Abstract
Many bacterial species are composed of multiple lineages distinguished by extensive variation in gene content. These often cocirculate in the same habitat, but the evolutionary and ecological processes that shape these complex populations are poorly understood. Addressing these questions is particularly important for Streptococcus pneumoniae, a nasopharyngeal commensal and respiratory pathogen, because the changes in population structure associated with the recent introduction of partial-coverage vaccines have substantially reduced pneumococcal disease. Here we show that pneumococcal lineages from multiple populations each have a distinct combination of intermediate-frequency genes. Functional analysis suggested that these loci may be subject to negative frequency-dependent selection (NFDS) through interactions with other bacteria, hosts or mobile elements. Correspondingly, these genes had similar frequencies in four populations with dissimilar lineage compositions. These frequencies were maintained following substantial alterations in lineage prevalences once vaccination programmes began. Fitting a multilocus NFDS model of post-vaccine population dynamics to three genomic datasets using Approximate Bayesian Computation generated reproducible estimates of the influence of NFDS on pneumococcal evolution, the strength of which varied between loci. Simulations replicated the stable frequency of lineages unperturbed by vaccination, patterns of serotype switching and clonal replacement. This framework highlights how bacterial ecology affects the impact of clinical interventions.
Similar content being viewed by others
Studies of population genomics have revealed that many bacterial species exhibit extensive variation in their ‘accessory’ genomes. While neutral evolutionary models can account for such diversity1,2,3, allowing for heterogeneity in the evolutionary rate between genes significantly improves their fit to genomic data4,5,6, consistent with selection causing differences in gene content7. If recombination rates are sufficiently high, selection can alter the distribution of individual genes8. However, lower levels of recombination are associated with chromosome-wide sweeps, such that niche specialization at one or more loci can result in largely clonal ‘ecotypes’9,10. Similarly, a recent model has suggested that selection acting on a high proportion of the genome could partition even freely recombining bacteria into highly diverged ‘metabolic types’11. As well as adaptation to particular niches, this latter model11 considered antigenic loci to be under NFDS, the situation in which alleles are most beneficial to genotypes when they are rare. This is based on the assumption that antigens become more costly when common, because they are more frequently recognized by acquired immune responses.
Such NFDS has been proposed to explain the extensive antigenic diversity of the nasopharyngeal colonizer and respiratory pathogen Streptococcus pneumoniae (pneumococcus)11,12,13. This variation makes anti-pneumococcal vaccine development challenging. The first licensed conjugate vaccine (PCV7) targeted seven of over ninety serotypes14, and consequently was associated with ‘serotype replacement’ as vaccine types were replaced by non-vaccine types (NVTs), with no substantial overall change in carriage rates15. This was driven by both serotype switching, the replacement of vaccine types by NVTs that differed at few loci other than those which determined the serotype, and clonal replacement of vaccine types by distantly related NVTs. These population dynamics are now amenable to detailed study, having been tracked by genomic surveillance of isolates carried by children in both Massachusetts (USA)14 and Southampton (UK)16,17, and isolates from invasive pneumococcal disease in adults in Nijmegen (the Netherlands)18. Here we use the distribution of the accessory genome across isolates to develop a gene frequency-based model of bacterial population structure based on multiple NFDS mechanisms19,20.
Results
Enrichment of loci under frequency-dependent selection in the accessory genome
Previous analyses of 5,442 clusters of orthologous genes (COGs) in the Massachusetts pneumococcal population suggested that those present at intermediate frequencies were important in distinguishing sequence clusters21. To identify functions that were enriched in this set of genes, the 1,112 COGs present in 5% to 95% of isolates and 1,194 core COGs14 were annotated by integrating multiple analyses (Fig. 1a and Supplementary Datasets 1, 2). The most substantial difference was in mobile genetic elements (MGEs; Fisher’s exact test; odds ratio (OR) = 336; two-sided P < 2.2 × 10−16). However, few of these genes were ‘cargo’ beneficial to the host bacterium, and were instead likely to be parasitic, consistent with the distribution of prophages between pneumococci21,22. Correspondingly, restriction modification systems that protect against MGE infection accounted for 2.4% of the intermediate-frequency genes, but were absent from the core COGs. These are most often advantageous when rare, such that the donor of an infecting MGE is unlikely to have the same system19, but typically futile when ubiquitous. Therefore, the coexistence of lineages probably involves competition between bacteria and MGEs through ‘kill-the-winner’ dynamics, a form of NFDS in which an increase in a genotype’s frequency would be associated with a counterbalancing rise in the prevalence of MGE genotypes able to infect such cells23.
