Abstract
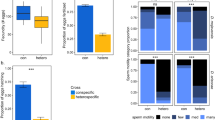
Males and females have different fitness optima but share the vast majority of their genomes, causing an inherent genetic conflict between the two sexes that must be resolved to achieve maximal population fitness. We show that two tandem duplicate genes found specifically in Drosophila melanogaster are sexually antagonistic, but rapidly evolved sex-specific functions and expression patterns that mitigate their antagonistic effects. We use copy-specific knockouts and rescue experiments to show that Apollo (Apl) is essential for male fertility but detrimental to female fertility, in addition to its important role in development, while Artemis (Arts) is essential for female fertility but detrimental to male fertility. Further analyses show that Apl and Arts have essential roles in spermatogenesis and oogenesis. These duplicates formed ~200,000 years ago, underwent a strong selective sweep and lost most expression in the antagonized sex. These data provide direct evidence that gene duplication allowed rapid mitigation of sexual conflict by allowing Apl and Arts to evolve essential sex-specific reproductive functions and complementary expression in male and female gonads.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Lande, R. Sexual dimorphism, sexual selection, and adaptation in polygenic characters. Evolution 34, 292–305 (1980).
Partridge, L. & Hurst, L. D. Sex and conflict. Science 281, 2003–2008 (1998).
Bonduriansky, R. & Chenoweth, S. F. Intralocus sexual conflict. Trends Ecol. Evol. 24, 280–288 (2009).
Parsch, J. & Ellegren, H. The evolutionary causes and consequences of sex-biased gene expression. Nat. Rev. Genet. 14, 83–87 (2013).
Gallach, M. & Betrán, E. Intralocus sexual conflict resolved through gene duplication. Trends Ecol. Evol. 26, 222–228 (2011).
Connallon, T. & Clark, A. G. The resolution of sexual antagonism by gene duplication. Genetics 187, 919–937 (2011).
Wyman, M. J., Cutter, A. D. & Rowe, L. Gene duplication in the evolution of sexual dimorphism. Evolution 66, 1556–1566 (2012).
Ohno, S. Evolution by Gene Duplication (Springer, Berlin, 1970).
Innan, H. & Kondrashov, F. The evolution of gene duplications: classifying and distinguishing between models. Nat. Rev. Genet. 11, 97–108 (2010).
Long, M., VanKuren, N. W., Chen, S. & Vibranovski, M. D. New gene evolution: little did we know. Annu. Rev. Genet. 47, 307–333 (2013).
Chippindale, A. K., Gibson, J. R. & Rice, W. R. Negative genetic correlation for adult fitness between sexes reveals ontogenetic conflict in Drosophila. Proc. Natl Acad. Sci. USA 98, 1671–1675 (2001).
Morrow, E. H., Stewart, A. D. & Rice, W. R. Assessing the extent of genome-wide intralocus sexual conflict via experimentally enforced gender-limited selection. J. Evol. Biol. 21, 1046–1054 (2008).
Innocenti, P. & Morrow, E. H. The sexually antagonistic genes of Drosophila melanogaster. PLoS Biol. 8, e1000335 (2010).
Obbard, D. J. et al. Estimating divergence dates and substitution rates in the Drosophila phylogeny. Mol. Biol. Evol. 29, 3459–3473 (2012).
Keightley, P. D. et al Analysis of the genome sequences of three Drosophila melanogaster spontaneous mutation accumulation lines. Genome Res. 19, 1195–1201 (2009).
Rogers, R. L. & Hartl, D. L. Chimeric genes as a source of rapid evolution in Drosophila melanogaster. Mol. Biol. Evol. 29, 517–29 (2012).
Hudson, R. R., Kreitman, M. & Aguadé, M. A test of neutral molecular evolution based on nucleotide data. Genetics 116, 153–159 (1987).
Ford, M. J. & Aquadro, C. F. Selection on X-linked genes during speciation in the Drosophila athabasca complex. Genetics 144, 689–703 (1996).
Timinszky, G. et al. The importin-β P446L dominant-negative mutant protein loses RanGTP binding ability and blocks the formation of intact nuclear envelope. J. Cell Sci. 115, 1675–1687 (2002).
Gorlich, D., Seewald, M. J. & Ribbeck, K. Characterization of Ran-driven cargo transport and the RanGTPase system by kinetic measurements and computer simulation. EMBO J. 22, 1088–1100 (2003).
Harel, A. & Forbes, D. J. Importin-β: conducting a much larger cellular symphony. Mol. Cell 16, 319–330 (2004).
Gates, J. Drosophila egg chamber elongation: insights into how tissues and organs are shaped. Fly 6, 213–227 (2012).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Rice, W. R. Sex chromosomes and the evolution of sexual dimorphism. Evolution 38, 735–742 (1984).
Boughman, J. W. in The Princeton Guide to Evolution (eds Losos, J. B. et al.) 520–528 (Princeton University Press, Princeton, 2014).
Pavlicev, M. & Wagner, G. P. A model of developmental evolution: selection, pleiotropy and compensation. Trends Ecol. Evol. 27, 316–322 (2012).
Zera, A. J. & Harshman, L. G. The physiology of life history tradeoffs in animals. Annu. Rev. Ecol. Syst. 32, 95–126 (2001).
Harshman, L. G. & Hoffmann, A. A. Laboratory selection experiments in Drosophila: what do they really tell us? Trends Ecol. Evol. 15, 32–36 (2000).
Ranz, J. M., Castillo-Davis, C. I., Meiklejohn, C. D. & Hartl, D. L. Sex-dependent gene expression and evolution of the Drosophila transcriptome. Science 300, 1742–1745 (2003).
