Abstract
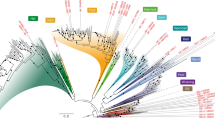
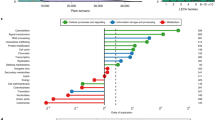
Establishing a unified timescale for the early evolution of Earth and life is challenging and mired in controversy because of the paucity of fossil evidence, the difficulty of interpreting it and dispute over the deepest branching relationships in the tree of life. Surprisingly, it remains perhaps the only episode in the history of life where literal interpretations of the fossil record hold sway, revised with every new discovery and reinterpretation. We derive a timescale of life, combining a reappraisal of the fossil material with new molecular clock analyses. We find the last universal common ancestor of cellular life to have predated the end of late heavy bombardment (>3.9 billion years ago (Ga)). The crown clades of the two primary divisions of life, Eubacteria and Archaebacteria, emerged much later (<3.4 Ga), relegating the oldest fossil evidence for life to their stem lineages. The Great Oxidation Event significantly predates the origin of modern Cyanobacteria, indicating that oxygenic photosynthesis evolved within the cyanobacterial stem lineage. Modern eukaryotes do not constitute a primary lineage of life and emerged late in Earth’s history (<1.84 Ga), falsifying the hypothesis that the Great Oxidation Event facilitated their radiation. The symbiotic origin of mitochondria at 2.053–1.21 Ga reflects a late origin of the total-group Alphaproteobacteria to which the free living ancestor of mitochondria belonged.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Dos Reis, M., Donoghue, P. C. J. & Yang, Z. Bayesian molecular clock dating of species divergences in the genomics era. Nat. Rev. Genet. 17, 71–80 (2016).
Wacey, D. Early Life on Earth: a Practical Guide Vol. 31 (Springer, New York, 2009).
Inoue, J., Donoghue, P. C. J. & Yang, Z. The impact of the representation of fossil calibrations on Bayesian estimation of species divergence times. Syst. Biol. 59, 74–89 (2009).
Parham, J. F. et al. Best practices for justifying fossil calibrations. Syst. Biol. 61, 346–359 (2012).
Warnock, R. C. M., Parham, J. F., Joyce, W. G., Lyson, T. R. & Donoghue, P. C. J. Calibration uncertainty in molecular dating analyses: there is no substitute for the prior evaluation of time priors. Proc. R. Soc. B 282, 20141013 (2014).
Davín, A. A. et al. Gene transfers can date the tree of life. Nat. Ecol. Evol. 2, 904–909 (2018).
Lozano-Fernandez, J., dos Reis, M., Donoghue, P. C. J. & Pisani, D. RelTime rates collapse to a strict clock when estimating the timeline of animal diversification. Genome Biol. Evol. 9, 1320–1328 (2017).
Pisani, D. & Liu, A. G. Animal evolution: only rocks can set the clock. Curr. Biol. 25, R1079–R1081 (2015).
Spang, A. et al. Complex archaea that bridge the gap between prokaryotes and eukaryotes. Nature 521, 173–179 (2015).
Williams, T. A., Foster, P. G., Cox, C. J. & Embley, T. M. An archaeal origin of eukaryotes supports only two primary domains of life. Nature 504, 231–236 (2013).
Dodd, M. S. et al. Evidence for early life in Earth’s oldest hydrothermal vent precipitates. Nature 543, 60–64 (2017).
Pflug, H. D. & Jaeschke-Boyer, H. Combined structural and chemical analysis of 3,800-Myr-old microfossils. Nature 280, 483–486 (1979).
Nutman, A. P., Bennett, V. C., Friend, C. R. L., Van Kranendonk, M. J. & Chivas, A. R. Rapid emergence of life shown by discovery of 3,700-million-year-old microbial structures. Nature 537, 535–538 (2016).
Rosing, M. T. 13C-depleted carbon microparticles in >3700-Ma sea-floor sedimentary rocks from West Greenland. Science 283, 674–676 (1999).
Mojzsis, S. J.et al. Evidence for life on Earth before 3,800 million years ago. Nature 384, 55–59 1996).
Van Zuilen, M. A., Lepland, A. & Arrhenius, G. Reassessing the evidence for the earliest traces of life. Nature 418, 627–630 (2002).
