but also in Cortinarius, Amanita and Lepiota, among others (for the complete list of shifts see Supplementary Data 5). Some of these represent the most taxonomically diverse agaric genera, such as Cortinarius with >2,000 (ref. 16) species described. Accelerated diversification in these groups is consistent with patterns seen in adaptive radiations36,37. Indeed, many clades show a diversification slowdown through time, a key sign of adaptive radiations36 (Supplementary Data 6), suggesting that many of the detected shifts may feature rapid adaptive filling of available niches. Among the 85 detected shifts in diversification rates, there were several that denoted old clades with low diversity, in which the shift represents a diversification slowdown rather than acceleration. For example, the genus Typhula, comprising species with minute club-shaped fruiting bodies, is an ancient group (dated at around 76 Myr) with a low diversity and an inferred diversification slowdown along its stem branch. Such parameters may indicate clades of relict lineages rooting deeply in Agaricales history but having only a few extant representatives. Taken together, these analyses provide a global view on bursts and slowdowns in the diversification history of mushroom-forming fungi and should facilitate a better understanding of adaptive diversification events in fungi, of which only a single example has been explicitly demonstrated6.
We next asked how morphological innovations may have contributed to shape the extant diversity of Agaricomycetes. Because fruiting body complexity is an important adaptive trait13,14, we mapped the evolution of fruiting body morphologies on the phylogeny and examined its impact on diversification rates. We performed ancestral state reconstructions and summarized inferred ancestral states through time across ten chronograms (Fig. 3b). In line with previous studies13,14, our reconstructions suggest ancestors with resupinate (crust-like) fruiting bodies for most early nodes along the backbone of the Agaricomycetes. The early dominance of resupinate morphologies was rapidly taken over by an array of more complex morphologies in the late Jurassic (~170–180 Myr), of which the pileate-stipitate type appears most dominant. This pattern could have been driven by the radiation of Agaricomycetidae, which contains the majority of pileate-stipitate species and started to diversify around this period, although pileate-stipitate species evolved in 12 orders as well, including the Russulales, Cantharellales and Gloeophyllales, among others. In the meanwhile, resupinate lineages may have simply continued to diversify at their previous rate, but, because of the radiation of species with pileate-stipitate fruiting bodies their dominance was diminished at the scale of the entire class. Consistent with this view, gains of the pileate-stipitate morphology are estimated to be significantly more likely (7–50 times, log(Bayes factor), logBF = 47) than their loss, indicating a trend towards pileate-stipitate morphologies, as reported earlier13. In fact, the pileate-stipitate morphology evolved convergently in at least 33 clades, suggesting it represents a stable attractor in the morphospace. Nevertheless, the probability of the loss of pileate-stipitate morphology is significantly larger than zero (logBF = 157.2), reflecting evolutionary transformations of pileate-stipitate fruiting bodies happening in many groups (Fig. 3c and Supplementary Data 1 and 7). We tested the impact of fruiting body morphologies on diversification rate in a character-state-dependent speciation and extinction framework38,39. Clades with pileate-stipitate fruiting bodies have a significantly higher speciation rate (P < 10−16, likelihood ratio test, LRT) than clades with other morphologies (Fig. 3d and Supplementary Data 7). This may be because the pileate-stipitate morphology provides support and protection for developing spores (for example, from precipitation) and enhances their dispersal efficiency by raising sporogenous tissue above ground-level; these advantages may explain how pileate-stipitate fruiting bodies can promote diversification and suggest they represent a key innovation for mushroom-forming fungi
Conclusions
This study represents a global analysis of the dynamic evolutionary history of mushroom-forming fungi. The diversity of Agaricomycetes might have been shaped by both class-wide diversification rate shifts and explosive clade-specific events. We identified a major class-wide radiation of Agaricomycetes in the Jurassic, which possibly followed a warming and humid climate in this period. The onset of this radiation coincided with the expansion of angiosperms and followed patterns of gymno- and later angiosperm expansion during the Mesozoic-Cenozoic radiation40. Our analyses provide evidence for several clade-specific adaptive radiations in the class, offering a possible explanation for why some genera contain a single species, while others contain thousands. Our data display strong signal for a mass extinction event coinciding with increased extinction rates at the Jurassic/Cretaceous boundary, however, no such evidence was found in the Agaricomycetes for the much more influential Cretaceous–Tertiary mass extinction event. The evolution of pileate-stipitate fruiting bodies, that is, morphologies with cap and stipe might be a key driver of evolutionary radiations, suggesting that the convergent evolution of the classic mushroom morphology contributed to shaping extant diversity in the Agaricomycetes. We predict that further data on traits and ecological opportunities that generated the spectacular diversity of mushroom-forming fungi will help to elucidate the general principles of mushroom evolution and integrate this ecologically important group into the record of major biodiversification events.
Methods
Molecular biology methods
Genomic DNA was extracted from 0.1–10 mg of dried herbarium specimens using the DNeasy Plant Mini kit (Qiagen) or the Nucleospin Plant II mini kit (Macherey-Nagel) following the manufacturers’ instructions. PCR amplification was performed as described previously6,15 on the LSU of the nuclear ribosomal RNA with the LR7 and LROR primers. Sequencing was performed commercially using the same primers as used for PCR. Sanger reads were assembled to a contig using the Pregap and Gap4 programs of the Staden package43 and checked manually against the electropherograms.
Taxon sampling
We sampled herbarium specimens so that taxon sampling be proportional across clades and geographic regions. To that end, we initially obtained estimates of the known diversity for each of the genera from Species Fungorum44 (http://www.speciesfungorum.org), the Dictionary of the Fungi16 and Funga Nordica45. Taxonomic information from these sources was compiled for all genera of Agaricomycetes. Using this compilation, we calculated the number of target species that needed to be sampled for each genus to obtain a final dataset of ~5,000 species (~30% sampling of total diversity) in which sampling of species in each genus is proportional to their described diversity. Although the three sources differed in their completeness, geographic scope (Dictionary of the Fungi and Species Fungorum being global databases and Funga Nordica being, taxonomically, the most stringent of the three) and taxonomic concept, it gave us general guidance on how to keep taxon sampling proportional across genera. When sampling species within genera, attempts have been made to obtain specimens from undersampled geographic regions—in particular, Africa, Asia, Australasia and South America.
Sequence alignment
Multiple sequence alignment was carried out for the LSU, ef-1a and RPB2 loci separately using the Probabilistic Alignment Kit (PRANK release 140603)46,47. An iterative alignment refinement strategy as described in Tóth et al.48 was employed: maximum-likelihood gene trees computed from preliminary alignments (using RAxML, see below) were used as guide trees for the next round of multiple alignment for PRANK. After three rounds of iterative refinement, the alignments were further corrected manually using a text editor. Manual curation was restricted to correcting homologous regions erroneously juxtaposed by PRANK. Alignments of individual sequences were concatenated into a superalignment. The concatenated alignment is available at Dryad (accession number doi:10.5061/dryad.gc2k9r), GenBank accession numbers for all sequences used are shown in Supplementary Data 1.