Ellegren, H. & Parsch, J. The evolution of sex-biased genes and sex-biased gene expression. Nat. Rev. Genet. 8, 689–698 (2007).
Zhou, Q. et al. On the origin of new genes in Drosophila. Genome Res. 18, 1446–1455 (2008).
Zhang, Y. E., Vibranovski, M. D., Krinsky, B. H. & Long, M. Age-dependent chromosomal distribution of male-biased genes in Drosophila. Genome Res. 20, 1526–1533 (2010).
Chen, S. et al. Frequent recent origination of brain genes shaped the evolution of foraging behavior in Drosophila. Cell Rep. 1, 118–132 (2012).
Rogers, R. L. et al. Landscape of standing variation for tandem duplications in Drosophila yakuba and Drosophila simulans. Mol. Biol. Evol. 31, 1750–1766 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Pool, J. et al. Population genomics of sub-saharan Drosophila melanogaster: African diversity and non-African admixture. PLoS Genet. 8, e1003080 (2012).
Dietzl, G. et al. A genome-wide transgenic RNAi library for conditional gene inactivation in Drosophila. Nature 448, 151–156 (2007).
Green, E. W., Fedele, G., Giorgini, F. & Kyriacou, C. P. A Drosophila RNAi collection is subject to dominant phenotypic effects. Nat. Methods 11, 222–223 (2014).
Bassett, A. & Liu, J. L. CRISPR/Cas9 mediated genome engineering in Drosophila. Methods 69, 128–136 (2014).
flyCRISPR Optimal Target Finder (University of Wisconsin, accessed 1 January 2016); http://tools.flycrispr.molbio.wisc.edu/targetFinder/
Port, F., Chen, H.-M., Lee, T. & Bullock, S. L. Optimized CRISPR/Cas tools for efficient germline and somatic genome engineering in Drosophila. Proc. Natl Acad. Sci. USA 111, E2967–E2976 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–360 (2012).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Trapnell, C. et al. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nat. Biotechnol. 31, 46–53 (2013).
Kibanov, M. V., Kotov, A. A. & Olenina, L. V. Multicolor fluorescence imaging of whole-mount Drosophila testes for studying spermatogenesis. Anal. Biochem. 436, 55–64 (2013).
Rathke, C. et al. Distinct functions of Mst77F and protamines in nuclear shaping and chromatin condensation during Drosophila spermiogenesis. Eur. J. Cell Biol. 89, 326–338 (2010).
McKenna, A. et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Van der Auwera, G. A. et al. in Current Protocols in Bioinformatics Vol. 43, 11.10.1–11.10.33 (John Wiley & Sons, 2013).
Langley, C. H. et al. Genomic variation in natural populations of Drosophila melanogaster. Genetics 192, 533–598 (2012).
Blanchette, M. et al. Aligning multiple genomic sequences with the threaded blockset aligner. Genome Res. 14, 708–715 (2004).
Harris, R. S. Improved Pairwise Alignment of Genomic DNA. PhD thesis, Pennsylvania State University (2007).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Abascal, F., Zardoya, R. & Telford, M. J. TranslatorX: multiple alignment of nucleotide sequences guided by amino acid translations. Nucleic Acids Res. 38, 7–13 (2010).
Welch, B. L. The generalization of ‘Student’s’ problem when several different population variances are involved. Biometrika 34, 28–35 (1947).
Shapiro, S. S. & Wilk, M. B. An analysis of variance test for normality (complete samples). Biometrika 52, 591–611 (1965).
R Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, 2004).
van der Linde, K., Houle, D., Spicer, D. S. & Steppan, S. J. A supermatrix-based molecular phylogeny of the family Drosophilidae. Genet. Res. 92, 25–38 (2010).
Acknowledgements
We thank R. Renkawitz-Pohl for providing antibodies; S. Horne-Badovinac, C. Stevenson, T. Davis and I. Rebay for help with staining, microscopy and discussion; G.Y.-C. Lee for advice on population genomics analyses; and members of the Long lab, M. Kreitman, R. Hudson, E. Ferguson and L. Harshman for valuable discussion. N.W.V. was supported by the National Institutes of Health (NIH) Genetics and Regulation Training Grant T32GM007197 and a National Science Foundation (NSF) Graduate Research Fellowship. M.L. was supported by NSF1026200, NIH R01GM100768-01A1 and NIH R01GM116113.
Author information
Authors and Affiliations
Contributions
N.W.V. and M.L. designed the study. N.W.V. collected and analysed data with M.L. N.W.V. and M.L. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–6, Supplementary Tables 1–9.
Rights and permissions
About this article
Cite this article
VanKuren, N.W., Long, M. Gene duplicates resolving sexual conflict rapidly evolved essential gametogenesis functions. Nat Ecol Evol 2, 705–712 (2018). https://doi.org/10.1038/s41559-018-0471-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-018-0471-0
This article is cited by
-
Mitochondrially mediated RNA interference, a retrograde signaling system affecting nuclear gene expression
Heredity (2024)
-
Evolution and function of developmentally dynamic pseudogenes in mammals
Genome Biology (2022)
-
Ancient homomorphy of molluscan sex chromosomes sustained by reversible sex-biased genes and sex determiner translocation
Nature Ecology & Evolution (2022)
-
A Simple Evolutionary Model of Genetic Robustness After Gene Duplication
Journal of Molecular Evolution (2022)
-
Nuclear transport genes recurrently duplicate by means of RNA intermediates in Drosophila but not in other insects
BMC Genomics (2021)