a, Functional classification of the 1,112 intermediate-frequency and 1,194 core COGs in the Massachusetts pneumococcal population, as detailed in Supplementary Datasets 1, 2. Each barchart compares the functional category frequencies in the intermediate-frequency and core COGs. Categories are grouped as likely to be under NFDS resulting from bacterium–MGE interactions (pink), bacterium–bacterium interactions (blue), or bacterium-host interactions (green). The chart with orange segments shows the frequencies of loci with roles in general metabolism or signal transduction, or that otherwise could not be classified. b, Population structure of the 4,127 isolates from Massachusetts (Mass), Southampton (Soton), Nijmegen and Maela (Supplementary Dataset 3). The maximum likelihood phylogeny was generated from 1,447 core gCOGs. The adjacent columns contain a row for each genome, and represent the population in which the bacterium was isolated, its susceptibility to PCV7-induced immunity, and sequence cluster classification. c, Comparison of core genome divergence, quantified as the cophenetic distance between isolates in the core genome phylogeny, and the accessory genome divergence, quantified as the Jaccard distance between the gCOG content of isolates. Each point represents a pairwise comparison between randomly sampled isolates (excluding the polyphyletic SC0), which was coloured orange if the isolates belonged to the same sequence cluster; purple if they belonged to different sequence clusters but were both encapsulated; and dark blue otherwise, revealing the presence of some genetically divergent unencapsulated genotypes. Isocontour lines quantify the distribution of points in each category.
Functional annotation of the intermediate-frequency genes also suggested that direct interference competition between bacteria was likely to be important in maintaining a diversity of lineages24. Bacteriocins, which mediate interstrain competition25, were significantly enriched in the accessory genome relative to the core (Fisher’s exact test; OR = 24.0; two-sided P < 2.2 × 10−16). Although regulatory components of the bacteriocin-like peptide (blp) locus were conserved across the population, most of the gene cluster was composed of various combinations of bacteriocin and immunity protein genes, many of which were found in multiple loci26. Despite this diversity, each of the previously described fifteen monophyletic sequence clusters14 was typically associated with one distinctive blp allele (Supplementary Fig. 1), with the exception of sequence cluster (SC)3 and SC14, which did not coexist for long owing to vaccine-induced population dynamics14.
Sequence clusters also varied in their complement of rarer bacteriocin biosynthesis gene clusters, including pneumocyclicin27 and pneumolancidin28; two loci associated with the TprA/PhrA quorum-sensing system29, and other putative operons (Supplementary Fig. 1). No individual gene cluster replicated the diversity of the blp locus; instead, sequence variation often corresponded to disruptive mutations in bacteriocin structural or biosynthetic genes. Assuming phenotypes can be reliably inferred from the gene clusters, such mutations result in bacteria immune to the relevant bacteriocin, but unable to kill competitors. These immune non-producers cocirculate with producer cells carrying the putatively fully functional allele, and susceptible cells completely lacking the gene cluster. Analogous variation with respect to individual bacteriocins is probably present between the blp loci, given their diverse complements of production and immunity genes. If both biosynthesis and immunity functions are costly, these phenotypes can coexist through rock–paper–scissors NFDS dynamics, as producers kill susceptible cells, immune non-producers outcompete producers, and susceptible cells outcompete immune non-producers30. Therefore, the distinctive overall bacteriocin-production profile of strains may be shaped by NFDS acting on multiple loci.