Genome and transcriptome sequencing and assembly
Sequencing
The genomes of Coprinopsis marcescibilis, Crucibulum laeve, Dendrothele bispora, Heliocybe sulcata, Peniophora sp., Pluteus cervinus, Polyporus arcularius and Pterula gracilis were sequenced using a combination of fragment and long-mate pair Illumina libraries. In addition, the genome of Coprinellus micaceus was sequenced using Pacific Biosciences platform. Fragment Illumina genomic libraries were produced for 100 ng of DNA was sheared to 270 bp using the Covaris LE220 and size selected using SPRI beads (Beckman Coulter). The fragments were treated with end repair, A-tailing and ligation of Illumina compatible adaptors (IDT, Inc.) using the KAPA-Illumina library creation kit (KAPA biosystems). For long-mate pair libraries (C. laeve, D. bispora, H. sulcata, P. arcularius and P. gracilis), 5 μg of DNA was sheared using the Covaris g-TUBE and gel size selected for 4 kb. The sheared DNA was treated with end repair and ligated with biotinylated adaptors containing loxP. The adaptor ligated DNA fragments were circularized via recombination by a Cre excision reaction (NEB). The circularized DNA templates were then randomly sheared using the Covaris LE220 (Covaris). The sheared fragments were treated with end repair and A-tailing using the KAPA-Illumina library creation kit (KAPA Biosystems) followed by immobilization of mate pair fragments on strepavidin beads (Invitrogen). Illumina compatible adaptors (IDT, Inc.) were ligated to the mate pair fragments and eight cycles—or in the case of Heliocybe sulcata ten cycles—of PCR were used to enrich for the final library (KAPA Biosystems). For C. micaceus unamplified libraries were generated using the Pacific Biosciences standard template preparation protocol. Then, 5 μg of genomic DNA was used to generate each library and the DNA was sheared using Covaris g-Tubes to generate sheared fragments of 10 kb in length. The sheared DNA fragments were then prepared using the Pacific Biosciences SMRTbell template preparation kit, where the fragments were treated with DNA damage repair, had their ends repaired so that they were blunt-ended and 5’ phosphorylated. Pacific Biosciences hairpin adaptors were then ligated to the fragments to create the SMRTbell template for sequencing. The SMRTbell templates were then purified using exonuclease treatments and size selected using AMPure PB beads.
All transcriptomes in this study were sequenced using Illumina RNA-Seq. Stranded complementary DNA libraries were generated using the Illumina Truseq Stranded RNA LT kit. Messenger RNA was purified from 1 μg of total RNA using magnetic beads containing poly-T oligos. mRNA was fragmented and reversed transcribed using random hexamers and SSII (Invitrogen) followed by second strand synthesis. The fragmented complementary DNA was treated with end-pair, A-tailing, adaptor ligation and ten cycles or, in the case of C. marcescibilis, C. laeve and P. cervinus, eight cycles of PCR.
The prepared Illumina libraries were quantified using KAPA Biosystem’s next-generation sequencing library quantitative PCR kit and run on a Roche LightCycler 480 real-time PCR instrument. The quantified libraries were then multiplexed with other libraries, and the pool of libraries was then prepared for sequencing on the Illumina HiSeq sequencing platform using a TruSeq paired-end cluster kit, v.4 and Illumina’s cBot instrument to generate a clustered flow cell for sequencing. Sequencing of the flow cell was performed on the Illumina HiSeq2500 sequencer using HiSeq TruSeq SBS sequencing kits, v.4, following a 2 × 150 indexed run recipe. In the case of C. micaceus, Dendrothele bispora, Peniophora sp. and P. arcularius, TruSeq paired-end cluster kit, v.3, and TruSeq SBS sequencing kits, v.3, were used and in the cases of C. laeve, H. sulcata and P. gracilis a 2 × 100 indexed run recipe was performed.
For PacBio sequencing primer was annealed to the SMRTbell templates and Version P5 sequencing polymerase was bound to them. The prepared SMRTbell template libraries were then sequenced on a Pacific Biosciences RSII sequencer using Version C3 chemistry and 1 × 240 min sequencing movie run times.
Assembly and annotation
Genomic reads from each pair of Illumina libraries were QC filtered for artefact/process contamination and assembled together with AllPathsLG v.R4940349. Illumina reads of stranded RNA-seq data were used as input for de novo assembly of RNA contigs and assembled into consensus sequences using Rnnotator (v.3.4)50. The Pacific Biosciences sequence data were filtered with smrtanalysis assembled together with Falcon version 2014082 (ref. 51), improved with finisherSC52 and polished with Quiver. All genomes were annotated using the JGI Annotation Pipeline and made available via the JGI fungal portal MycoCosm53. Genome assemblies and annotation were also deposited at DDBJ/EMBL/GenBank.
Phylogenomic analysis of 104 Agaricomycotina
Phylogenomic analysis
The genome sequences of 104 species (Supplementary Table 1) were included in the phylogenomic analysis of Agaricomycotina. Two representatives of Tremellomycetes (Tremella mesenterica (Treme1), Cryptococcus neoformans var. grubii H99 (Cryne_H99_1)) were used as outgroups.
We performed all-versus-all blast using mpiBLAST 1.6.0 (ref. 54) with default parameters, then identified gene families using the Markov clustering algorithm MCL 14-137 (ref. 55) with an inflation parameter of 2.0. We first identified gene families that contained 52–208 genes (50–200% of the species included in the study), then performed multiple sequence alignment using PRANK 140603(ref. 47). Ambiguously aligned regions were removed from the alignments using Gblocks 091b56 with the settings -p = yes -b2 = 26 -b3 = 10 -b4 = 5 -b5 = h -t = p -e = .gbl. Next, we screened for gene families that contained a single representative gene for each species, or ones that contained inparalogs but no deep paralogs. Deep paralogs were identified following Nagy et al.57. Gene trees were inferred using the PTHREADS version of RAxML 8.1.2 (ref. 58) using the PROTGAMMAWAG model and the standard algorithm. A single inparalog, closest to the root on the basis of root-to-tip patristic distances was retained for each species. Gene families in which 75% or more of the species were represented were concatenated into a supermatrix.
We used RAxML 8.1.2 to perform maximum-likelihood analysis and bootstrapping on the concatenated dataset, under a WAG model with gamma-distributed rate heterogeneity partitioned by gene. We ran 100 bootstrap replicates using the rapid hill-climbing algorithm. Because in initial analyses Peniophora sp., Tubaria furfuracea and Clavaria fumosa were placed on long terminal branches, we performed the following steps to clean the alignments of overly divergent sequences (either due to sequencing errors or pseudogenes). We screened all individual gene trees and excluded proteins with a terminal branch length >2.75 times longer than the average branch length of the gene tree. Altogether, 1,294 (in 542 gene families) protein sequences were excluded from the analyses.
Sensitivity analysis
The robustness of the dataset was tested by eliminating incrementally higher numbers of fast-evolving sites using six levels of stringency in Gblocks 091b56. Using these parameters, we eliminated 8.5% (-b1 = 78 -b2 = 78 -b3 = 10 -b4 = 10), 24.3% (-b1 = 78 -b2 = 78 -b3 = 8 -b4 = 15), 36.7% (-b1 = 88 -b2 = 88 -b3 = 8 -b4 = 15), 46.4% (-b1 = 95 -b2 = 95 -b3 = 8 -b4 = 15), 50.1% (-b1 = 100 -b2 = 100 -b3 = 8 -b4 = 15) and 55.2% (-b2 = 104 -b3 = 5 -b4 = 20) of the least reliably aligned regions of the alignments, resulting in trimmed concatenated datasets with 129,886, 107,496, 89,732, 76,153, 70,862 and 63,309 amino acid sites, respectively. We performed maximum-likelihood phylogenetic inference for each of the reduced datasets in RAxML as described above.
Maximum likelihood analyses of the 5,284-taxon dataset
Maximum-likelihood trees for the 5,284-taxon dataset were inferred using the parallel version of RAxML v.8.1.2 (ref. 58) under the generalized time-reversible model with gamma-distributed rate heterogeneity (four categories) with three partitions corresponding to the LSU, ef1-α and rpb2 loci. The phylogenomic tree was used as a backbone monophyly constraint. We performed 245 maximum-likelihood inferences and tested whether these trees adequately represented the plausible set of topologies given the alignment. This was done to ensure that phylogenetic uncertainty is properly taken into account in subsequent comparative analyses. If our tree set contains all plausible topologies, then the rolling average of pairwise Robinson–Foulds distances should show a saturation as a function of increasing the number of trees. To this end, we computed Robinson–Foulds distance for each pair of trees for incrementally larger numbers of trees using R package ‘phangorn’ v.2.0.2 (ref. 59). We then plotted the rolling average, maximum and minimum values as a function of the number of trees in R.
