Abstract
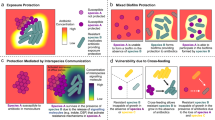
Microbes rarely exist in isolation, rather, they form intricate multi-species communities that colonize our bodies and inserted medical devices. However, the efficacy of antimicrobials is measured in clinical laboratories exclusively using microbial monocultures. Here, to determine how multi-species interactions mediate selection for resistance during antibiotic treatment, particularly following drug withdrawal, we study a laboratory community consisting of two microbial pathogens. Single-species dose responses are a poor predictor of community dynamics during treatment so, to better understand those dynamics, we introduce the concept of a dose-response mosaic, a multi-dimensional map that indicates how species’ abundance is affected by changes in abiotic conditions. We study the dose-response mosaic of a two-species community with a ‘Gene × Gene × Environment × Environment’ ecological interaction whereby Candida glabrata, which is resistant to the antifungal drug fluconazole, competes for survival with Candida albicans, which is susceptible to fluconazole. The mosaic comprises several zones that delineate abiotic conditions where each species dominates. Zones are separated by loci of bifurcations and tipping points that identify what environmental changes can trigger the loss of either species. Observations of the laboratory communities corroborated theory, showing that changes in both antibiotic concentration and nutrient availability can push populations beyond tipping points, thus creating irreversible shifts in community composition from drug-sensitive to drug-resistant species. This has an important consequence: resistant species can increase in frequency even if an antibiotic is withdrawn because, unwittingly, a tipping point was passed during treatment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
19 September 2018
In the version of this Article originally published, the following sentence was missing from the Acknowledgements: “R.E.B. is an EPSRC Healthcare Technologies Impact Fellow EP/N033671/1; I.G. is funded by ERC Consolidator grant 647292 MathModExp; A.J.P.B., N.A.R.G. and A.T. were funded by BBSRC grant BB/F00513X/1; K.H., I.G., S.N. and E.C. were funded by BBSRC grant BB/F005210/2.” This text has now been added.
References
Payne, D. J., Gwynn, M. N., Holmes, D. J. & Pompliano, D. L. Drugs for bad bugs: confronting the challenges of antibacterial discovery. Nat. Rev. Drug. Discov. 6, 29–40 (2007).
Mira, P. M. et al. Rational design of antibiotic treatment plans: a treatment strategy for managing evolution and reversing resistance. PLoS. ONE 10, 1–25 (2015).
Kollef, M. H. Is antibiotic cycling the answer to preventing the emergence of bacterial resistance in the intensive care unit? Clin. Infect. Dis. 43 (Suppl. 2), S82–S88 (2006).
Sundqvist, M. Reversibility of antibiotic resistance. Ups. J. Med. Sci. 119, 142–148 (2014).
Lee, J. et al. Control of extended-spectrum beta-lactamase-producing Escherichia coli and Klebsiella pneumoniae in a children’s hospital by changing antimicrobial agent usage policy. J. Antimicrob. Chemother. 60, 629–637 (2007).
Rahal, J. J. et al. Class restriction of cephalosporin use to control total cephalosporin resistance in nosocomial Klebsiella. JAMA 280, 1233–1237 (1998).
Cook, P. P., Catrou, P. G., Christie, J. D., Young, P. D. & Polk, R. E. Reduction in broad-spectrum antimicrobial use associated with no improvement in hospital antibiogram. J. Antimicrob. Chemother. 53, 853–859 (2004).
Nijssen, S. et al. Effects of reducing beta-lactam antibiotic pressure on intestinal colonization of antibiotic-resistant gram-negative bacteria. Intensive Care Med. 36, 512–519 (2010).
Chong, Y. et al. Antibiotic rotation for febrile neutropenic patients with hematological malignancies: clinical significance of antibiotic heterogeneity. PLoS ONE 8, e54190 (2013).
Takesue, Y. et al. Impact of a hospital-wide programme of heterogeneous antibiotic use on the development of antibiotic-resistant Gram-negative bacteria. J. Hosp. Infect. 75, 28–32 (2010).
