Abstract
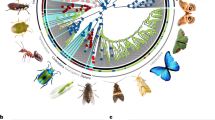
Plant secondary metabolites play important ecological and evolutionary roles, most notably in the deterrence of natural enemies. The classical theory explaining the evolution of plant chemical diversity is that new defences arise through a pairwise co-evolutionary arms race between plants and their specialized natural enemies. However, plant species are bombarded by dozens of different herbivore taxa from disparate phylogenetic lineages that span a wide range of feeding strategies and have distinctive physiological constraints that interact differently with particular plant metabolites. How do plant defence chemicals evolve under such multiple and potentially contrasting selective pressures imposed by diverse herbivore communities? To tackle this question, we exhaustively characterized the chemical diversity and insect herbivore fauna from 31 sympatric species of Amazonian Protieae (Burseraceae) trees. Using a combination of phylogenetic, metabolomic and statistical learning tools, we show that secondary metabolites that were associated with repelling herbivores (1) were more frequent across the Protieae phylogeny and (2) were found in average higher abundance than other compounds. Our findings suggest that generalist herbivores can play an important role in shaping plant chemical diversity and support the hypothesis that chemical diversity can also arise from the cumulative outcome of multiple diffuse interactions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lokvam, J. & Kursar, T. A. Divergence in structure and activity of phenolic defenses in young leaves of two co-occurring Inga species. J. Chem. Ecol. 31, 2563–2580 (2005).
Ehrlich, P. & Raven, P. H. Butterflies and plants: a study in coevolution. Evolution 18, 586–608 (1964).
Speed, M. P., Fenton, A., Jones, M. G., Ruxton, G. D. & Brockhurst, M. A. Coevolution can explain defensive secondary metabolite diversity in plants. New Phytol. 208, 1251–1263 (2015).
Berenbaum, M. R. Chemical mediation of coevolution: phylogenetic evidence for Apiaceae and associates. Ann. Mo. Bot. Gard. 88, 45–59 (2001).
Edger, P. P. et al. The butterfly plant arms-race escalated by gene and genome duplications. Proc. Natl Acad. Sci. USA 112, 8362–8366 (2015).
Alejo-Armijo, A. et al. Antimicrobial and antibiofilm activities of procyanidins extracted from laurel wood against a selection of foodborne microorganisms. Int. J. Food Sci. Technol. 52, 679–686 (2017).
Wise, M. J. & Rausher, M. D. Evolution of resistance to a multiple-herbivore community: genetic correlations, diffuse coevolution, and constraints on the plant’s response to selection. Evolution 67, 1767–1779 (2013).
Volf, M. et al. Community structure of insect herbivores is driven by conservatism, escalation and divergence of defensive traits in Ficus. Ecol. Lett. 21, 83–92 (2018).
Lankau, R. A. Specialist and generalist herbivores exert opposing selection on a chemical defense. New Phytol. 175, 176–184 (2007).
Volf, M., Hrcek, J., Julkunen-Tiitto, R. & Novotny, V. To each its own: differential response of specialist and generalist herbivores to plant defence in willows. J. Anim. Ecol. 84, 1123–1132 (2015).
Janzen, D. H. When is it coevolution? Evolution 34, 611–612 (1980).
Iwao, K. & Rausher, M. D. Evolution of plant resistance to multiple herbivores: quantifying diffuse coevolution. Am. Nat. 149, 316–335 (1997).
Futuyma, D. J. & Agrawal, A. A. Macroevolution and the biological diversity of plants and herbivores. Proc. Natl Acad. Sci. USA 106, 18054–18061 (2009).
Opitz, S. E. W. & Müller, C. Plant chemistry and insect sequestration. Chemoecology 19, 117–154 (2009).
Reinecke, A. & Hilker, M. in Annual Plant Reviews: Insect–Plant Interactions Vol. 47 (eds Voelckel, C. & Jander, G.) 115–153 (John Wiley & Sons, Chichester, 2014).
Johnson, M. T. J., Ives, A. R., Ahern, J. & Salminen, J. P. Macroevolution of plant defenses against herbivores in the evening primroses. New Phytol. 203, 267–279 (2014).
Kursar, T. A. et al. The evolution of antiherbivore defenses and their contribution to species coexistence in the tropical tree genus Inga. Proc. Natl Acad. Sci. USA 106, 18073–18078 (2009).
Richards, L. A. et al. Phytochemical diversity drives plant–insect community diversity. Proc. Natl Acad. Sci. USA 112, 10973–10978 (2015).