Molecular clock dating of the 5,284-taxon dataset
Due to computational limitations, we focused on ten maximum-likelihood trees that maximize topological diversity (as inferred on the basis of Robinson–Foulds distances) as representative trees for molecular clock analysis. These were chosen by first calculating pairwise Robinson–Foulds distances between trees, then hierarchically clustering trees using Ward’s clustering method (as implemented in the hclust R function). The ten maximum-likelihood trees for subsequent analyses were chosen evenly from the resulting clusters of trees. To overcome the computational limitations of inferring accurate chronograms for thousand-tip phylogenies, we adopted a two-step strategy. First, PhyloBayes 4.1b60 was used to infer parameters of a birth-death model and divergence times of trees on 10% subsampled versions of the 5,284-taxon dataset. Subsampled datasets were obtained by randomly deleting 90% of the species, but forcing key descendants of fossil-calibrated nodes to be retained in the subsampled alignments. Then we used the ages and parameters estimated in PhyloBayes as input parameters for FastDate development version61 (provided by T. Flouri) for analysing the complete 5,284-species dataset.
PhyloBayes analyses were run using the 10% subsampled dataset, a birth-death prior on divergence times, an uncorrelated gamma multiplier relaxed clock model and a CAT-poisson substitution model with a gamma distribution on the rate across sites. A uniformly distributed prior was applied to fossil calibration times. All analyses were run until convergence, typically 15,000 cycles. Convergence of chains was assessed by visually inspecting the likelihood values of the trees and the tree height parameter. We sampled every tree from the posterior and after discarding the first 7,000 samples as burn-in we summarized the posterior estimates using the readdiv function of PhyloBayes.
Next, we used FastDate, a program that implements the speed dating algorithm described in Akerborg et al.61. FastDate was run on the complete trees (5,284 species) with the node ages constrained to the values of the 95% highest posterior densities of the ages inferred by PhyloBayes. FastDate analyses were run with time discretized into 1,000 intervals and the ratio of sampled extant individuals set to 0.14.
Fossil calibrations
The extensive sampling density of our tree allowed us to use more fossil calibration points than any study before and to place them more precisely within the tree. Eight fossils were used as calibration points (Supplementary Table 2) for the crown node of the specified taxa.
We excluded a number of prospective well-preserved fossils for reasons of redundantly calibrating an already calibrated node (Appianoporites vancouverensis—Hymenochaetales62, Protomycena electra—marasmioid clade63, Cyathus dominicanus—Nidulariaceae64, Fomes idahoensis—Polyporales65. Geastrum tepexensis66 was not considered as it could be assigned to either of the two earth-star clades, one in the Phallomycetidae (Geastraceae) or the other in the Boletales (Astraeus). For other mushroom fossils, the taxonomic affiliations were deemed too uncertain.
To investigate if there is any conflict between the fossil calibration points we conducted a fossil cross-validation analysis following Near et al.67. For this, we ran PhyloBayes analyses on one of the 10% subsampled trees with the same settings as mentioned above, using a single fossil calibration point at a time. This resulted in eight independent molecular dating analyses. To quantify conflict between single fossil calibrations, first we calculated the sum of the square differences between molecular age estimates and fossil ages:
where x is the fossil calibration point used and Di is the difference between molecular age estimate and fossil age for fossil i. We ordered the SS values of each of the eight analyses in descending order and calculated the average squared deviation for all fossil calibrations:
Next, the analysis with the highest SS value was removed and s was recalculated. We continued this process until only two analyses remained. In parallel, we performed one-tailed F-tests to check whether the removal of the fossil had a significant effect on the variance of s. The principle of this procedure is that during the stepwise removal of fossils, s should decrease by small constant values and variance should not decrease significantly
Genome-based molecular clock analyses
We examined the robustness of our molecular age estimates using phylogenomic methods as an alternative to the 5,284 species phylogeny. Specifically, we tested whether the differences between our and previous17,18 molecular clock estimates for Agaricomycete orders were attributable to differences in the dataset or analytical method used or a difference in how precisely fossils could be placed on the tree. To this end, we performed a series of molecular clock analyses on our phylogenomic dataset that resembled that of Kohler and Floudas in terms of topology and taxon sampling density, but differed in the placement of some fossil calibrations. A key difference was that our phylogenomic tree included the most recent common ancestor (MRCA) of the Hymenochaetales and that of the Suillaceae, which allowed us to calibrate the MRCA of these clades, as opposed to their stem nodes by Kohler et al.18 and Floudas et al.17 Further, our analysis includes the MRCA of the Nidulariaceae, a relatively recent node that can be calibrated by two well-preserved fossils described recently64
Phylogenomic dataset
We used a smaller, more conserved subset of the 568-gene and 104-species phylogenomic dataset (which was computationally not tractable in these analyses). First, we selected the first 70 most conserved genes of the 568-gene dataset by calculating the mean genetic distances for each gene using the dist.alignment function of the seqinR R package v.3.4-5 (ref. 68). To enable a more accurate placement of fossil calibration points we added additional three species (Cyathus striatus, Pycnoporus cinnabarinus, Suillus brevipes) to this dataset (Supplementary Table 1) and excluded two taxa that harboured ambiguous positions. We searched homologous sequences in the additional genomes using blastp v.2.7.1 (ref. 69) with one randomly selected gene from each of the 70 gene families as query. We selected the best hit (smallest E-value) as a one-to-one ortholog if the second best hit had a significantly worse E-value (by 20 orders of magnitude). Protein clusters were aligned by PRANK v.100802 (refs. 46,47) using default settings. Next, conserved blocks of the alignments were selected using Gblocks v.0.91b56 with default settings except for the minimum length of a block that was set to 5 and gap positions in half of the sequences were allowed. A phylogenomic tree was constructed by RAxML v.8.2.11 (ref. 58) under the WAG + G substitution model partitioned by gene.
Calibrations
To dissect sources of differences in molecular age estimates, we ran analyses under three fossil calibration schemes (Supplementary Data 2) and the 105-species phylogenomic tree. First, we used the same fossil calibration scheme as for the 5,284-species phylogenetic dataset (‘Default calibration scheme’). Next, we replicated the analyses of Kohler et al.18 on our tree, using the fossil calibration points from Kohler et al. (‘Kohler et al. calibration scheme 1’). For this, we placed the suilloid ectomycorrhiza fossil in the split of Suillinae/Paxillinae/Sclerodermatinae and Archaeomarasmius leggettii in the MRCA of Gymnopus luxurians and Schizophyllum commune with uniformly distributed 40–60 Myr and 70–110 Myr ago time priors, respectively. Finally, we used the calibrations used by Kohler et al. (scheme 2) but placed the two fossils in the MRCAs of the Suillaceae and marasmioid clade, respectively (Kohler et al.18 calibration scheme 2’). In all analyses we constrained the age of the root to be between 300 Myr and 600 Myr ago.
Penalized likelihood analysis in r8s
We ran a series of molecular clock analyses in r8s v.1.81 (ref. 70). A cross-validation analysis was performed to determine the optimal smoothing parameter (λ) by testing values across seven orders of magnitude starting from 10−3. The additive penalty function was applied and the optimization was run 25 times starting from independent starting points. In one optimization step, after reaching an initial solution, the solution was perturbed and the truncated Newton optimization was rerun 20 times. We compared the results of previous studies to that of analyses across seven ancestral nodes in Agaricomycotina (Supplementary Data 2).
Bayesian molecular clock dating
We used the mcmctree method implemented in PAML v.4.8a71. The independent-rates clock model, a WAG substitution model and approximate likelihood calculation72 were used. The birth rate, the death rate and the sampling fraction of the birth-death process were set to 1, 1 and 0.14, respectively. The shape and the concentration parameter of the gamma-Dirichlet prior for the drift rate coefficient (σ2) was set to 1 and three different scale parameters were tested (10, 100, 1,000) to see their effect on the time estimates. The substitution rates of each gene were estimated by codeml under a global clock model, to set the parameters of the gamma-Dirichlet prior for the overall rate. By calculating the mean substitution rate of the loci and examining the density plot of the rates we set up a prior that reasonably fitted the data: the shape parameter, the scale parameter and the concentration parameter were set to 5, 90.7441 and 1, respectively, resulting in an average substitution rate per site per time unit of 0.055. We set the time unit to 100 Myr and applied uniform priors on eight fossil calibrations with lower and upper hard bounds. MCMC analysis was run for 80,000 iterations, discarding the first 20,000 iterations as a burn-in and sampling every 30th tree from the posterior. After three independent analyses were run the convergence of log-likelihood values was visually inspected and the estimated ages were compared between replicates.