Hashino, S. et al. Clinical impact of cycling the administration of antibiotics for febrile neutropenia in Japanese patients with hematological malignancy. Eur. J. Clin. Microbiol. Infect. Dis. 31, 173–178 (2012).
Sarraf-Yazdi, S. et al. A 9-year retrospective review of antibiotic cycling in a surgical intensive care unit. J. Surg. Res. 176, e73–e78 (2012).
Gruson, D. et al. Rotation and restricted use of antibiotics in a medical intensive care unit. Am. J. Respir. Crit. Care. Med. 162, 837–843 (2000).
Van Loon, H. J. et al. Antibiotic rotation and development of Gram-negative antibiotic resistance. Am. J. Respir. Crit. Care. Med. 171, 480–487 (2004).
Warren, D. et al. Cycling empirical antimicrobial agents to prevent emergence of antimicrobial-resistant Gram-negative bacteria among intensive care unit patients. Crit. Care Med. 32, 2450–2456 (2004).
Andersson, D. I. & Hughes, D. Antibiotic resistance and its cost: is it possible to reverse resistance? Nat. Rev. Microbiol. 8, 260–271 (2010).
Gonze, D., Lahti, L., Raes, J. & Faust, K. Multi-stability and the origin of microbial community types. ISME J. 11, 2159–2166 (2017).
Panda, S. et al. Short-term effect of antibiotics on human gut microbiota. PLoS ONE 9, e95476 (2014).
Jakobsson, H. E. et al. Short-term antibiotic treatment has differing long-term impacts on the human throat and gut microbiome. PLoS ONE 5, e9836 (2010).
Dethlefsen, L., McFall-Ngai, M. & Relman, Da An ecological and evolutionary perspective on human-microbe mutualism and disease. Nature 449, 811–818 (2007).
Antonopoulos, D. A. et al. Reproducible community dynamics of the gastrointestinal microbiota following antibiotic perturbation. Infect. Immun. 77, 2367–2375 (2009).
Perez-Cobas, A. E. et al. Gut microbiota disturbance during antibiotic therapy: a multi-omic approach. Gut 62, 1591–1601 (2013).
Dethlefsen, L. & Relman, D. A. Incomplete recovery and individualized responses of the human distal gut microbiota to repeated antibiotic perturbation. Proc. Natl Acad. Sci. USA 108 (Suppl.), 4554–4561 (2011).
McFarland, L. V., Elmer, G. W. & Surawicz, C. M. Breaking the cycle: treatment strategies for 163 cases of recurrent clostridium difficile disease. Am. J. Gastroenterol. 97, 1769–1775 (2002).
Cousin, L., Berre, M. L., Launay-Vacher, V., Izzedine, H. & Deray, G. Dosing guidelines for fluconazole in patients with renal failure. Nephrol. Dial. Transplant. 18, 2227–2231 (2003).
Ashbee, H. R. et al. Therapeutic drug monitoring (TDM) of antifungal agents: guidelines from the British Society for Medical Mycology. J. Antimicrob. Chemother. 69, 1162–1176 (2014).
Havey, T. C., Fowler, R. A., Pinto, R., Elligsen, M. & Daneman, N. Duration of antibiotic therapy for critically ill patients with bloodstream infections: a retrospective cohort study. Can. J. Infect. Dis. Med. Microbiol. 24, 129–137 (2013).
Cowart, S. L. & Stachura, M. E. Glucosuria (Butterworth Publishers, Boston, MA, 1990).
Carlotti, A. P. C. P. et al. A hyperglycaemic hyperosmolar state in a young child: diagnostic insights from a quantitative analysis. QJM 100, 125–137 (2007).
Manoj, G., George, M. R., Dipu, R. & Jishnu, J. The survival story of a diabetic ketoacidosis patient with blood sugar levels of 1985 mg/dl. Asian J. Med. Sci. 8, 60–61 (2017).
Ho, K.-m. & Cheng, T.-s. Common superficial fungal infections, a short review. Med. Bull. 15, 23–27 (2010).
