Abstract
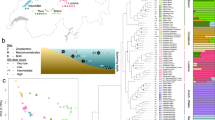
The interplay between divergence and phenotypic plasticity is critical to our understanding of a species’ adaptive potential under rapid climate changes. We investigated divergence and plasticity in natural populations of the Pacific oyster Crassostrea gigas with a congeneric oyster Crassostrea angulata from southern China used as an outgroup. Genome re-sequencing of 371 oysters revealed unexpected genetic divergence in a small area that coincided with phenotypic divergence in growth, physiology, heat tolerance and gene expression across environmental gradients. These findings suggest that selection and local adaptation are pervasive and, together with limited gene flow, influence population structure. Genes showing sequence differentiation between populations also diverged in transcriptional response to heat stress. Plasticity in gene expression is positively correlated with evolved divergence, indicating that plasticity is adaptive and favoured by organisms under dynamic environments. Divergence in heat tolerance—partly through acetylation-mediated energy depression—implies differentiation in adaptive potential. Trade-offs between growth and survival may play an important role in local adaptation of oysters and other marine invertebrates.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The whole-genome re-sequencing and transcriptome datasets were deposited in the Sequence Read Archive (SRA) database under the accession number PRJNA394055. The proteome and acetylome data are available via the ProteomeXchange Consortium with the identifier PXD008057.
References
Sanford, E. & Kelly, M. W. Local adaptation in marine invertebrates. Annu. Rev. Mar. Sci. 3, 509–535 (2011).
Somero, G. N. The physiology of global change: linking patterns to mechanisms. Annu. Rev. Mar. Sci. 4, 39–61 (2012).
Bozinovic, F., Calosi, P. & Spicer, J. I. Physiological correlates of geographic range in animals. Annu. Rev. Ecol. Evol. Syst. 42, 155–179 (2011).
Nadeau, C. P., Urban, M. C. & Bridle, J. R. Climates past, present, and yet-to-come shape climate change vulnerabilities. Trends Ecol. Evol. 32, 786–800 (2017).
Conover, D. O., Clarke, L. M., Munch, S. B. & Wagner, G. N. Spatial and temporal scales of adaptive divergence in marine fishes and the implications for conservation. J. Fish Biol. 69, 21–47 (2006).
Zhang, G. et al. Molecular basis for adaptation of oysters to stressful marine intertidal environments. Annu. Rev. Anim. Biosci. 4, 357–381 (2016).
Hice, L. A., Duffy, T. A., Munch, S. B. & Conover, D. O. Spatial scale and divergent patterns of variation in adapted traits in the ocean. Ecol. Lett. 15, 568–575 (2012).
Burford, M. O., Scarpa, J., Cook, B. J. & Hare, M. P. Local adaptation of a marine invertebrate with a high dispersal potential: evidence from a reciprocal transplant experiment of the eastern oyster Crassostrea virginica. Mar. Ecol. Prog. Ser. 505, 161–175 (2014).
Hebert, A. S. et al. Calorie restriction and SIRT3 trigger global reprogramming of the mitochondrial protein acetylome. Mol. Cell 49, 186–199 (2013).
Pfennig, D. W. et al. Phenotypic plasticity’s impacts on diversification and speciation. Trends Ecol. Evol. 25, 459–467 (2010).
Agrawal, A. A. Phenotypic plasticity in the interactions and evolution of species. Science 294, 321–326 (2001).
Fischer, E. K., Ghalambor, C. K. & Hoke, K. L. Can a network approach resolve how adaptive vs nonadaptive plasticity impacts evolutionary trajectories? Integr. Comp. Biol. 56, 877–888 (2016).
Ghalambor, C. K. et al. Non-adaptive plasticity potentiates rapid adaptive evolution of gene expression in nature. Nature 525, 372–375 (2015).
Schneider, R. F. & Meyer, A. How plasticity, genetic assimilation and cryptic genetic variation may contribute to adaptive radiations. Mol. Ecol. 26, 330–350 (2017).
Schaum, C. E. & Collins, S.Plasticity predicts evolution in a marine alga. Proc. R. Soc. B 281, 20141486 (2014).
Calosi, P., De Wit, P., Thor, P. & Dupont, S. Will life find a way? Evolution of marine species under global change. Evol. Appl. 9, 1035–1042 (2016).
Bay, R. A. et al. Predicting responses to contemporary environmental change using evolutionary response architectures. Am. Nat. 189, 463–473 (2017).
Chevin, L. M., Lande, R. & Mace, G. M. Adaptation, plasticity, and extinction in a changing environment: towards a predictive theory. PLoS Biol. 8, e1000357 (2010).
