Abstract
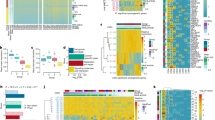
The evolutionary events that cause colorectal adenomas (benign) to progress to carcinomas (malignant) remain largely undetermined. Using multi-region genome and exome sequencing of 24 benign and malignant colorectal tumours, we investigate the evolutionary fitness landscape occupied by these neoplasms. Unlike carcinomas, advanced adenomas frequently harbour sub-clonal driver mutations—considered to be functionally important in the carcinogenic process—that have not swept to fixation, and have relatively high genetic heterogeneity. Carcinomas are distinguished from adenomas by widespread aneusomies that are usually clonal and often accrue in a ‘punctuated’ fashion. We conclude that adenomas evolve across an undulating fitness landscape, whereas carcinomas occupy a sharper fitness peak, probably owing to stabilizing selection.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Raw data are available via the European Genome-Phenome Archive (https://ega-archive.org/) accession code: EGAS00001003066.
References
Morson, B. C. Evolution of cancer of the colon and rectum. Cancer 34, 845–849 (1974).
Ashton-Rickardt, P. G. et al. High frequency of APC loss in sporadic colorectal carcinoma due to breaks clustered in 5q21–22. Oncogene 4, 1169–1174 (1989).
Powell, S. M. et al. APC mutations occur early during colorectal tumorigenesis. Nature 359, 235–237 (1992).
Fearon, E. R. & Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell 61, 759–767 (1990).
Jones, S. et al. Comparative lesion sequencing provides insights into tumor evolution. Proc. Natl Acad. Sci. USA 105, 4283–4288 (2008).
Smith, G. et al. Mutations in APC, Kirsten-ras, and p53—alternative genetic pathways to colorectal cancer. Proc. Natl Acad. Sci. USA 99, 9433–9438 (2002).
Muzny, D. M. et al. Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, 330–337 (2012).
Sottoriva, A. et al. A Big Bang model of human colorectal tumor growth. Nat. Genet. 47, 209–216 (2015).
Williams, M. J., Werner, B., Barnes, C. P., Graham, T. A. & Sottoriva, A. Identification of neutral tumor evolution across cancer types. Nat. Genet. 48, 238–244 (2016).
Wright, S. The roles of mutation, inbreeding, crossbreeding and selection in evolution. In Proc. 6th International Congress of Genetics Vol. 1 356–366 (1932).
Yap, T. A., Gerlinger, M., Futreal, P. A., Pusztai, L. & Swanton, C. Intratumor heterogeneity: seeing the wood for the trees. Sci. Transl. Med. 4, 127ps10 (2012).
Blum, M. G. B. & François, O. On statistical tests of phylogenetic tree imbalance: the Sackin and other indices revisited. Math. Biosci. 195, 141–153 (2005).
Fischer, A., Illingworth, C. J., Campbell, P. J. & Mustonen, V. EMu: probabilistic inference of mutational processes and their localization in the cancer genome. Genome Biol. 14, R39 (2013).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Katainen, R. et al. CTCF/cohesin-binding sites are frequently mutated in cancer. Nat. Genet. 47, 818–821 (2015).
Quirke, P. et al. DNA aneuploidy in colorectal adenomas. Br. J. Cancer 53, 477–481 (1986).
Jones, A. M. et al. Analysis of copy number changes suggests chromosomal instability in a minority of large colorectal adenomas. J. Pathol. 213, 249–256 (2007).
Wang, H., Liang, L., Fang, J.-Y. & Xu, J. Somatic gene copy number alterations in colorectal cancer: new quest for cancer drivers and biomarkers. Oncogene 35, 2011–2019 (2016).
Durinck, S. et al. Temporal dissection of tumorigenesis in primary cancers. Cancer Discov. 1, 137–143 (2011).
Newman, S. et al. The relative timing of mutations in a breast cancer genome. PLoS ONE 8, e64991 (2013).
Toyota, M. et al. CpG island methylator phenotype in colorectal cancer. Proc. Natl Acad. Sci. USA 96, 8681–8686 (1999).
Kim, T.-M. et al. Subclonal genomic architectures of primary and metastatic colorectal cancer based on intratumoral genetic heterogeneity. Clin. Cancer Res. 21, 4461–4472 (2015).
Uchi, R. et al. Integrated multiregional analysis proposing a new model of colorectal cancer evolution. PLoS Genet. 12, e1005778 (2016).
Suzuki, Y. et al. Multiregion ultra-deep sequencing reveals early intermixing and variable levels of intratumoral heterogeneity in colorectal cancer. Mol. Oncol. 11, 124–139 (2017).
Kim, T.-M. et al. Clonal origins and parallel evolution of regionally synchronous colorectal adenoma and carcinoma. Oncotarget 6, 27725–27735 (2015).
Stachler, M. D. et al. Paired exome analysis of Barrett’s esophagus and adenocarcinoma. Nat. Genet. 47, 1047–1055 (2015).
Maley, C. C. et al. Genetic clonal diversity predicts progression to esophageal adenocarcinoma. Nat. Genet. 38, 468–473 (2006).
Ross-Innes, C. S. et al. Whole-genome sequencing provides new insights into the clonal architecture of Barrett’s esophagus and esophageal adenocarcinoma. Nat. Genet. 47, 1038–1046 (2015).