Wenzel, R. P. & Gennings, C. Bloodstream infections due to Candida species in the intensive care unit: identifying especially high-risk patients to determine prevention strategies. Clin. Infect. Dis. 41 (Suppl. 6), S389–S393 (2005).
Brown, G. D. et al. Hidden killers: human fungal infections. Sci. Transl. Med. 4, 165rv13 (2012).
Kett, D. H., Azoulay, E., Echeverria, P. M. & Vincent, J.-L. Candida bloodstream infections in intensive care units: analysis of the extended prevalence of infection in intensive care unit study. Crit. Care Med. 39, 665–670 (2011).
Pappas, P. G. et al. Guidelines for treatment of candidiasis. Clin. Infect. Dis. 38, 161–189 (2004).
Rex, J. H. et al. Development of interpretive breakpoints for antifungal susceptibility testing: conceptual framework and analysis of in vitro–in vivo correlation data for fluconazole, itraconazole, and Candida infections. Subcommittee on Antifungal Susceptibility Testing of the National Committee for Clinical Laboratory Standards. Clin. Infect. Dis. 24, 235–247 (1997).
Lortholary, O. et al. Recent exposure to caspofungin or fluconazole influences the epidemiology of candidemia: a prospective multicenter study involving 2,441 patients. Antimicrob. Agents Chemother. 55, 532–538 (2011).
Hibbing, M. E., Fuqua, C., Parsek, M. R. & Peterson, S. B. Bacterial competition: surviving and thriving in the microbial jungle. Nat. Rev. Microbiol. 8, 15–25 (2010).
Scheffer, M., Carpenter, S., Foley, Ja, Folke, C. & Walker, B. Catastrophic shifts in ecosystems. Nature 413, 591–596 (2001).
Lenton, T. M. et al. Tipping elements in the Earth’s climate system. Proc. Natl Acad. Sci. USA 105, 1786–1793 (2008).
Wissel, C. A universal law of the characteristic return time near thresholds. Oecologia 65, 101–107 (1984).
Wiesenfeld, K. & Mcnamara, B. Small-signal amplification in bifurcating dynamical systems. Phys. Rev. A 33, 629–642 (1986).
Dai, L., Vorselen, D., Korolev, K. S. & Gore, J. Generic indicators for loss of resilience before a tipping point leading to population collapse. Science 336, 1175–1177 (2012).
Scheffer, M. et al. Early-warning signals for critical transitions. Nature 461, 53–59 (2009).
Dakos, V. et al. Slowing down as an early warning signal for abrupt climate change. Proc. Natl Acad. Sci. USA 105, 14308–14312 (2008).
Lenton, T. M. Early warning of climate tipping points. Nat. Clim. Change 1, 201–209 (2011).
Carpenter, S. R. & Brock, W. A. Rising variance: a leading indicator of ecological transition. Ecol. Lett. 9, 308–315 (2006).
Guttal, V. & Jayaprakash, C. Changing skewness: an early warning signal of regime shifts in ecosystems. Ecol. Lett. 11, 450–460 (2008).
Baillie, G. S. & Douglas, L. J. Effect of growth rate on resistance of Candida albicans biofilms to antifungal agents. Antimicrob. Agents Chemother. 42, 1900–1905 (1998).
Basson, N. J. Competition for glucose between Candida albicans and oral bacteria grown in mixed culture in a chemostat. J. Med. Microbiol. 49, 969–975 (2000).
Huang, M., McClellan, M., Berman, J. & Kao, K. C. Evolutionary dynamics of Candida albicans during in vitro evolution. Eukaryot. Cell 10, 1413–1421 (2011).
Ene, I. V., Brunke, S., Brown, A. J. P. & Hube, B. Metabolism in fungal pathogenesis. Cold Spring Harb. Perspect. Med. 4, a019695 (2014).
MacLean, R. C. & Gudelj, I. Resource competition and social conflict in experimental populations of yeast. Nature 441, 498–501 (2006).