Analyses of diversification and character evolution
Character coding
We discretely coded three characters, the presence of a cap, fruiting body type and substrate preference for the 5,284 species. Fruiting body types were coded as one of six types and data were compiled from the literature. The six character states distinguished were: (0) no fruiting body; (1) resupinate; (2) agaricoid; (3) cyphelloid; (4) gasteroid/secotioid and (5) coralloid/clavarioid. Transitional morphologies, or species for which no clear decision could be made on fruiting body type were coded as uncertain. Resupinate fruiting bodies were defined as crust-like or effused, flat morphologies that follow the morphology of the substrate, but irrespective of thickness or hymenophore type. The agaricoid type was defined as having a distinguishable stipe and cap. Cyphelloid species were coded following Bodensteiner73. The gasteroid types included species with closed fruiting bodies that produce spores internally. No distinction was made between secotioid, sequestrate and false-truffle morphologies and all were coded as gasteroid. Fruiting bodies with club-shaped or branched, erect morphology but no differentiated cap were considered coralloid/clavarioid.
The presence of a cap, that is pileate-stipitate morphology, was coded on the basis of literature data as either absent (state 0) or present (state 1). Species with rudimentary or reduced caps were coded as uncertain.
We coded species’ preference for substrate on the basis of literature data (a detailed listing of resources used is not given) as gymnosperm, angiosperm or both (that is, generalist species). Species were considered gymno- or angiosperm specialists if >90% of records were for either plant group, or their habitat preferences in taxonomic literature were described by the terms ‘exclusively’, ‘predominantly’, ‘mostly’ or occurring on the least dominant substrate ‘rarely’, ‘very rarely’ or ‘exceptionally’. On the other hand, species having records on both gymno- and angiosperm substrates, described in the literature as generalists or given multiple gymno- or angiosperm substrate plant species were coded as generalists. Species with insufficient data or no data at all were treated as missing data. Soil-inhabiting saprotrophs were coded on the basis of their association with either gymno- or angiosperms, if a clear preference was reported in the literature.
Tests of the LDG hypothesis
Global positioning system (GPS) data
We downloaded 5,884,445 fungal GPS records from the GBIF database, representing 4,429 Agaricomycetidae species. For 848 species, a location based on the province, country and/or biogeographic realm74 was obtained from the literature. In cases where only the province, country or biogeographic realm was available, we took the centroid of the area as a proxy for GPS location. All GPS data were handled and processed in R75. Noise was added to identical GPS coordinates using the jitter function in R.
Analyses of geographic diversification
State-dependent diversification rates were estimated in two ways: (1) using centroid latitudes as continuous traits (for example, Sánchez-Ramírez et al.35) and (2) using discrete areas. We used the data mentioned above to first calculate a centroid coordinate (RGEOS R package) in cases where multiple GPS records exist per species. Then we took the centroid latitude for each species. For singletons, we simply took the latitude of each record. We also used the GPS database to calculate a standard deviation for latitude per species. For species with a single record, we took the mean standard deviation of species with multiple records. We fitted multiple Quantitative State Speciation and Extinction (QuaSSE76) models, using a maximum-likelihood approach, to ten of the 5,284-species chronograms. Models included speciation rate changes as a constant function of the trait (no effect), as a linear function or as a Gaussian function, keeping the extinction rate constant76. We also added two models in which the extinction rate varied as linear or Gaussian functions, while speciation rate remained constant. To improve the efficiency of the algorithm given the scale of the tree, we applied the following settings to the models: method = ’fftC’, dt.max = 1 and nx = 256. Akaike information criterion values were averaged across phylogenies and compared between different models. We used the best supported model to interpolate speciation rates into a 1 × 1 degree global map made with Natural Earth. (Free vector and raster map data are available at www.naturalearthdata.com.)
To obtain discrete areas, first we divided all terrestrial WWF ecoregions into either tropical or extra-tropical. For tropical areas we considered: Tropical and Subtropical Moist Broadleaf Forests, Tropical and Subtropical Dry Broadleaf Forests, Tropical and Subtropical Grasslands, Savannas and Shrublands. All other terrestrial ecoregions were deemed extra-tropical. Ecoregion polygons were integrated (that is, dissolved) in a way that only two-state areas were found (Supplementary Note 7). For species with multiple GPS records, we extracted binary information about their presence in both areas. For records that were not found precisely within the geometry of the area, we took the area to which the distance to the GPS point was closer. To assign discrete states to each species with multiple records, our criterion was to assign a ‘widespread’ state (both areas) if records for the area with the minor frequency was higher than 0.2. In any other case, we assigned ‘tropical’ or ‘extra-tropical’ states to taxa in the area with the highest frequency. For taxa that had only biogeographic realm data, we coded them as follows: ‘widespread’: Neotropical, Afrotropical, Oceanian/Australia; ‘tropical’: Panamenian, Oriental and extra-tropical': Europe/Palaearctic/North Asia, Nearctic, Saharo-Arabian, Sino-Japanese. Using these three states for each taxon, we fitted multiple constrained models and one full model of Geographic State Speciation and Extinction (GeoSSE77) in a maximum-likelihood framework. The full model consisted of seven parameters: (1) speciation rate in the tropics, (2) speciation rate in the extra-tropical zone, (3) the allopatric speciation rate (speciation rate in both tropical and extra-tropical regions), (4) extinction rate in the tropics, (5) extinction rate in the extra-tropical zone, and dispersal rates to (6) the tropics and to (7) the extra-tropical zone. The constrained models consisted of: (1) speciation in the tropics is equal to speciation in the extra-tropical zone; (2) extinction in the tropics is equal to extinction in the extra-tropical zone and (3) dispersal to the tropics occurs at the same rate as dispersal to the extra-tropical zone. In each case, a single parameter for speciation, extinction, and dispersal was respectively estimated, allowing the other parameters of the model to vary. To explore the parameter space better, we re-estimated parameters using MCMC on the full model. In a similar way as above, we used Akaike information criterion scores to rank the models.
Analyses of character evolution
Model tests
To infer substitution models that best describe the evolution of agaricoid/non-agaricoid fruiting bodies we used maximum-likelihood and MCMC approaches implemented in BayesTraits 2.0 Linux 64 Quad Precision alternative build78 and the R package diversitree v.0.9–8 (ref. 39). The BiSSE model38 was used in diversitree. We performed model tests by first constraining the forward and reverse transition rates between non-agaricoid (state 0) and agaricoid type (state 1) to be equal (q01 = q10). We also separately constrained each of the rates to be zero (q01 = 0 or q10 = 0). Each of the constrained models were compared to the best fit model on the basis of log-likelihood values (maximum-likelihood analyses in BayesTraits), log-Bayes factors (MCMC analyses in BayesTraits) or LRT and Akaike information criterion scores (maximum-likelihood analyses of BiSSE model in diversitree)38,39,78,79. In BayesTraits, a difference of 2.00 log-likelihood units (maximum-likelihood approach) or a difference of 10 log marginal likelihood units (MCMC approach) was considered as significant support for a model over another, while in the BiSSE analyses significance was assessed using LRTs (P < 0.05).
BayesTraits analysis
BayesTraits analyses (maximum likelihood and MCMC) were performed on 245 phylogenetic trees using the MultiState module of the program. Before the final MCMC analyses, we tried several prior distributions (uniform, exponential, gamma and the hyper-prior versions of these) with different settings. On the basis of preliminary analyses, we found that the gamma distribution was most optimal, therefore in further analyses we used a gamma hyper-prior with different prior distributions for each parameter (Supplementary Note 5). All preliminary BayesTraits analyses were conducted with the following settings: 1,010,000 generations, 10,000 generations as burn-in and sampling every 500th generation. We forced Markov chains to spend 200,000 generations on each tree using the equaltree option, with 100,000 generations as burn-in and sampling every 500th generation. The marginal likelihood was estimated by the stepping stone method78,80 using 50 stones with a chain length of 5,000. All analyses in BayesTraits were repeated three times to check the congruence of independent runs.