Price, T. D., Qvarnstrom, A. & Irwin, D. E. The role of phenotypic plasticity in driving genetic evolution. Proc. R. Soc. Lond. B 270, 1433–1440 (2003).
Markov, A. V. & Ivnitsky, S. B. Evolutionary role of phenotypic plasticity. Moscow Univ. Biol. Sci. Bull. 71, 185–192 (2016).
Boyd, P. W. et al. Biological responses to environmental heterogeneity under future ocean conditions. Glob. Change Biol. 22, 2633–2650 (2016).
Dineshram, R. et al. Quantitative analysis of oyster larval proteome provides new insights into the effects of multiple climate change stressors. Glob. Change Biol. 22, 2054–2068 (2016).
Zhang, G. et al. The oyster genome reveals stress adaptation and complexity of shell formation. Nature 490, 49–54 (2012).
Wang, H., Zhang, G., Liu, X. & Guo, X. Classification of common oysters from north China. J. Shellfish Res. 27, 495–503 (2008).
Ju, X. & Xiong, X. Distributions and seasonal changes of water temperature in the Bohai Sea, Yellow Sea and East China Sea.Adv. Mar. Sci. 31, 55–68 (2013).
Weng, X., Zhang, Q., Zhang, Y. & Yang, Y. Characteristics of the daily variations of the temperature in the Bohai Sea, Yellow Sea and East China Sea. Mar. Sci. 6, 49–54 (1993).
Guo, X., He, Y., Zhang, L., Lelong, C. & Jouaux, A. Immune and stress responses in oysters with insights on adaptation. Fish Shellfish Immunol. 46, 107–119 (2015).
Zhang, L. & Guo, X. Development and validation of single nucleotide polymorphism markers in the eastern oyster Crassostrea virginica Gmelin by mining ESTs and resequencing. Aquaculture 302, 124–129 (2010).
Qi, H. et al. Construction and evaluation of a high-density SNP array for the Pacific oyster (Crassostrea gigas). PLoS ONE 12, e0174007 (2017).
Guan, B. & Mao, H. A note on circulation of the East China Sea. Chin. J. Oceanol. Limnol. 1, 5–16 (1982).
Zhan, A. et al. Fine-scale population genetic structure of Zhikong scallop (Chlamys farreri): do local marine currents drive geographical differentiation? Mar. Biotechnol. 11, 223–235 (2009).
Kong, L., Bai, J. & Li, Q. Comparative assessment of genomic SSR, EST–SSR and EST–SNP markers for evaluation of the genetic diversity of wild and cultured Pacific oyster, Crassostrea gigas Thunberg. Aquaculture 420–421, S85–S91 (2014).
Li, Q., Yu, H. & Yu, R. Genetic variability assessed by microsatellites in cultured populations of the Pacific oyster (Crassostrea gigas) in China. Aquaculture 259, 95–102 (2006).
Qin, Y., Zhao, Y. & Zhao, S. Geology of the Bohai Sea (Science Press, Beijing, 1985).
Moreau, P. et al. Autophagy plays an important role in protecting Pacific oysters from OsHV-1 and Vibrio aestuarianus infections. Autophagy 11, 516–526 (2015).
Duncan, R. E. & Hershey, J. W. B. Protein-synthesis and protein-phosphorylation during heat-stress, recovery, and adaptation. J. Cell Biol. 109, 1467–1481 (1989).
Helmuth, B. et al. Climate change and latitudinal patterns of intertidal thermal stress. Science 298, 1015–1017 (2002).
Sussarellu, R. et al. Molecular and cellular response to short-term oxygen variations in the Pacific oyster Crassostrea gigas. J. Exp. Mar. Biol. Ecol. 412, 87–95 (2012).
Guévélou, E. et al. Regulation of a truncated isoform of AMP-activated protein kinase α (AMPKα) in response to hypoxia in the muscle of Pacific oyster Crassostrea gigas. J. Comp. Physiol. B 183, 597–611 (2013).
Pan, T.-C. F., Applebaum, S. L. & Manahan, D. T. Experimental ocean acidification alters the allocation of metabolic energy. Proc. Natl Acad. Sci. USA 112, 4696–4701 (2015).
Falfushynska, H. I., Phan, T. & Sokolova, I. M. Long-term acclimation to different thermal regimes affects molecular responses to heat stress in a freshwater clam Corbicula fluminea. Sci. Rep. 6, 39476 (2016).
Roy, K., Jablonski, D. & Martien, K. K. Invariant size–frequency distributions along a latitudinal gradient in marine bivalves. Proc. Natl Acad. Sci. USA 97, 13150–13155 (2000).