Horita, J. & Berndt, M. E. Abiogenic methane formation and isotopic fractionation under hydrothermal conditions. Science 285, 1055–1057 (1999).
Lepland, A., Arrhenius, G. & Cornell, D. Apatite in early Archean Isua supracrustal rocks, southern West Greenland: its origin, association with graphite and potential as a biomarker. Precambrian Res. 118, 221–241 (2002).
Sugitani, K. et al. Early evolution of large micro‐organisms with cytological complexity revealed by microanalyses of 3.4 Ga organic‐walled microfossils. Geobiology 13, 507–521 (2015).
Sugitani, K. et al. Biogenicity of morphologically diverse carbonaceous microstructures from the ca. 3400 Ma Strelley Pool Formation, in the Pilbara Craton, Western Australia. Astrobiology 10, 899–920 (2010).
Sugitani, K., Mimura, K., Nagaoka, T., Lepot, K. & Takeuchi, M. Microfossil assemblage from the 3400 Ma Strelley Pool Formation in the Pilbara Craton, Western Australia: results form a new locality. Precambrian Res. 226, 59–74 2013).
Wacey, D., Kilburn, M. R., Saunders, M., Cliff, J. & Brasier, M. D. Microfossils of sulphur-metabolizing cells in 3.4-billion-year-old rocks of Western Australia. Nat. Geosci. 4, 698–702 (2011).
Lepot, K. et al. Texture-specific isotopic compositions in 3.4 Gyr old organic matter support selective preservation in cell-like structures. Geochim. Cosmochim. Acta 112, 66–86 (2013).
Wacey, D., McLoughlin, N., Whitehouse, M. J. & Kilburn, M. R. Two coexisting sulfur metabolisms in a ca. 3400 Ma sandstone. Geology 38, 1115–1118 (2010).
Wacey, D. Stromatolites in the ~3400 Ma Strelley Pool Formation, Western Australia: examining biogenicity from the macro-to the nano-scale. Astrobiology 10, 381–395 (2010).
Abramov, O. & Mojzsis, S. J. Microbial habitability of the Hadean Earth during the late heavy bombardment. Nature 459, 419–422 (2009).
Butterfield, N. J. Bangiomorpha pubescens n. gen., n. sp.: implications for the evolution of sex, multicellularity, and the Mesoproterozoic/Neoproterozoic radiation of eukaryotes. Paleobiology 26, 386–404 (2000).
Sánchez-Baracaldo, P., Raven, J. A., Pisani, D. & Knoll, A. H. Early photosynthetic eukaryotes inhabited low-salinity habitats. Proc. Natl Acad. Sci. USA 114, E7737–E7745 (2017).
Bengston, S. et al. Three-dimensional preservation of cellular and subcellular structures suggests 1.6 billion-year-old crown-group red algae. PLoS Biol. 15, e2000735 (2017).
Lamb, D. M., Awramik, S. M., Chapman, D. J. & Zhu, S. Evidence for eukaryotic diversification in the ∼1800 million-year-old Changzhougou Formation, North China. Precambrian Res. 173, 93–104 (2009).
Zaremba-Niedzwiedzka, K. et al. Asgard archaea illuminate the origin of eukaryotic cellular complexity. Nature 541, 353–358 (2017).
Warnock, R. C. M., Yang, Z. & Donoghue, P. C. J. Exploring uncertainty in the calibration of the molecular clock. Biol. Lett. 8, 156–159 (2012).
Lanfear, R., Calcott, B., Ho, S. Y. W. & Guindon, S. PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol. Biol. Evol. 29, 1695–1701 (2012).
Woese, C. R. & Fox, G. E. Phylogenetic structure of the prokaryotic domain: the primary kingdoms. Proc. Natl Acad. Sci. USA 74, 5088–5090 (1977).
Drummond, A. J., Ho, S. Y. W., Phillips, M. J. & Rambaut, A. Relaxed phylogenetics and dating with confidence. PLoS Biol. 4, e88 (2006).
Chapman, C. R., Cohen, B. A. & Grinspoon, D. H. What are the real constraints on the existence and magnitude of the late heavy bombardment? Icarus 189, 233–245 (2007).