NFDS can also result from competition for resources20,31. A particular nutrient import strategy, either optimized for different nutrients31 or different concentrations of the same nutrient32, will become less advantageous as it becomes more common, as a consequence of more intense competition for the same resource24. While nutrient importers account for 11.5% of the core COGs, because many are universally necessary, they also make up 9.35% of the intermediate-frequency COGs. Therefore, these genes are significantly enriched relative to general metabolic genes in the latter category (Fisher’s exact test, OR = 2.48, two-sided P = 2.61 × 10−8). This suggests that NFDS may sustain multiple nutrient-acquisition strategies in the population as a consequence of interstrain competition for resources.
Antibiotic resistance, also variable between isolates, could be affected by similar competition33. If resistant bacteria are considered adapted to hosts consuming antibiotics, but suffering a cost in untreated hosts, then resistance will be most effective as a resource acquisition strategy where rare, because of the lessened competition with other strains. This could directly result in NFDS, although there are alternative explanations for the coexistence of sensitive and resistant pneumococci that instead imply NFDS through other mechanisms34.
A further functional category that was enriched in the intermediate-frequency COGs relative to the core genome were genes encoding the biosynthesis of immunogenic structures, such as surface proteins35 or the capsule36 (Fisher’s exact test, OR = 2.56, two-sided P = 9.23 × 10−10). These can be under NFDS as long as alleles are immunologically distinguishable, a criterion met by the serotype-defining capsule36, as well as accessory antigens that are typically either present as large surface structures, or completely absent, such as the pili37.
Therefore, multiple disparate functions enriched in the intermediate-frequency genes relative to the core genome can each be understood as being subject to NFDS, albeit through different processes. While no NFDS mechanism could be identified for 32.1% of the intermediate-frequency COGs, this category is likely to include both metabolic enzymes and signal transduction proteins linked to loci under NFDS on genomic islands, and loci under NFDS that cannot be identified as such, owing to incomplete functional information. To test whether these inferences applied to other pneumococcal populations in a similar manner, further genomic datasets were compared to those from Massachusetts.
Population similarities in frequencies of genes, but not genotypes
Overall, 4,127 isolates were combined from available reference sequences, Massachusetts, Southampton, Nijmegen and the Maela refugee camp in Thailand, where the population is unvaccinated38 (Supplementary Dataset 3). This analysis identified 11,049 ‘global’ COGs (gCOGs), from which a ‘relaxed’ core of 1,447 gCOGs was extracted to generate a maximum-likelihood phylogeny (Fig. 1b and Supplementary Fig. 2). Notably, there was little evidence of genetic isolation-by-distance, as both vaccine-type status and country of isolation had a polyphyletic distribution, indicating a history of recombination and frequent international migration.
The core alignment was also used to define 74 sequence clusters. Plotting the pairwise core genome divergence of isolates, represented by their cophenetic separation in the tree, against their accessory genome divergence, calculated as the Jaccard distance between the gCOG content of isolates, showed that members of the same sequence cluster were substantially more similar in their accessory, as well as core, genomes (Fig. 1c). These differences between lineages were probably biologically meaningful, as they represented a significant proportion of the accessory genome and were preserved despite international dissemination of some genotypes and ongoing horizontal DNA transfer. Although some of the previously identified atypical unencapsulated lineages were associated with extensive private gene content21, sequence clusters of encapsulated pneumococci each contained few unique accessory loci. The mean numbers of gCOGs present in ≥95% of the isolates in a given sequence cluster, but not meeting this criterion in any other sequence cluster in the same population, were only 16.75 in Massachusetts, 19.94 in Southampton, 19.46 in Nijmegen and 15.02 Maela (Supplementary Fig. 2). The distinctiveness of the sequence clusters instead resulted from the polyclonal distribution of the 1,731 intermediate-frequency gCOGs, present in between 5% and 95% of the pre-vaccination isolates in at least one population (Supplementary Fig. 2). Therefore, a long history of recombination was reflected in intermediate-frequency loci that were associated with multiple lineages, with each lineage in turn defined by a unique combination of intermediate-frequency loci.
