Abstract
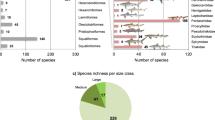
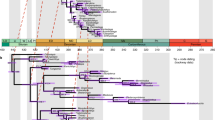
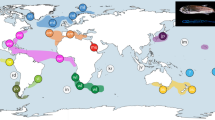
The Cretaceous–Palaeogene (K–Pg) mass extinction is linked to the rapid emergence of ecologically divergent higher taxa (for example, families and orders) across terrestrial vertebrates, but its impact on the diversification of marine vertebrates is less clear. Spiny-rayed fishes (Acanthomorpha) provide an ideal system for exploring the effects of the K–Pg on fish diversification, yet despite decades of morphological and molecular phylogenetic efforts, resolution of both early diverging lineages and enormously diverse subclades remains problematic. Recent multilocus studies have provided the first resolved phylogenetic backbone for acanthomorphs and suggested novel relationships among major lineages. However, these new relationships and associated timescales have not been interrogated using phylogenomic approaches. Here, we use targeted enrichment of >1,000 ultraconserved elements in conjunction with a divergence time analysis to resolve relationships among 120 major acanthomorph lineages and provide a new timescale for acanthomorph radiation. Our results include a well-supported topology that strongly resolves relationships along the acanthomorph backbone and the recovery of several new relationships within six major percomorph subclades. Divergence time analyses also reveal that crown ages for five of these subclades, and for the bulk of the species diversity in the sixth, coincide with the K–Pg boundary, with divergences between anatomically and ecologically distinctive suprafamilial clades concentrated in the first 10 million years of the Cenozoic.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Erwin, D. H. Lessons from the past: biotic recoveries from mass extinctions. Proc. Natl Acad. Sci. USA 98, 5399–5403 (2001).
Krug, A. Z., Jablonski, D. & Valentine, J. W. Signature of the end-Cretaceous mass extinction in the modern biota. Science 323, 767–771 (2009).
Jarvis, E. D. et al. Whole-genome analyses resolve early branches in the tree of life of modern birds. Science 346, 1320–1331 (2014).
Prum, R. O. et al. A comprehensive phylogeny of birds (Aves) using targeted next-generation DNA sequencing. Nature 526, 569–573 (2015).
O’Leary, M. A. et al. The placental mammal ancestor and the post-K–Pg radiation of placentals. Science 339, 662–667 (2013).
Longrich, N. R., Bhullar, B.-A. S. & Gauthier, J. A. Mass extinction of lizards and snakes at the Cretaceous–Paleogene boundary. Proc. Natl Acad. Sci. USA 109, 21396–21401 (2012).
Benton, M. J. Diversification and extinction in the history of life. Science 268, 52–58 (1995).
Near, T. J. et al. Phylogeny and tempo of diversification in the superradiation of spiny-rayed fishes. Proc. Natl Acad. Sci. USA 110, 12738–12743 (2013).
Sanciangco, M. D., Carpenter, K. E. & Betancur-R, R. Phylogenetic placement of enigmatic percomorph families (Teleostei: Percomorphaceae). Mol. Phylogenet. Evol. 94, 565–576 (2016).
Santini, F., Harmon, L. J., Carnevale, G. & Alfaro, M. E. Did genome duplication drive the origin of teleosts? A comparative study of diversification in ray-finned fishes. BMC Evol. Biol. 9, 194 (2009).
Near, T. J. et al. Resolution of ray-finned fish phylogeny and timing of diversification. Proc. Natl Acad. Sci. USA 109, 13698–13703 (2012).
Patterson, C. & Smith, A. B. Periodicity in extinction: the role of systematics. Ecology 70, 802–811 (1989).
MacLeod, N. et al. The Cretaceous–Tertiary biotic transition. J. Geol. Soc. Lond. 154, 265–292 (1997).
Cavin, L. in Geological and Biological Effects of Impact Events (eds Buffetaut, E. & Koeberl, C.) 141–158 (Springer, Berlin & Heidelberg, 2002).
Friedman, M. Explosive morphological diversification of spiny-finned teleost fishes in the aftermath of the end-Cretaceous extinction. Proc. R. Soc. B 277, 1675–1683 (2010).
Friedman, M. Ecomorphological selectivity among marine teleost fishes during the end-Cretaceous extinction. Proc. Natl Acad. Sci. USA 106, 5218–5223 (2009).
Sibert, E. C. & Norris, R. D. New age of fishes initiated by the Cretaceous–Paleogene mass extinction. Proc. Natl Acad. Sci. USA 112, 8537–8542 (2015).
Patterson, C. An overview of the early fossil record of the acanthomorphs. Bull. Mar. Sci. 52, 29–59 (1993).
Johnson, D. G. Percomorph phylogeny: progress and problems. Bull. Mar. Sci. 52, 3–28 (1993).
Johnson, D. G. & Patterson, C. Percomorph phylogeny: a survey of acanthomorphs and a new proposal. Bull. Mar. Sci. 52, 554–626 (1993).