Hayes, J. M. in Early life on Earth Vol. 84, 220–236 (Columbia University Press, New York, 1994).
Ueno, Y., Yamada, K., Yoshida, N., Maruyama, S. & Isozaki, Y. Evidence from fluid inclusions for microbial methanogenesis in the early Archaean era. Nature 440, 516–519 (2006).
Wolfe, J. & Fournier, G. P. Horizontal gene transfer constrains the timing of methanogen evolution. Nat. Ecol. Evol. 2, 897–903 (2018).
Evans, P. N. et al. Methane metabolism in the archaeal phylum Bathyarchaeota revealed by genome-centric metagenomics. Science 350, 434–438 (2015).
Vanwonterghem, I. et al. Methylotrophic methanogenesis discovered in the archaeal phylum Verstraetearchaeota. Nat. Microbiol. 1, 16170 (2016).
Williams, T. A. et al. Integrative modeling of gene and genome evolution roots the archaeal tree of life. Proc. Natl Acad. Sci. USA 114, 4602–4611 (2017).
Weiss, M. C. et al. The physiology and habitat of the last universal common ancestor. Nat. Microbiol. 1, 16116 (2016).
Sousa, F. L., Nelson-Sathi, S. & Martin, W. F. One step beyond a ribosome: the ancient anaerobic core. Biochim. Biophys. Acta 1857, 1027–1038 (2016).
Borrel, G., Adam, P. S. & Gribaldo, S. Methanogenesis and the Wood–Ljungdahl pathway: an ancient, versatile, and fragile association. Genome Biol. Evol. 8, 1706–1711 (2016).
Lyons, T. W., Reinhard, C. T. & Planavsky, N. J. The rise of oxygen in Earth’s early ocean and atmosphere. Nature 506, 307–315 (2014).
Schirrmeister, B. E., de Vos, J. M., Antonelli, A. & Bagheri, H. C. Evolution of multicellularity coincided with increased diversification of cyanobacteria and the Great Oxidation Event. Proc. Natl Acad. Sci. USA 110, 1791–1796 (2013).
Knoll, A. H. & Nowak, M. A. The timetable of evolution. Sci. Adv. 3, e1603076 (2017).
Shih, P. M., Hemp, J., Ward, L. M., Matzke, N. J. & Fischer, W. W. Crown group Oxyphotobacteria postdate the rise of oxygen. Geobiology 15, 19–29 (2017).
Kurland, C. G., Collins, L. J. & Penny, D. Genomics and the irreducible nature of eukaryote cells. Science 312, 1011–1014 (2006).
Chernikova, D., Motamedi, S., Csürös, M., Koonin, E. V. & Rogozin, I. B. A late origin of the extant eukaryotic diversity: divergence time estimates using rare genomic changes. Biol. Direct 6, 26 (2011).
Eme, L., Sharpe, S. C., Brown, M. W. & Roger, A. J. On the age of eukaryotes: evaluating evidence from fossils and molecular clocks. CSH Perspect. Biol. 6, a016139 (2014).
Parfrey, L. W., Lahr, D. J. G., Knoll, A. H. & Katz, L. A. Estimating the timing of early eukaryotic diversification with multigene molecular clocks. Proc. Natl Acad. Sci. USA 108, 13624–13629 (2011).
Shih, P. M. & Matzke, N. J. Primary endosymbiosis events date to the later Proterozoic with cross-calibrated phylogenetic dating of duplicated ATPase proteins. Proc. Natl Acad. Sci. USA 110, 12355–12360 (2013).
McInerney, J. O., O’Connell, M. J. & Pisani, D. The hybrid nature of the Eukaryota and a consilient view of life on Earth. Nat. Rev. Microbiol. 12, 449–455 (2014).
Roger, A. J., Muñoz-Gómez, S. A. & Kamikawa, R. The origin and diversification of mitochondria. Curr. Biol. 27, R1177–R1192 (2017).
Pittis, A. A. & Gabaldón, T. Late acquisition of mitochondria by a host with chimeric prokaryotic ancestry. Nature 531, 101–104 (2016).
Martin, W. F. et al. Late mitochondrial origin is an artifact. Genome Biol. Evol. 9, 373–379 (2017).