Diversitree analysis
We used ten chronograms to analyse trait-dependent diversification under the BiSSE model implemented in the R package diversitree v.0.9–10 (refs. 38,39). Maximum-likelihood search started from the point in the parameter space determined by the function starting.point.bisse. Bayesian MCMC was performed using an exponential prior, defined as 1/(2r), where r is the character independent diversification rate. State-specific sampling fractions were defined on the basis of data from Species Fungorum44 (see Accounting for random and incomplete taxon sampling). We first optimized the MCMC sampler’s step size argument by running 100 generations, then we ran MCMC analyses for 20,000 generations with burn-in set to 10%. The convergence of the chains was visually checked on the basis of likelihood and parameter values.
Ancestral character-state reconstruction
We reconstructed the ancestral states using stochastic character mapping as implemented in phytools v.06–20 (ref. 81), to reveal the dynamics of fruiting body types and substrate preference through time. This method is a modified version of a previously published algorithm82, which samples discrete character histories from the posterior probability distribution. We performed the analysis with the make.simmap function on ten time-calibrated trees under a Markov model with all rates different. The stochastic character histories were simulated 5,000 times. We plotted state posterior probabilities through time using a custom R script. Briefly, we summarized character-state posterior probabilities through the time scale of our ten chronograms split into 100 bins. The plot was created using ggplot and geom_area functions of ggplot2 v.2.2.1 (ref. 83). Ancestral probability distributions of substrate preference for the MRCAs of each of the orders and some additional clades were plotted as pie charts using the nodelabels function of ape v.4.1 (ref. 84).
Diversification rate analyses
Accounting for random and incomplete taxon sampling
To meet the assumption of random or complete sampling of the BAMM and BiSSE models20,39, we specified the sampling fraction of each genus in accordance with the described number of species based on Species Fungorum44. To gather information on described species, we screened all orders of the Agaricomycetes, Dacrymycetes and Tremellomycetes in Species Fungorum and gathered all species with a custom java program (available from the authors on request). We took into account taxonomic and nomenclatural synonymy if indicated by Species Fungorum. We accounted for incomplete taxon sampling by two strategies. First, we assigned specific sampling fractions to the character states (BiSSE model) or to genera (BAMM model). In these cases we used the built-in correction of the BiSSE and BAMM models20,85 to account for missing species in our phylogeny. Second, because we could have unintentionally oversampled certain genera (for example, because of better availability of specimens), we statistically tested for oversampling and generated a pruned phylogeny in which each genus is represented in proportion of its described diversity. We adjusted taxon sampling in our phylogeny by iteratively deleting species from genera that were oversampled relative to the mean sampling fraction of the tree, until sampling fractions of each genera corresponded their known size as judged by a comparison to Species Fungorum. To do this, we performed a hypergeometric test (P < 0.05) at each iteration of species removal. Elimination of species was stopped when oversampling disappeared (P > 0.05, hypergeometric test). With this procedure we wanted to produce a ‘skeletal tree’ (sensu FitzJohn et al.85), where species were both evenly and randomly sampled from every genera85.
Trait-dependent diversification rate analyses
We performed analyses under the BiSSE model to examine the effect of agaricoid fruiting body type on diversification rate. BiSSE analyses were carried out in diversitree v.09–10 as described above, with the following differences. We performed model tests to examine if the presence of a cap influenced speciation and extinction rates. To this end, we first constrained state-specific speciation or extinction rates to be equal (λ0 = λ1 or μ0 = μ1) then constrained both the speciation rates and extinction rates to be equal (λ0 = λ1 and μ0 = μ1). Models were compared by LRT in R.
Trait-independent diversification rate analyses
We used BAMM v.2.5.0 (ref. 20) to examine rate heterogeneity across lineages and detect shifts in diversification rates. We analysed ten chronograms and ran MCMC analyses for 100 million generations using four independent chains per analysis with 50 million generations as burn-in. Prior parameters were optimized using the setBAMMpriors function in BAMMtools v.2.1.6 (ref. 86), except for the prior on the expected number of shifts, which was set to 270 on the basis of preliminary runs. We accounted for incomplete taxon sampling as described above. We checked the convergence of chains by visually inspecting the convergence of likelihoods, by calculating the effective sample size and the Geweke’s diagnostic87 of log-likelihoods, numbers of shifts and evolutionary rate parameters, using functions in CODA 0.19-1 (ref. 88). Geweke’s diagnostic tests for the quasi equality of the means of parameters sampled at different parts of the MCMC analysis; it means that, if the beginning and end regions of the post-burn-in MCMC sample do not differ to a statistically significant extent (P > 0.05), then the Markov chain is considered to be in a stationary phase. To ensure that a shift is highly supported by the data and the prior had negligible contribution, we examined only core shifts; that is, those with a prior-to-posterior marginal odds ratio exceeding 5 (ref. 86).
We compared core shifts across ten chronograms to obtain a consensus view on shifts that can be detected in all or most of the trees. To this end, we first empirically identified taxonomically similar clades for which a core shift was inferred, taking into account topological differences among trees. Then we noted whether the mean diversification rate change after the shift is positive or negative (that is, rate acceleration or deceleration). There were cases when several core shifts were inferred on adjacent branches around the MRCA of a given clade. These come from distinct shift configurations sampled during the MCMC analysis and correspond to the same signal of rate variation in the data but were located to adjacent branches of the tree. In such cases, we chose the shift with the highest posterior probability, noting that the biological reality of increasing rates might be spread out across a few adjacent branches around the one with the highest posterior. Finally, shifts were referred to as congruent core shifts if they were highly congruent across four or more trees and had strong tree support (mean posterior probability >0.5).
Next, we assessed whether posterior probabilities of congruent core shifts reached convergence in a fashion similar to the ‘cumulative’ function of AWTY89. For this, we calculated the posterior probability of each core shift at each generation, as if the analysis was stopped at that point and plotted posterior probabilities as a function of generation using the ggplot2 v.2.2.1 package83. We also took into account the distinct shift configurations by examining whether a rate increase, after a congruent core shift was explained by a rate decrease after another core shift. First, we determined congruent core shift—core shift pairs that could potentially be mutually exclusive to each other. These shift pairs were determined in one tree, usually in the tree where a congruent core shift had high posterior probability. Then we calculated two kinds of posterior proportions: one for co-occurring shift pairs and one for each of the single occurrence of the shifts. We said that two shifts were mutually exclusive to each other if only a negligible co-occurrence was presented and the direction of the diversification rate change was different between the single occurrence posterior samples.
To reveal tree-wide evolutionary patterns we calculated average net rates through time using the getRateThroughTimeMatrix function and plotted by the plotRateThroughTime function of the BAMM package.
Validation of BAMM estimates
There have been critics of the BAMM method recently90,91. Meyer and Wiens91 found that the method-of-moments estimator yielded stronger relationship between true and estimated diversification rates than BAMM, particularly in smaller clades. Therefore, to validate the BAMM results we used the method-of-moments estimator on clades with congruent core shifts. The method-of-moments approach estimates diversification rates from species richness and clade age while it can incorporate incomplete taxon sampling and extinction rate by assuming a relative extinction fraction (ε = extinction rate / speciation rate)92. We used the bd.ms function in geiger 2.0.6 (ref. 93). The analyses were performed on all the 85 clades with congruent core shifts, using three different relative extinction fractions (0, 0.45, 9) and calculating with both stem and crown ages. We calculated unsampled species fractions using the genus-specific sampling fractions we determined for BAMM analyses. For the same set of clades we calculated average net diversification and net speciation rates from BAMM data using the getCladeRates function. Then we performed linear regressions between method-of-moments estimates with different settings and average net diversification or net speciation rates from BAMM. We examined the adjusted r2 and the P values of the models to evaluate the correspondence between the two methods.