Basset, Y. Diversity and abundance of insect herbivores foraging on seedlings in a rainforest in Guyana.Ecol. Entomol. 24, 245–259 (1999).
Dyer, L. A. et al. Host specificity of Lepidoptera in tropical and temperate forests. Nature 448, 696–699 (2007).
Novotny, V. et al. Low beta diversity of herbivorous insects in tropical forests. Nature 448, 692–695 (2007).
Jones, C. G. & Lawton, J. H. Plant chemistry and insect species richness of British umbellifers. J. Anim. Ecol. 60, 767–777 (1991).
Tibshirani, R. Regression shrinkage and selection via the lasso.J. R. Stat. Soc. B Stat. Methodol. 58, 267–288 (1996).
Rasmann, S., Bennett, A., Biere, A., Karley, A. & Guerrieri, E.Root symbionts: powerful drivers of plant above- and belowground indirect defenses. Insect Sci. 24, 947–960 (2017).
Langenheim, J. H. Plant Resins: Chemistry, Evolution, Ecology, and Ethnobotany (Timber Press, Portland, 2003).
Andrew, R. L., Peakall, R., Wallis, I. R. & Foley, W. J. Spatial distribution of defense chemicals and markers and the maintenance of chemical variation. Ecology 88, 716–728 (2007).
Firn, R. D. & Jones, C. G. in Phytochemical Diversity and Redundancy in Ecological Interactions (eds Romeo, J. T., Saunders, J. A. & Barbosa, P.) 295–312 (Springer, New York, 1996).
Coley, P. D., Endara, M.-J. & Kursar, T. A. Consequences of interspecific variation in defenses and herbivore host choice for the ecology and evolution of Inga, a speciose rainforest tree. Oecologia https://doi.org/10.1007/s00442-018-4080-z (2018).
Fine, P. V. A., Daly, D. C., Villa, F. G., Mesones, I. & Cameron, K. M. The contribution of edaphic heterogeneity to the evolution and diversity of Burseraceae trees in the western Amazon. Evolution 59, 1464–1478 (2005).
Folmer, O., Black, M., Hoeh, W., Lutz, R. & Vrijenhoek, R.DNA primers for amplification of mitochondrial cytochrome c oxidase subunit 1 from diverse metazoan invertebrates.Mol. Mar. Biol. Biotechnol. 3, 294–299 (1994).
Hebert, P. D. N., Penton, E. H., Burns, J. M., Janzen, D. H. & Hallwachs, W. Ten species in one: DNA barcoding reveals cryptic species in the neotropical skipper butterfly Astraptes fulgerator. Proc. Natl Acad. Sci. USA 101, 14812–14817 (2004).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Schliep, K. P. phangorn: phylogenetic analysis in R. Bioinformatics 27, 592–593 (2011).
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003).
Miller, M. A., Pfeiffer, W. & Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In 2010 Gateway Computing Environments Workshop (GCE) 1–8 (IEEE, 2010).
Miller, M. A., Pfeiffer, W. & Schwartz, T. Extreme digital discovery. In Proc. 2011 TeraGrid Conference 1–8 (ACM, 2011).
Salazar, D., Jaramillo, A. & Marquis, R. J. The impact of plant chemical diversity on plant–herbivore interactions at the community level. Oecologia 181, 1199–1208 (2016).
Salazar, D., Jaramillo, A. & Marquis, R. J. Chemical similarity and local community assembly in the species rich tropical genus Piper. Ecology 97, 3176–3183 (2016).
Lokvam, J. & Fine, P. V. A. An oxidized squalene derivative from Protium subserratum Engl. (Engl.) growing in Peru. Molecules 17, 7451–7457 (2012).
Lokvam, J., Metz, M. R., Takeoka, G. R., Nguyen, L. & Fine, P. V. A. Habitat-specific divergence of procyanidins in Protium subserratum (Burseraceae). Chemoecology 25, 293–302 (2015).
Davies, T. The new automated mass spectrometry deconvolution and identification system (AMDIS). Spectrosc. Eur. 10, 24–27 (1998).
The NIST Mass Spectral Search Program for the NIST/EPA/NIH Mass Spectral Library (National Institute of Standards and Technology, Gaithersburg, 2017).
Horai, H. et al. MassBank: a public repository for sharing mass spectral data for life sciences. J. Mass Spectrom. 45, 703–714 (2010).
Stein, S. E. An integrated method for spectrum extraction and compound identification from gas chromatography/mass spectrometry data. J. Am. Soc. Mass Spectrom. 10, 770–781 (1999).