Davesne, D. et al. The phylogenetic intrarelationships of spiny-rayed fishes (Acanthomorpha, Teleostei, Actinopterygii): fossil taxa increase the congruence of morphology with molecular data. Front. Ecol. Evol. 4, 129 (2016).
Alfaro, M. E. et al. Nine exceptional radiations plus high turnover explain species diversity in jawed vertebrates. Proc. Natl Acad. Sci. USA 106, 13410–13414 (2009).
Nelson, J. S. Fishes of the World 4th edn (Wiley, Hoboken, 2006).
Betancur, R. et al. The tree of life and a new classification of bony fishes. PLoS Curr. 5, https://doi.org/10.1371/currents.tol.53ba26640df0ccaee75bb165c8c26288 (2013).
Wainwright, P. C. et al. The evolution of pharyngognathy: a phylogenetic and functional appraisal of the pharyngeal jaw key innovation in labroid fishes and beyond. Syst. Biol. 61, 1001–1027 (2012).
Faircloth, B. C., Sorenson, L., Santini, F. & Alfaro, M. E. A phylogenomic perspective on the radiation of ray-finned fishes based upon targeted sequencing of ultraconserved elements (UCEs). PLoS ONE 8, e65923 (2013).
Bejerano, G. et al. Ultraconserved elements in the human genome. Science 304, 1321–1325 (2004).
Faircloth, B. C. et al. Ultraconserved elements anchor thousands of genetic markers spanning multiple evolutionary timescales. Syst. Biol. 61, 717–726 (2012).
Setiamarga, D. H. E. et al. Divergence time of the two regional medaka populations in Japan as a new time scale for comparative genomics of vertebrates. Biol. Lett. 5, 812–816 (2009).
Yamanoue, Y., Miya, M., Inoue, J. G., Matsuura, K. & Nishida, M. The mitochondrial genome of spotted green pufferfish Tetraodon nigroviridis (Teleostei: Tetraodontiformes) and divergence time estimation among model organisms in fishes. Genes Genet. Syst. 81, 29–39 (2006).
Yang, Z. & Rannala, B. Bayesian estimation of species divergence times under a molecular clock using multiple fossil calibrations with soft bounds. Mol. Biol. Evol. 23, 212–226 (2006).
Zheng, Y., Peng, R., Kuro-o, M. & Zeng, X. Exploring patterns and extent of bias in estimating divergence time from mitochondrial DNA sequence data in a particular lineage: a case study of salamanders (order Caudata). Mol. Biol. Evol. 28, 2521–2535 (2011).
Brown, J. W. & Van Tuinen, M. (eds) in Living Dinosaurs: The Evolutionary History of Modern Birds 306–324 (Wiley, Chichester, 2011).
Smith, B. T. & Klicka, J. Examining the role of effective population size on mitochondrial and multilocus divergence time discordance in a songbird. PLoS ONE 8, e55161 (2013).
Friedman, M. & Sallan, L. C. Five hundred million years of extinction and recovery: a Phanerozoic survey of large-scale diversity patterns in fishes. Palaeontology 55, 707–742 (2012).
Miya, M. et al. Evolutionary origin of the scombridae (tunas and mackerels): members of a Paleogene adaptive radiation with 14 other pelagic fish families. PLoS ONE 8, e73535 (2013).
Harrington, R. C. et al. Phylogenomic analysis of carangimorph fishes reveals flatfish asymmetry arose in a blink of the evolutionary eye. BMC Evol. Biol. 16, 224 (2016).
Carnevale, G. & Johnson, G. D. A Cretaceous cusk-eel (Teleostei, Ophidiiformes) from Italy and the Mesozoic diversification of percomorph fishes. Copeia 103, 771–791 (2015).
Schwarzhans, W. in Mesozoic Fishes—Systematics and Paleoecology (ed. Viohl, G.) 417–431 (F. Pfeil, München, 1996).
Faircloth, B. C. & Glenn, T. C. Not all sequence tags are created equal: designing and validating sequence identification tags robust to indels. PLoS ONE 7, e42543 (2012).
Glenn, T. C. et al. Adapterama I: universal stubs and primers for thousands of dual-indexed Illumina libraries (iTru & iNext). Preprint at https://doi.org/10.1101/049114 (2016).
Blumenstiel, B. et al. Targeted exon sequencing by in-solution hybrid selection. Curr. Protoc. Hum. Genet. 66, 18.4.1–18.4.24 (2010).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Faircloth, B. C. PHYLUCE is a software package for the analysis of conserved genomic loci. Bioinformatics 32, 786–788 (2015).
Lanfear, R., Calcott, B., Ho, S. Y. W. & Guindon, S. Partitionfinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol. Biol. Evol. 29, 1695–1701 (2012).
Lanfear, R., Calcott, B., Kainer, D., Mayer, C. & Stamatakis, A. Selecting optimal partitioning schemes for phylogenomic datasets. BMC Evol. Biol. 14, 82 (2014).