Andrews, S. FastQC: a quality control tool for high throughput sequence data (Babraham Bioinformatics, 2013); http://www.bioinformatics.babraham.ac.uk/projects/fastqc/
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics. 25, 2078–2079 (2009).
Rimmer, A. et al. Integrating mapping-, assembly- and haplotype-based approaches for calling variants in clinical sequencing applications. Nat Genet. 46, 912–918 (2014).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6, 80–92 (2012).
Hasan, M. S., Wu, X. & Zhang, L.Performance evaluation of indel calling tools using real short-read data.Hum. Genom. 9, 20 (2015).
Narzisi, G. et al. Accurate de novo and transmitted indel detection in exome-capture data using microassembly. Nat. Methods 11, 1033–1036 (2014).
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform. 14, 178–192 (2013).
Fischer, A., Vázquez-García, I., Illingworth, C. J. R. & Mustonen, V. High-definition reconstruction of clonal composition in cancer. Cell Rep. 7, 1740–1752 (2014).
Werner, B., Traulsen, A., Sottoriva, A. & Dingli, D. Detecting truly clonal alterations from multi-region profiling of tumours. Sci. Rep. 7, 44991 (2017).
Nik-Zainal, S. et al. The life history of 21 breast cancers. Cell 149, 994–1007 (2012).
Paradis, E., Claude, J. & Strimmer, K.APE: analyses of phylogenetics and evolution in R language.Bioinformatics 20, 289–290 (2004).
Bortolussi, N., Durand, E., Blum, M. & Francois, O. apTreeshape: statistical analysis of phylogenetic tree shape. Bioinformatics 22, 363–364 (2005).
Acknowledgements
S.J.L., T.A.G. (A19771) and I.P.M.T. (A27327) are funded by Cancer Research UK. We acknowledge core funding provided to the Wellcome Trust Centre for Human Genetics from the Wellcome Trust (090532/Z/09/Z). T.A.G. and S.J.L. were also supported by the Bowel and Cancer Research small grant scheme. T.A.G. was also supported by the Wellcome Trust (202778/Z/16/Z). V.M. was supported in part by funding from the Wellcome Trust (098051). M. Kovac was supported by the Krebsliga beider Basel (grant no. KLBB-12-2013) and the University of Basel (‘Förderung exzellenter Nachwuchsforschender’). A-M.B. also acknowledges funding from Cancer Research UK (A14895). D.C.W. is supported by the Li Ka Shing Foundation. X.J. and I.P.M.T. are supported by an ERC advanced grant (EVOCAN-340560). The S:CORT study is funded by the MRC and Cancer Research UK. K.H is supported by Krebsliga Zentralschweiz. A.S. is supported by the Wellcome Trust (202778/B/16/Z), Cancer Research UK (A22909) and the Chris Rokos Fellowship in Evolution and Cancer. This work was also supported a Wellcome Trust award to the Centre for Evolution and Cancer (105104/Z/14/Z). J.E.E. was funded by the National Institute for Health Research (NIHR) Oxford Biomedical Research Centre (BRC). V.H.K. was funded by the Swiss National Science Foundation (P2SKP3_168322 / 1 and P2SKP3_168322 / 2). D.T. acknowledges funding from the EPSRC (grant no.: EP/F500351/1).
Author information
Authors and Affiliations
Consortia
Contributions
I.P.M.T., T.A.G. and S.J.L. conceived and designed the study. R.G., J.E.E., L.M.W., K.H., S.J.L. and I.P.M.T. provided the samples. H.D., A.-M.B., S.B. and L.C. performed the experiments. W.C., M. Kovac, V.M., P.M., R.A. and D.C.W. performed the bioinformatics analysis. W.C. and D.T. performed the mathematical analysis. C.G., A.R.A. and V.H.K. performed the image analysis. M.J., M.R.-J. and L.M.W. performed the pathology assessment. E.D., T.M. and the S:CORT consortium provided reference data. W.C., M. Kovac, V.M., D.T., R.A., V.H.K., X.J., D.C.W., Y.F., M.Kovacova, S.A., A.S., S.J.L., T.A.G. and I.P.M.T. analysed the data. W.C., A.S., S.J.L., T.A.G. and I.P.M.T. performed the evolutionary analysis. W.C., T.A.G. and I.P.M.T. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary figures 1–9; Supplementary modelling; Supplementary table legends
Supplementary tables
Supplementary tables 1–7
Rights and permissions
About this article
Cite this article
Cross, W., Kovac, M., Mustonen, V. et al. The evolutionary landscape of colorectal tumorigenesis. Nat Ecol Evol 2, 1661–1672 (2018). https://doi.org/10.1038/s41559-018-0642-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-018-0642-z
This article is cited by
-
Computational validation of clonal and subclonal copy number alterations from bulk tumor sequencing using CNAqc
Genome Biology (2024)
-
Aneuploidy and complex genomic rearrangements in cancer evolution
Nature Cancer (2024)
-
Evolving copy number gains promote tumor expansion and bolster mutational diversification
Nature Communications (2024)
-
Deterministic evolution and stringent selection during preneoplasia
Nature (2023)
-
Biallelic FBXW7 knockout induces AKAP8-mediated DNA damage in neighbouring wildtype cells
Cell Death Discovery (2023)