Suzuki, R. & Shimodaira, H. Pvclust: an R package for assessing the uncertainty in hierarchical clustering. Bioinformatics 22, 1540–1542 (2006).
R Development Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, 2015).
Pluskal, T., Castillo, S., Villar-Briones, A. & Orešič, M. MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinformatics 11, 395 (2010).
Kembel, S. W. et al. Picante: R tools for integrating phylogenies and ecology. Bioinformatics 26, 1463–1464 (2010).
Blomberg, S. P., Garland, T. & Ives, A. R. Testing for phylogenetic signal in comparative data: behavioral traits are more labile. Evolution 57, 717–745 (2003).
Dixon, P. VEGAN, a package of R functions for community ecology. J. Veg. Sci. 14, 927–930 (2003).
Dray, S. & Dufour, A.-B. The ade4 package: implementing the duality diagram for ecologists. J. Stat. Softw. 22, 1–20 (2007).
Friedman, J., Hastie, T. & Tibshirani, R.Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Meiners, T. Chemical ecology and evolution of plant–insect interactions: a multitrophic perspective.Curr. Opin. Insect Sci. 8, 22–28 2015).
Wäschke, N., Meiners, T. & Rostás, M. in Chemical Ecology of Insect Parasitoids (eds Wajnberg, E. & Colazza, S.) 37–63 (John Wiley & Sons, Chichester, 2013).
Erb, M. & Robert, C. A. M. Sequestration of plant secondary metabolites by insect herbivores: molecular mechanisms and ecological consequences. Curr. Opin. Insect Sci. 14, 8–11 (2016).
Heckel, D. G. in Annual Plant Reviews: Insect–Plant Interactions Vol. 47 (eds Voelckel, C. & Jander, G.) 77–114 (John Wiley & Sons, Chichester, 2014).
Pentzold, S., Zagrobelny, M., Roelsgaard, P. S., Møller, B. L. & Bak, S. The multiple strategies of an insect herbivore to overcome plant cyanogenic glucoside defence. PLoS ONE 9, e91337 (2014).
Jones, C. G., Firn, R. D. & Malcolm, S. B. On the evolution of plant secondary chemical diversity [and discussion]. Phil. Trans. R. Soc. Lond. B 333, 273–280 (1991).
Acknowledgements
We thank R. Marquis, M. Metz, N. Whiteman and C. Marshall for comments on the manuscript; G. Takeoka (USDA, Albany), P. Oboyski (Essig Museum), L. Smith (Evolutionary Genetics Laboratory at the University of California, Berkeley) and N. Tsutsui for advice, laboratory space and sample archiving; E. Hendrickson, B. Ho, C. Chong, S. Visvanathan, E. Suh and S. Sharma for help with the laboratory work; and D. Vásquez and C. Villacorta for field assistance. We also thank C. Rivera for help with research permits at the Allpahuayo-Mishana National Reserve. Funding for this project was provided by the NSF DEB (award number 1254214).
Author information
Authors and Affiliations
Contributions
D.S., J.L., P.dV., P.V.A.F. and I.M. contributed to the design, analysis and preparation of the manuscript. I.M., M.V.P. and J.M.A.Z. coordinated the fieldwork and data collection. D.S. and P.V.A.F. wrote the article with contributions from all co-authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods, Supplementary Figures 1–11, Supplementary Table 2, Supplementary References
Supplementary Table 1
Herbivore samples supplementary information. Includes the Essig Museum of Entomology, University of California, Berkeley, Essig Museum and Genbank accession numbers
Rights and permissions
About this article
Cite this article
Salazar, D., Lokvam, J., Mesones, I. et al. Origin and maintenance of chemical diversity in a species-rich tropical tree lineage. Nat Ecol Evol 2, 983–990 (2018). https://doi.org/10.1038/s41559-018-0552-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-018-0552-0
This article is cited by
-
Efficacy and selectivity of Sextonia rubra wood extracts and formulation in the control of Aedes aegypti strains
Journal of Pest Science (2024)
-
Metabolomic Evenness Underlies Intraspecific Differences Among Lineages of a Wetland Grass
Journal of Chemical Ecology (2023)
-
A review of Neotropical Burseraceae
Brazilian Journal of Botany (2022)
-
Phytochemistry reflects different evolutionary history in traditional classes versus specialized structural motifs
Scientific Reports (2021)
-
Diversification of terpenoid emissions proposes a geographic structure based on climate and pathogen composition in Japanese cedar
Scientific Reports (2021)