Sussarellu, R. et al. Oyster reproduction is affected by exposure to polystyrene microplastics. Proc. Natl Acad. Sci. USA 113, 2430–2435 (2016).
Tomanek, L. Proteomics to study adaptations in marine organisms to environmental stress. J. Proteomics 105, 92–106 (2014).
De Wit, P., Dupont, S. & Thor, P. Selection on oxidative phosphorylation and ribosomal structure as a multigenerational response to ocean acidification in the common copepod Pseudocalanus acuspes. Evol. Appl. 9, 1112–1123 (2016).
Pörtner, H. O. Oxygen- and capacity-limitation of thermal tolerance: a matrix for integrating climate-related stressor effects in marine ecosystems. J. Exp. Biol. 213, 881–893 (2010).
Schulte, P. M. The effects of temperature on aerobic metabolism: towards a mechanistic understanding of the responses of ectotherms to a changing environment. J. Exp. Biol. 218, 1856–1866 (2015).
Sokolova, I. M., Frederich, M., Bagwe, R., Lannig, G. & Sukhotin, A. A. Energy homeostasis as an integrative tool for assessing limits of environmental stress tolerance in aquatic invertebrates. Mar. Environ. Res. 79, 1–15 (2012).
Jombart, T., Devillard, S. & Balloux, F. Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet. 11, 94 (2010).
Siljestam, M. & Ostman, O. The combined effects of temporal autocorrelation and the costs of plasticity on the evolution of plasticity. J. Evol. Biol. 30, 1361–1371 (2017).
DeWitt, T. J., Sih, A. & Wilson, D. S. Costs and limits of phenotypic plasticity. Trends Ecol. Evol. 13, 77–81 (1998).
Ghalambor, C. K., McKay, J. K., Carroll, S. P. & Reznick, D. N. Adaptive versus non-adaptive phenotypic plasticity and the potential for contemporary adaptation in new environments. Funct. Ecol. 21, 394–407 (2007).
Thor, P. & Dupont, S. Transgenerational effects alleviate severe fecundity loss during ocean acidification in a ubiquitous planktonic copepod. Glob. Change Biol. 21, 2261–2271 (2015).
Li, Z., Li, X., Wang, Z., Shen, Q. W. & Zhang, D. Antemortem stress regulates protein acetylation and glycolysis in postmortem muscle. Food Chem. 202, 94–98 (2016).
Li, A., Li, L., Song, K., Wang, W. & Zhang, G. Temperature, energy metabolism, and adaptive divergence in two oyster subspecies. Ecol. Evol. 7, 6151–6162 (2017).
Guo, X., Li, Q., Wang, Q. Z. & Kong, L. F. Genetic mapping and QTL analysis of growth-related traits in the Pacific oyster. Mar. Biotechnol. 14, 218–226 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Li, H. et al. The sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Watterson, G. A. On the number of segregating sites in genetical models without recombination. Theor. Popul. Biol. 7, 256–276 (1975).
Hartl, D. L. The molecular approach to evolution: molecular evolutionary genetics. Science 237, 782 (1987).
Vilella, A. J., Blanco-Garcia, A., Hutter, S. & Rozas, J. VariScan: analysis of evolutionary patterns from large-scale DNA sequence polymorphism data. Bioinformatics 21, 2791–2793 (2005).
Barrett, J. C., Fry, B., Maller, J. & Daly, M. J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265 (2005).
Wang, W. et al. Development of calibration models for rapid determination of chemical composition of Pacific oyster (Crassostrea gigas) by near infrared reflectance spectroscopy. J. Shellfish Res. 34, 303–309 (2015).
Freitas, V. et al. Temperature tolerance and energetics: a dynamic energy budget-based comparison of North Atlantic marine species. Phil. Trans. R. Soc. B 365, 3553–3565 (2010).
Zhu, Q., Zhang, L., Li, L., Que, H. & Zhang, G. Expression characterization of stress genes under high and low temperature stresses in the Pacific oyster, Crassostrea gigas. Mar. Biotechnol. 18, 176–188 (2016).
Meng, J. et al. Genome and transcriptome analyses provide insight into the euryhaline adaptation mechanism of Crassostrea gigas. PLoS ONE 8, e58563 (2013).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
R Development Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2016).
Hadfiel, J. D. MCMC methods for multi-response generalized linear mixed models: the MCMCglmm R package. J. Stat. Softw. 33, 1–22 (2010).