Detecting mass extinction events
We performed analyses using the compound Poisson process on mass extinction times model (CoMET)27,94, to examine tree-wide variation in diversification rate and occurrence of mass extinctions during the evolution of mushroom-forming fungi. First, we conducted a model comparison on ten chronograms using the CoMET model with a constant rate birth-death process implemented in the TESS 2.1.0. R package95. We compared models with and without mass extinction events on the basis of marginal likelihoods. We allowed the occurrence of a mass extinction event along the entire time span of a tree and we set the survival probability of species to 0.1 (corresponding to 10% of the species surviving a mass extinction event). The overall sampling fraction of species was set to 0.14. MCMC analyses were run for 20,000 generations and the first 2,000 posterior samples were discarded as a burn-in. Marginal likelihoods were estimated using stepping stone simulation using 100 stepping stones.
Then, we performed reversible jump Markov chain Monte Carlo (rjMCMC) analyses under the CoMET model to sample from the space of episodically varying birth-death processes with mass extinction events. In this analysis, we also tested the significance of the occurrence of a mass extinction event by performing model tests within time intervals. Bayes factor values BF > 10 or lnBF > 6 were considered strong support for a model, following Höhna et al.96 and Kass and Raftery97. We examined the sensitivity of the posterior probabilities to different prior settings in preliminary analyses of a randomly chosen tree. We examined models with two or ten expected mass extinction events, with 30, 100 or 270 expected rate changes and with survival probabilities of 0.001, 0.05 or 0.3. All of these preliminary analyses were run for 1.6 million generations with 100,000 generations as burn-in. We used a log-normal prior on speciation and extinction rates with a mean of 0.2 and 0.15 and a standard deviation of 0.5 and 0.5, respectively. In the final analyses we used all ten chronograms and an empirical hyper-prior on rate parameters on the basis of 200,000 iterations with 100,000 burn-in. The priors on the number of expected mass extinction and the expected rate changes were set to 2 and 30, respectively, on the basis of results of preliminary analyses. We set the survival probability to 0.05. Analyses were run for 3 million generations with 1 million generations as burn-in. The convergence of the analyses was checked by visually inspecting the log-likelihood values and by computing the effective sample size and the Geweke diagnostic for the log-likelihoods, the number of speciation rate shifts, the number of extinction rate shifts and the number of mass extinction events using the effectiveSize and the geweke.diag function of the CODA 0.19.-1 package88, respectively. To check the convergence of interval-specific parameters we used the tess.plot.singlechain.diagnostics function of the TESS package.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
New sequence data generated for this study were deposited at DDBJ/EMBL/GenBank under the accession numbers MK277471–MK278670 and MK299400–MK299412. Trees and alignments have been deposited in Dryad Digital Repository (accession number: doi:10.5061/dryad.gc2k9r). Genome assembly and annotation of C. micaceus FP101781, C. marcescibilis CBS121175, C. laeve CBS166.37, Dendrothele bispora CBS962.96, H. sulcata OMC1185, Peniophora sp. Cont, P. cervinus NL-1719, P. arcularius HBB13444 and P. gracilis CBS309.79 were deposited at DDBJ/EMBL/GenBank under the accession numbers QPFP00000000, QPFQ00000000, QPFR00000000, QPKH00000000, QPFL00000000, LOAU00000000, QPFM00000000, QPFN00000000 and QPFO00000000. All custom code is available from the authors upon request.
References
Jetz, W. & Pyron, R. A. The interplay of past diversification and evolutionary isolation with present imperilment across the amphibian tree of life. Nat. Ecol. Evol. 2, 850–858 (2018).
Jetz, W., Thomas, G. H., Joy, J. B., Hartmann, K. & Mooers, A. O. The global diversity of birds in space and time. Nature 491, 444–448 (2012).
Rabosky, D. L. et al. Rates of speciation and morphological evolution are correlated across the largest vertebrate radiation. Nat. Commun. 4, 1958 (2013).
Alfaro, M. E. et al. Explosive diversification of marine fishes at the Cretaceous–Palaeogene boundary. Nat. Ecol. Evol. 2, 688–696 (2018).
Alfaro, M. E. et al. Nine exceptional radiations plus high turnover explain species diversity in jawed vertebrates. Proc. Natl Acad. Sci. USA 106, 13410–13414 (2009).
Nagy, L. G. et al. The evolution of defense mechanisms correlate with the explosive diversification of autodigesting coprinellus mushrooms (Agaricales, fungi). Syst. Biol. 61, 595–607 (2012).
Wilson, A. W., Hosaka, K. & Mueller, G. M. Evolution of ectomycorrhizas as a driver of diversification and biogeographic patterns in the model mycorrhizal mushroom genus Laccaria. New Phytol. 213, 1862–1873 (2017).
Wilson, A. W., Binder, M. & Hibbett, D. S. Diversity and evolution of ectomycorrhizal host associations in the sclerodermatineae (Boletales, Basidiomycota). New Phytol. 194, 1079–1095 (2012).
Sánchez-Ramírez, S., Tulloss, R. E., Amalfi, M. & Moncalvo, J. M. Palaeotropical origins, boreotropical distribution and increased rates of diversification in a clade of edible ectomycorrhizal mushrooms (Amanita section Caesareae). J. Biogeogr. 42, 351–363 (2015).
Sánchez-Garcia, M. & Matheny, P. B. Is the switch to an ectomycorrhizal state an evolutionary key innovation in mushroom-forming fungi? A case study in the Tricholomatineae (Agaricales). Evolution 71, 51–65 (2017).
Wilson, A. W., Binder, M. & Hibbett, D. S. Effects of gasteroid fruiting body morphology on diversification rates in three independent clades of fungi estimated using binary state speciation and extinction analysis. Evolution 65, 1305–1322 (2011).
Hibbett, D. S. After the gold rush, or before the flood? Evolutionary morphology of mushroom-forming fungi (Agaricomycetes) in the early 21st century. Mycol. Res. 111, 1001–1018 (2007).
Hibbett, D. S. Trends in morphological evolution in homobasidiomycetes inferred using maximum likelihood: a comparison of binary and multistate approaches. Syst. Biol. 53, 889–903 (2004).
Hibbett, D. S. & Binder, M. Evolution of complex fruiting-body morphologies in homobasidiomycetes. Proc. Biol. Sci. 269, 1963–1969 (2002).
Matheny, P. B. et al. Major clades of Agaricales: a multilocus phylogenetic overview. Mycologia 98, 982–995 (2006).
Kirk, P, Cannon, P. & Minter, D. Dictionary of the Fungi (CABI International, Wallingford, UK, 2011).
Floudas, D. et al. The Paleozoic origin of enzymatic lignin decomposition reconstructed from 31 fungal genomes. Science 336, 1715–1719 (2012).
Kohler, A. et al. Convergent losses of decay mechanisms and rapid turnover of symbiosis genes in mycorrhizal mutualists. Nat. Genet. 47, 410–415 (2015).
Lutzoni, F. et al. Contemporaneous radiations of fungi and plants linked to symbiosis. Nat. Commun. 9, 5451 (2018).
Rabosky, D. L. Automatic detection of key innovations, rate shifts, and diversity-dependence on phylogenetic trees. PLoS ONE 9, e89543 (2014).
McKenna, D. D., Sequeira, A. S., Marvaldi, A. E. & Farrell, B. D. Temporal lags and overlap in the diversification of weevils and flowering plants. Proc. Natl Acad. Sci. USA 106, 7083–7088 (2009).
Looney, B. P., Ryberg, M., Hampe, F., Sánchez-García, M. & Matheny, P. B. Into and out of the tropics: global diversification patterns in a hyperdiverse clade of ectomycorrhizal fungi. Mol. Ecol. 25, 630–647 (2016).
Krah, F.-S. et al. Evolutionary dynamics of host specialization in wood-decay fungi. BMC Evol. Biol. 18, 119 (2018).
Beaulieu, J. M. & O’Meara, B. C. Extinction can be estimated from moderately sized molecular phylogenies. Evolution 69, 1036–1043 (2015).