Fidel, P. L. Jr, Vazquez, J. A. & Sobel, J. D. Candida glabrata: review of epidemiology, pathogenesis, and clinical disease with comparison to C. albicans. Clin. Microbiol. Rev. 12, 80–96 (1999).
Ray, D. et al. Prevalence of Candida glabrata and its response to boric acid vaginal suppositories in comparison with oral fluconazole in patients with diabetes and vulvovaginal candidiasis. Diabetes Care 30, 312–317 (2007).
Sonnenburg, E. D. et al. Specificity of polysaccharide use in intestinal bacteroides species determines diet-induced microbiota alterations. Cell 141, 1241–1252 (2010).
David, L. A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505, 559–563 (2014).
Metzler-Zebeli, B. U., Lange, J. C., Zijlstra, R. T. & Gänzle, M. G. Dietary non-starch polysaccharides alter the abundance of pathogenic clostridia in pigs. Livest. Sci. 152, 31–35 (2013).
Allison, K. R., Brynildsen, M. P. & Collins, J. J. Metabolite-enabled eradication of bacterial persisters by aminoglycosides. Nature 473, 216–220 (2011).
Peng, B. et al. Exogenous alanine and/or glucose plus kanamycin kills antibiotic-resistant bacteria. Cell. Metab. 21, 249–261 (2015).
Zampieri, M. et al. Metabolic constraints on the evolution of antibiotic resistance. Mol. Syst. Biol. 13, 917 (2017).
Milne, S. W., Cheetham, J., Lloyd, D., Aves, S. & Bates, S. Cassettes for PCR-mediated gene tagging in Candida albicans utilizing nourseothricin resistance. Yeast 3, 833–841 (2011).
Mansfield, B. E. et al. Azole drugs are imported by facilitated diffusion in Candida albicans and other pathogenic fungi. PLoS Pathog. 6, 11 (2010).
Botev, Z. I., Grotowski, J. F. & Kroese, D. P. Kernel density estimation via diffusion. Ann. Stat. 38, 2916–2957 (2010).
Acknowledgements
In memory of our friend and colleague Ken Haynes who sadly passed away on 19 March 2018. R.E.B. is an EPSRC Healthcare Technologies Impact Fellow EP/N033671/1; I.G. is funded by ERC Consolidator grant 647292 MathModExp; A.J.P.B., N.A.R.G. and A.T. were funded by BBSRC grant BB/F00513X/1; K.H., I.G., S.N. and E.C. were funded by BBSRC grant BB/F005210/2.
Author information
Authors and Affiliations
Contributions
I.G. and R.E.B. conceived the idea. R.E.B., I.G. and E.C. designed all experiments (apart from Supplementary Fig. 5). T.C.W. designed the experiment in Slementary Fig. 5. E.C., S.N., A.R.S., A.T., B.D.E., K.H., N.A.R.G. and A.J.P.B. carried out experiments. I.G. and R.E.B. developed and numerically simulated the mathematical model. R.E.B., I.G., E.C., T.C.W., K.H., N.A.R.G. and A.J.P.B. discussed the results. R.E.B., E.C. and I.G. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary figures 1–12; supplementary experimental details; supplementary modelling
Rights and permissions
About this article
Cite this article
Beardmore, R.E., Cook, E., Nilsson, S. et al. Drug-mediated metabolic tipping between antibiotic resistant states in a mixed-species community. Nat Ecol Evol 2, 1312–1320 (2018). https://doi.org/10.1038/s41559-018-0582-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-018-0582-7
This article is cited by
-
Seeking patterns of antibiotic resistance in ATLAS, an open, raw MIC database with patient metadata
Nature Communications (2022)
-
Repeatable ecological dynamics govern the response of experimental communities to antibiotic pulse perturbation
Nature Ecology & Evolution (2020)
-
Glyoxylate cycle gene ICL1 is essential for the metabolic flexibility and virulence of Candida glabrata
Scientific Reports (2019)
-
Handling unpredictable ecosystems
Nature Ecology & Evolution (2018)