Aberer, A. J., Kobert, K. & Stamatakis, A. ExaBayes: massively parallel Bayesian tree inference for the whole-genome era. Mol. Biol. Evol. 31, 2553–2556 (2014).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Phillips, M. J. & Penny, D. The root of the mammalian tree inferred from whole mitochondrial genomes. Mol. Phylogenet. Evol. 28, 171–185 (2003).
Phillips, M. J., Delsuc, F. & Penny, D. Genome-scale phylogeny and the detection of systematic biases. Mol. Biol. Evol. 21, 1455–1458 (2004).
Seo, T.-K. Calculating bootstrap probabilities of phylogeny using multilocus sequence data. Mol. Biol. Evol. 25, 960–971 (2008).
Mirarab, S. & Warnow, T. ASTRAL-II: coalescent-based species tree estimation with many hundreds of taxa and thousands of genes. Bioinformatics 31, i44–i52 (2015).
Sukumaran, J. & Holder, M. T. DendroPy: a Python library for phylogenetic computing. Bioinformatics 26, 1569–1571 (2010).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Dos Reis, M. & Yang, Z. Approximate likelihood calculation on a phylogeny for Bayesian estimation of divergence times. Mol. Biol. Evol. 28, 2161–2172 (2011).
Smith, S., Brown, J. W. & Walker, J. F. So many genes, so little time: comments on divergence-time estimation in the genomic era. Preprint at https://doi.org/10.1101/114975 (2017).
Huerta-Cepas, J., Serra, F. & Bork, P. ETE 3: reconstruction, analysis, and visualization of phylogenomic data. Mol. Biol. Evol. 33, 1635–1638 (2016).
Vaidya, G., Lohman, D. J. & Meier, R. SequenceMatrix: concatenation software for the fast assembly of multi-gene datasets with character set and codon information. Cladistics 27, 171–180 (2011).
Lanfear, R., Frandsen, P. B., Wright, A. M., Senfeld, T. & Calcott, B. PartitionFinder 2: new methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol. Biol. Evol. 34, 772–773 (2017).
Chen, W.-J. et al. New insights on early evolution of spiny-rayed fishes (Teleostei: Acanthomorpha). Front. Mar. Sci. 1, 1–17 (2014).
Acknowledgements
We thank J. Johnson for the illustrations used in the figures. This work was partially supported by National Science Foundation grants DEB-0842397 (to M.E.A.); DEB-1242260, DEB-1655624 (to B.C.F.); and startup funds from Louisiana State University (to B.C.F.). DNA alignment was supported in part by resources and technical expertise from the Georgia Advanced Computing Resource Center—a partnership between the University of Georgia’s Office of the Vice President for Research and Office of the Vice President for Information Technology. Phylogenetic analysis portions of this research were conducted using high-performance computing resources provided by Louisiana State University (http://www.hpc.lsu.edu).
Author information
Authors and Affiliations
Contributions
M.E.A. conceived the study. M.E.A., B.C.F., R.C.H., M.F. and T.J.N. designed the study. L.S., R.C.H. and B.C.F. collected the sequence data from enriched UCE loci. M.F. selected appropriate calibration points. B.C.F., M.E.A., D.Č. and C.H.O. performed the data analyses. M.E.A. and B.C.F. wrote the manuscript with help from R.C.H., L.S., M.F., D.Č. and T.J.N. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Information, Methods, Results, References and Figures
Supplementary Table 1
Species, family, institutional or personal voucher code, preparation method (Prep.), enrichment method (Enrich.), sequencing platform (Plat.), and NCBI BioSample and SRA accession information for each individual used in this study
Supplementary Table 2
Statistics for trimmed sequence reads collected from each library enriched for ultraconserved element loci
Supplementary Table 3
Statistics for all contigs assembled from trimmed sequence data collected from each library enriched for ultraconserved element loci
Supplementary Table 4
Statistics for ultraconserved element (UCE) contigs assembled from trimmed sequence data collected from each library enriched for ultraconserved element loci
Rights and permissions
About this article
Cite this article
Alfaro, M.E., Faircloth, B.C., Harrington, R.C. et al. Explosive diversification of marine fishes at the Cretaceous–Palaeogene boundary. Nat Ecol Evol 2, 688–696 (2018). https://doi.org/10.1038/s41559-018-0494-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-018-0494-6
This article is cited by
-
Convergent genomic signatures associated with vertebrate viviparity
BMC Biology (2024)
-
Ecological niche conservatism spurs diversification in response to climate change
Nature Ecology & Evolution (2024)
-
Range-wide population genomics of common seadragons shows secondary contact over a former barrier and insights on illegal capture
BMC Biology (2023)
-
Evolution of the germline mutation rate across vertebrates
Nature (2023)
-
Genomics of cold adaptations in the Antarctic notothenioid fish radiation
Nature Communications (2023)