Rabosky, D. L. Challenges in the estimation of extinction from molecular phylogenies: a response to Beaulieu and O’Meara. Evolution 70, 218–228 (2016).
Rabosky, D. L. Extinction rates should not be estimated from molecular phylogenies. Evolution 64, 1816–1824 (2010).
May, M. R., Hohna, S. & Moore, B. R. A Bayesian approach for detecting the impact of mass-extinction events on molecular phylogenies when rates of lineage diversification may vary. Methods Ecol. Evol. 7, 947–959 (2016).
Tennant, J. P., Mannion, P. D., Upchurch, P., Sutton, M. D. & Price, G. D. Biotic and environmental dynamics through the late jurassic-early cretaceous transition: evidence for protracted faunal and ecological turnover. Biol. Rev. 92, 776–814 (2017).
Casadevall, A. Fungi and the rise of mammals. PLoS Pathog. 8, e1002808 (2012).
Vajda, V. & McLoughlin, S. Fungal proliferation at the Cretaceous–Tertiary boundary. Science 303, 1489–1489 (2004).
Mittelbach, G. G. et al. Evolution and the latitudinal diversity gradient: speciation, extinction and biogeography. Ecology Lett. 10, 315–331 (2007).
Peay, K. G., Kennedy, P. G. & Talbot, J. M. Dimensions of biodiversity in the Earth mycobiome. Nat. Rev. Microbiol. 14, 434–447 (2016).
Shi, L.-L. et al. Variation in forest soil fungal diversity along a latitudinal gradient. Fungal Divers. 64, 305–315 (2014).
Tedersoo, L. et al. Global diversity and geography of soil fungi. Science 346, 1256688–1256688 (2014).
Sánchez-Ramírez, S., Etienne, R. S. & Moncalvo, J. M. High speciation rate at temperate latitudes explains unusual diversity gradients in a clade of Ectomycorrhizal fungi. Evolution 69, 2196–2209 (2015).
Gavrilets, S. & Losos, J. B. Adaptive radiation: contrasting theory with data. Science 323, 732–737 (2009).
Givnish, T. J. Adaptive radiation versus ‘radiation’ and ‘explosive diversification’: why conceptual distinctions are fundamental to understanding evolution. New Phytol. 207, 297–303 (2015).
Maddison, W. P., Midford, P. E. & Otto, S. P. Estimating a binary character’s effect on speciation and extinction. Syst. Biol. 56, 701–710 (2007).
Fitzjohn, R. G. Diversitree: Comparative phylogenetic analyses of diversification in R. Methods Ecol. Evol. 3, 1084–1092 (2012).
Berendse, F. & Scheffer, M. The angiosperm radiation revisited, an ecological explanation for Darwin’s ‘abominable mystery’. Ecol. Lett 12, 865–872 (2009).
Niklas, K. J., Tiffney, B. H. & Knoll, A. H. Patterns in vascular land plant diversification. Nature 303, 614–616 (1983).
Berner, R. A. Phanerozoic atmospheric oxygen: new results using the GEOCARBSULF model. Am. J. Sci. 309, 603–606 (2009).
Staden, R. The Staden sequence analysis package. Mol. Biotechnol. 5, 233–241 (1996).
Species Fungorum (CABI/Royal Botanic Gardens Kew, 2018).
Knudsen, H. & Vesterholt, J. Funga Nordica: Agaricoid, Boletoid and Cyphelloid Genera (Nordsvamp:Copenhagen, 2008).
Loytynoja, A. & Goldman, N. Phylogeny-aware gap placement prevents errors in sequence alignment and evolutionary analysis. Science 320, 1632–1635 (2008).
Loytynoja, A. & Goldman, N. From The Cover: an algorithm for progressive multiple alignment of sequences with insertions. Proc. Natl Acad. Sci. USA 102, 10557–10562 (2005).
Tóth, A. et al. Iteratively refined guide trees help improving alignment and phylogenetic inference in the mushroom family bolbitiaceae. PLoS ONE 8, e56143 (2013).
Gnerre, S. et al. High-quality draft assemblies of mammalian genomes from massively parallel sequence data. Proc. Natl Acad. Sci. USA 108, 1513–1518 (2011).
Martin, J. et al. Rnnotator: an automated de novo transcriptome assembly pipeline from stranded RNA-Seq reads. BMC Genomics 11, 663 (2010).
Chin, C.-S. et al. Phased diploid genome assembly with single-molecule real-time sequencing. Nat. Methods 13, 1050–1054 (2016).
Lam, K.-K., LaButti, K., Khalak, A. & Tse, D. FinisherSC: a repeat-aware tool for upgrading de novo assembly using long reads. Bioinformatics 31, 3207–3209 (2015).
Grigoriev, I. V. et al. MycoCosm portal: gearing up for 1000 fungal genomes. Nucleic Acids Res. 42(699), 704–(2014).
Darling, A., Carey, L. & Feng, W. The design, implementation, and evaluation of mpiBLAST. in ClusterWorld Conference & Expo and the 4th International Conference on Linux Clusters: The HPC Revolution 2003. LA-UR 03-2862 (2003).
van Dongen, S. Graph Clustering by Flow Simulation. PhD thesis, Univ. Utrecht (2000).
Talavera, G. & Castresana, J. Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst. Biol. 56, 564–577 (2007).
Nagy, L. G. et al. Comparative genomics of early-diverging mushroom-forming fungi provides insights into the origins of lignocellulose decay capabilities. Mol. Biol. Evol. 33, 959–970 (2016).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Schliep, K. P. phangorn: phylogenetic analysis in R. Bioinformatics 27, 592–593 (2011).
Lartillot, N., Lepage, T. & Blanquart, S. PhyloBayes 3: a Bayesian software package for phylogenetic reconstruction and molecular dating. Bioinformatics 25, 2286–2288 (2009).
Åkerborg, Ö., Sennblad, B. & Lagergren, J. Birth-death prior on phylogeny and speed dating. BMC Evol. Biol. 8, 77 (2008).
Smith, S. Y., Currah, R. S. & Stockey, R. A. Cretaceous and eocene poroid hymenophores from Vancouver Island, British Columbia. Mycologia 96, 180–186 (2004).
Hibbett, D. S., Grimaldi, D. & Donoghue, M. J. Fossil mushrooms from Miocene and Cretaceous ambers and the evolution of homobasidiomycetes. Am. J. Bot. 84, 981–991 (1997).
Poinar, G. Bird’s nest fungi (Nidulariales: Nidulariaceae) in Baltic and Dominican amber. Fungal Biol 118, 325–329 (2014).
BrownR. W A bracket fungus from the late Tertiary of southwestern Idaho. J. Washington Acad. Sci. 30, 422–424 (1940).
Magallon-Puebla, S. & Cevallos-Ferriz, S. R. S. A fossil earthstar (Geasteraceae; Gasteromycetes) from the late cenozoic of puebla, mexico. Am. J. Bot. 80, 1162–1167 (1993).
Near, T. J., Meylan, P. A. & Shaffer, H. B. Assessing concordance of fossil calibration points in molecular clock studies: an example using turtles. Am. Nat. 165, 137–146 (2005).
Charif, D. & Lobry, J. R. SeqinR 1.0-2: A Contributed Package to the R Project for Statistical Computing Devoted to Biological Sequences Retrieval and Analysis (Springer, Berlin, 2007); https://doi.org/10.1007/978-3-540-35306-5_10
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 1–9 (2009).
Sanderson, M. J. Estimating absolute rates of molecular evolution and divergence times: a penalized likelihood approach. Mol. Biol. Evol. 19, 101–109 (2002).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Thorne, J. L., Kishino, H. & Painter, I. S. Estimating the rate of evolution of the rate of molecular evolution. Mol. Biol. Evol. 15, 1647–1657 (1998).
Bodensteiner, P., Binder, M., Moncalvo, J. M., Agerer, R. & S Hibbett, D. Phylogenetic relationships of cyphelloid homobasidiomycetes. Mol. Phylogenet. Evol. 33, 501–515 (2004).