Acknowledgements
G.Z. and L.L. are supported by the National Natural Science Foundation of China (31530079 to G.Z. and 31572620 to L.L.). G.Z. is supported by the Strategic Priority Research Program of the ‘Western Pacific Ocean System: Structure, Dynamics and Consequences’ project (XDA 11020305), Blue Life Breakthrough Program of LMBB (MS2018NO02) of Qingdao National Laboratory for Marine Science and Technology, and Modern Agro-industry Technology Research System (CARS-49). X.G. is supported by the ‘Taishan Overseas Scholar Program’ of Shandong and USDA/NJAES project 1004475/NJ32920. We thank J. Yan for suggestions on the experimental design and data analyses, Q. Li, Z. Yu, C. Ke, Z. Zeng, Y. Ning and Y. Bao for sampling collection, and B. Yin and J. Qi for information on marine currents.
Author information
Authors and Affiliations
Contributions
L.L. and G.Z. conceived the study and designed the major scientific objectives. X.G. participated in the final data analysis and interpretation. L.L., W.W., H. Que, F.W., H. Qi, F.X., R.C., B.H., S. Zhang and Y. Li. participated in the collection of wild oysters. L.L., W.W., H. Qi., J.M., S.Z., C.L., T.W. and A.L. participated in oyster breeding for F1 and F2 generations. A.L., X.T., S.Liu, B.L., R.S. and Y. Liu. participated in larval rearing and adult management. J.M., C.L., T.W., X.T. and A.L. conducted DNA extraction and sequencing library preparation. K.S., S.Li, C.Z. and W.H. performed the genome sequencing. K.S., S.Li, C.Z., W.H., S. Zhao and A.L. contributed to the re-sequencing data analysis. A.L. and X.T. participated in the morphological measurements and sample collection for physiological and cellular determinations. A.L. conducted the laboratory experiments for measuring physiological and cellular parameters, as well as data analysis. A.L. conducted messenger RNA extraction for transcriptome and data analysis, and protein extraction for proteome and acetylome analysis. A.L. and C.B. contributed to data analysis of the proteome and acetylome. H.L. collected data on the monthly average SST. L.L., A.L., K.S., X.G., P.D.W., J.M., L.Z. and G.Z. did most of the writing and revision, with input from all authors. All authors approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Information
Supplementary Figures and Tables
Supplementary Table 3
Mapping statistics for Pacific oyster samples resequenced
Supplementary Table 4
SNP statistics in different genomic regions for each Pacific oyster sample
Supplementary Table 8
Genes from genomic regions under selection in Bohai and southern Yellow Seas and their FST values
Supplementary Table 9
GO enrichment analysis of genes from regions under selection in Bohai and southern Yellow Seas
Supplementary Table 10
Annotation of genes from regions under selection in Bohai and southern Yellow Seas
Supplementary Table 11
Summary statistics of transcriptome dataset
Supplementary Table 12
Genes exhibiting significant transcriptional response at 6 and 24 hr of heat shock in oysters from JZ and QD
Supplementary Table 13
Summary statistics of proteome and acetylome datasets
Supplementary Table 14
Numbers of proteins and acetyl sites showing differential abundance
Supplementary Table 16
Genes exhibiting both significantly differential expression in response to high temperature and differential plasticity at 6 and 24 hr in oysters from JZ and QD
Supplementary Table 17
Acetyl sites exhibiting differential plasticity in proteins of major energy metabolic pathways when oysters were exposed to high temperature for 6 and 24 hr
Supplementary Table 18
Expression level of proteins corresponding to acetyl sites exhibiting differential plasticity in key energy metabolic pathways
Rights and permissions
About this article
Cite this article
Li, L., Li, A., Song, K. et al. Divergence and plasticity shape adaptive potential of the Pacific oyster. Nat Ecol Evol 2, 1751–1760 (2018). https://doi.org/10.1038/s41559-018-0668-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-018-0668-2
This article is cited by
-
Estimating microhaplotype allele frequencies from low-coverage or pooled sequencing data
BMC Bioinformatics (2023)
-
Genome-wide sequencing identifies a thermal-tolerance related synonymous mutation in the mussel, Mytilisepta virgata
Communications Biology (2023)
-
Reproductive Phenology of the Eastern Oyster, Crassostrea virginica (Gmelin, 1791), Along a Temperate Estuarine Salinity Gradient
Estuaries and Coasts (2023)
-
Coping with harsh heat environments: molecular adaptation of metabolic depression in the intertidal snail Echinolittorina radiata
Cell Stress and Chaperones (2023)
-
Genomic Landscape of Rare Codon Usage at Start Region in the Pacific Oyster Genome
Journal of Ocean University of China (2023)