Pisani, D., Cotton, J. A. & McInerney, J. O. Supertrees disentangle the chimerical origin of eukaryotic genomes. Mol. Biol. Evol. 24, 1752–1760 (2007).
Ku, C. et al. Endosymbiotic origin and differential loss of eukaryotic genes. Nature 524, 427–432 (2015).
Lane, N. & Martin, W. The energetics of genome complexity. Nature 467, 929–934 (2010).
Benson, D. A. et al. GenBank. Nucleic Acids Res. 41, D36–D42 (2013).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Lartillot, N., Lepage, T. & Blanquart, S. PhyloBayes 3: a Bayesian software package for phylogenetic reconstruction and molecular dating. Bioinformatics 25, 2286–2288 (2009).
Kück, P. & Meusemann, K. FASconCAT: convenient handling of data matrices. Mol. Phylogenet. Evol. 56, 1115–1118 (2010).
Lartillot, N. & Philippe, H. A Bayesian mixture model for across-site heterogeneities in the amino-acid replacement process. Mol. Biol. Evol. 21, 1095–1109 (2004).
Aberer, A. J., Krompass, D. & Stamatakis, A. Pruning rogue taxa improves phylogenetic accuracy: an efficient algorithm and webservice. Syst. Biol. 62, 162–166 (2013).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
R Core Development Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2017).
Butterfield, N. J., Knoll, A. H. & Swett, K. A bangiophyte red alga from the Proterozoic of arctic Canada. Science 250, 104–108 (1990).
Rambaut, A., Drummond, A. J., Xie, D., Baele, G. & Suchard, M. A. Posterior summarisation in Bayesian phylogenetics using Tracer 1.7. Syst. Biol. https://doi.org/10.1093/sysbio/syy032 (2018).
Martijn, J., Vosseberg, J., Guy, L., Offre, P. & Ettema, T. J. G. Deep mitochondrial origin outside the sampled alphaproteobacteria. Nature 557, 101–105 (2018).
Esser, C. et al. A genome phylogeny for mitochondria among alpha-proteobacteria and a predominantly eubacterial ancestry of yeast nuclear genes. Mol. Biol. Evol. 21, 1643–1660 (2004).
Fitzpatrick, D. A., Creevey, C. J. & McInerney, J. O. Genome phylogenies indicate a meaningful α-proteobacterial phylogeny and support a grouping of the mitochondria with the Rickettsiales. Mol. Biol. Evol. 23, 74–85 (2006).
Acknowledgements
H.C.B. was supported by a NERC GW4 PhD studentship. J.W.C. was supported by a BBSRC SWBio PhD studentship. M.N.P. was supported by an 1851 Royal Commission Fellowship. P.C.J.D. was supported by BBSRC grant BB/N000919/1. T.A.W. is supported by a Royal Society Fellowship and NERC grant NE/P00251X/1.
Author information
Authors and Affiliations
Contributions
D.P., P.C.J.D. and T.A.W. designed the study. H.C.B. assembled the datasets and performed the phylogenetic and molecular clock analyses. M.N.P. and J.W.C. contributed further molecular clock analyses. H.C.B., D.P., P.C.J.D. and T.A.W. wrote the manuscript. All authors edited the manuscript and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary information, figures and tables
Rights and permissions
About this article
Cite this article
Betts, H.C., Puttick, M.N., Clark, J.W. et al. Integrated genomic and fossil evidence illuminates life’s early evolution and eukaryote origin. Nat Ecol Evol 2, 1556–1562 (2018). https://doi.org/10.1038/s41559-018-0644-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-018-0644-x
This article is cited by
-
Fossil-calibrated molecular clock data enable reconstruction of steps leading to differentiated multicellularity and anisogamy in the Volvocine algae
BMC Biology (2024)
-
Lost world of complex life and the late rise of the eukaryotic crown
Nature (2023)
-
Gene gain facilitated endosymbiotic evolution of Chlamydiae
Nature Microbiology (2023)
-
ATP synthase evolution on a cross-braced dated tree of life
Nature Communications (2023)
-
A biallelic variant of DCAF13 implicated in a neuromuscular disorder in humans
European Journal of Human Genetics (2023)