Holt, B. G. et al. An update of Wallace’s zoogeographic regions of the world. Science 339, 74–78 (2013).
R: A Language and Environment for Statistical Computing (R Core Team, 2018).
Fitzjohn, R. G. Quantitative traits and diversification. Syst. Biol. 59, 619–633 (2010).
Goldberg, E. E., Lancaster, L. T. & Ree, R. H. Phylogenetic inference of reciprocal effects between geographic range evolution and diversification. Syst. Biol. 60, 451–465 (2011).
Pagel, M. & Meade, A. BayesTraits v.3.0.1 (Reading Evolutionary Biology Group, 2007).
Pagel, M. Inferring the historical patterns of biological evolution. Nature 401, 877–884 (1999).
Xie, W., Lewis, P. O., Fan, Y., Kuo, L. & Chen, M. H. Improving marginal likelihood estimation for bayesian phylogenetic model selection. Syst. Biol. 60, 150–160 (2011).
Revell, L. J. phytools: an R package for phylogenetic comparative biology (and other things). Methods Ecol. Evol. 3, 217–223 (2012).
Huelsenbeck, J. P., Nielsen, R. & Bollback, J. P. Stochastic mapping of morphological characters. Syst. Biol. 52, 131–158 (2003).
Wickham, H. Ggplot2. Elegant Graphics for Data Analysis (Springer, New York, 2009); https://doi.org/10.1007/978-0-387-98141-3
Paradis, E., Claude, J. & Strimmer, K. APE: Analyses of phylogenetics and evolution in R language. Bioinformatics 20, 289–290 (2004).
Fitzjohn, R. G., Maddison, W. P. & Otto, S. P. Estimating trait-dependent speciation and extinction rates from incompletely resolved phylogenies. Syst. Biol. 58, 595–611 (2009).
Rabosky, D. L. et al. BAMMtools: an R package for the analysis of evolutionary dynamics on phylogenetic trees. Methods Ecol. Evol. 5, 701–707 (2014).
Geweke, J. Evaluating the accuracy of sampling-based approaches to the calculation of posterior moments. Bayesian Stat. 4, 169–193 (1992).
Plummer, M., Best, N., Cowles, K. & Vines, K. CODA: convergence diagnosis and output analysis for MCMC. R News 6, 7–11 (2006).
Nylander, J. A. A., Wilgenbusch, J. C., Warren, D. L. & Swofford, D. L. AWTY (are we there yet?): a system for graphical exploration of MCMC convergence in Bayesian phylogenetics. Bioinformatics 24, 581–583 (2008).
Moore, B. R., Höhna, S., May, M. R., Rannala, B. & Huelsenbeck, J. P. Critically evaluating the theory and performance of Bayesian analysis of macroevolutionary mixtures. Proc. Natl Acad. Sci. USA 113, 9569–9574 (2016).
MeyerA. L. S. & WiensJ. J. Estimating diversification rates for higher taxa: BAMM can give problematic estimate of rates and rate shifts. Evolution 72, 1–15 (2017).
Magallon, S. & Sanderson, M. J. Absolute diversification rates in angiosperms clades. Evolution 55, 1762–1780 (2001).
Harmon, L. J., Weir, J. T., Brock, C. D., Glor, R. E. & Challenger, W. GEIGER: investigating evolutionary radiations. Bioinformatics 24, 129–131 (2008).
Höhna, S. The time-dependent reconstructed evolutionary process with a key-role for mass-extinction events. J. Theor. Biol. 380, 321–331 (2015).
Höhna, S. Fast simulation of reconstructed phylogenies under global time-dependent birth-death processes. Bioinformatics 29, 1367–1374 (2013).
Hohna, S., May, M. R. & Moore, B. R. Phylogeny Simulation and Diversification Rate Analysis with TESS; https://cran.r-project.org/web/packages/TESS/vignettes/Bayesian_Diversification_Rate_Analysis.pdf 1–98 (2015).
Kass, R. E. & Raftery, A. E. Bayes factors. J. Am. Stat. Assoc. 90, 773–795 (1995).
Acknowledgements
We want to thank I. Nagy for help in writing a custom java program to retrieve species from the Species Fungorum database. This work has received funding from the Momentum Program of the Hungarian Academy of Sciences (contract no. LP2014/12, to L.G.N.) and from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (grant agreement no. 758161). The work by the U.S. Department of Energy Joint Genome Institute, a DOE Office of Science User Facility, is supported by the Office of Science of the U.S. Department of Energy under contract no. DE-AC02-05CH11231. T.V. was supported by the National Talent Program (contract no. NTP-NFTÖ-17-B-0337), by the UNKP-18-3 New National Excellence Program of the Ministry of Human Capacities and by the Straub Young Scientist scholarship from the Biological Research Centre, Hungarian Academy of Sciences. The support of V.A. was provided to the Moravian Museum by the Ministry of Culture of the Czech Republic (DKRVO, ref. MK000094862).
Author information
Authors and Affiliations
Contributions
B.D., J.G.S., B.K., J.U., S.K. and V.B. sequenced specimens, W.A, K.W.B., M.C., S.H., K.L.B., A.L., S.M., R.R., F.M.M. and I.V.G. sequenced, assembled, and annotated genomes. K.K., T.V. J.G.S. and L.G.N made multiple alignment, K.K. performed phylogenomic analyses. G.S. and T.V. conducted molecular dating. A.F.J., A.S., A.S., B.B., B.D., B.E.L., B.O.S., C.A., C.F., C.O., C.S., C.V., D.D., D.K., H.K., I.K., J.C., J.G., K.H., K.L., K.S., L.A., L.M., M.E.N., M.T., N.L.B., N.R., O.M., P.B., P.C., P.F., R.E.T., S.K., S.S.R., T.N., T.P., V.A., V.S., W.D. and Z.L. contributed specimens or sequences. K.K., T.V., B.D., V.P., V.B., C.F. and O.M. worked on character coding of species. S.S.R. tested the LDG hypothesis. M.S.G. performed BiSSE analyes. C.F. and T.V. conducted BayesTraits analyses. All other analyses were performed by T.V. L.G.N., K.K., T.V., M.S.G., S.S.R. and D.S.H. wrote the article. L.G.N. supervised the project. All authors read and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Notes 1–8; Supplementary Figures 1–10; Supplementary Tables 1–8; Captions for Supplementary Data 1–7
Supplementary Data 1
Data matrices of GenBank accession numbers of newly sequenced taxa, character states used for analyses of character evolution and taxon sampling statistics used in this study.
Supplementary Data 2
Comparison of results of different molecular dating analyses.
Supplementary Data 3
Transition rate matrices of ancestral character-state reconstruction (ASR) analyses and results of the ASR of fungal substrate preference.
Supplementary Data 4
Speciation, extinction and diversification rates through time obtained by iteratively removing orders and higher level clades from the phylogeny.
Supplementary Data 5
List of highly supported core shifts that are congruent across multiple trees.
Supplementary Data 6
Lineages through time (LTT) plots for the clades containing highly supported core shifts.
Supplementary Data 7
Parameters and detailed results of BayesTraits and BiSSE analyses.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Varga, T., Krizsán, K., Földi, C. et al. Megaphylogeny resolves global patterns of mushroom evolution. Nat Ecol Evol 3, 668–678 (2019). https://doi.org/10.1038/s41559-019-0834-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-019-0834-1
This article is cited by
-
Heterostigmatic mites (Acari: Prostigmata) associated with mushroom-forming fungi (Basidiomycota: Agaricomycetes), with description of a new species of microdispid mite (Microdispidae)
Biologia (2024)
-
Taxonomic characterisation of the Regianum clade (genus Tuber) and the trait evolution of spore size among true truffles
Mycological Progress (2024)
-
Pleurotus ostreatus as a model mushroom in genetics, cell biology, and material sciences
Applied Microbiology and Biotechnology (2024)
-
Mycological surveys reveal two new species of genus Lepiota (Agaricaceae) from the semi-arid climatic region of Punjab, Pakistan
Mycological Progress (2024)
-
Two new Agaricomycetes related to post-fire mosses
Mycological Progress (2024)