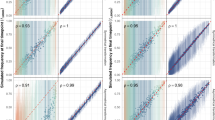
Despite the lineages representing discrete and distinct sets of genotypes, their prevalences were highly heterogeneous between the four populations, with a significant correlation only between those in Massachusetts and Southampton (Fig. 2a). In contrast, the frequencies of accessory gCOGs were strongly correlated between Massachusetts and every other population (Fig. 2b; Pearson's correlation, two-sided P < 10−15 in all comparisons). This suggests that pneumococcal populations are configured by genomic islands being maintained at equilibrium frequencies that are conserved between populations, consistent with their prevalence being influenced by NFDS19. A significant deviation between populations was the elevated frequency of Tn916 in Maela; this transposon underlies tetracycline resistance21, and hence the difference is likely to represent a location-specific selection pressure rather than drift39. Therefore, selection appears to shape pneumococcal populations to be similar in frequencies of genes, rather than genotypes.
a, The distribution of sequence clusters between populations; the frequency of each sequence cluster in Massachusetts is shown on the x axis, and the corresponding frequencies in Maela, Southampton and Nijmegen are shown on the y axes (from top to bottom). Red points represent sequence clusters predominantly (≥75%) composed of vaccine-type isolates across the compared populations; blue points represent sequence clusters predominantly (≥75%) composed of NVT isolates, and black points are mixed sequence clusters. b, The distribution of gCOGs between populations. The frequency of each intermediate-frequency gCOG in Massachusetts is shown on the x axis, and the corresponding frequencies in Maela, Southampton and Nijmegen are shown on the y axes. Only gCOGs present at a mean frequency between 5% and 95% across the two compared populations were included, and the corresponding points are coloured according to the functional annotation of COGs in Fig. 1a. The gCOGs encoded by Tn916, including the tetM tetracycline-resistance gene, are annotated in the comparison of the Massachusetts and Maela populations. c, The pre- and post-vaccination frequencies of sequence clusters in Massachusetts, Southampton and Nijmegen. Points are coloured as in a, showing the general decrease in the frequency of sequence clusters of vaccine types. d, The pre- and post-vaccination frequency of gCOGs in Massachusetts, Southampton and Nijmegen. Only gCOGs with an overall frequency between 5% and 95% in the relevant population were included. Points are coloured as in b. The reduced frequency of the wciN allele involved in the synthesis of vaccine-type capsules 6A and 6B is annotated. Each panel displays Pearson's correlation statistics, including two-sided P values.
Vaccination as a test of NFDS
The partial-coverage vaccines introduced to limit pneumococcal disease can be used as a natural experiment, to test whether loci expected to change in frequency due to association with vaccine types were actually maintained at equilibrium frequencies by NFDS. Although a significant correlation existed between pre- and post-PCV7 sequence cluster frequencies in the three vaccinated populations (Fig. 2c), divergence in population composition was driven by the replacement of some vaccine-type sequence clusters with distantly related NVT lineages. Across all comparisons of pre- and post-PCV7 populations, gCOG frequencies showed a stronger positive correlation. This stability in gene frequencies reflected the significant correlation between the post-PCV7 decrease in a gCOG’s absolute frequency in vaccine-type isolates, and the contemporaneous increase in its absolute frequency in NVT isolates (Supplementary Fig. 3), consistent with the NFDS hypothesis. The greatest deviation in the Massachusetts population was wciN, which is directly involved in the synthesis of the vaccine-targeted 6A and 6B capsules, reflecting differences in selection pressures between timepoints14. This suggested that the equilibrium frequencies of the intermediate-frequency gCOGs were likely to govern the post-vaccine restructuring of the population.
To quantify whether NFDS of intermediate-frequency gCOGs could explain changes in pneumococcal populations better than a neutral model, a discrete-time Wright–Fisher multilocus NFDS model was constructed in which the number of offspring produced by a genotype i at generation t, X i,t , was distributed as:
General density-dependent competition was parameterized by the number of pneumococci in the simulated population at generation t, N t, and the environment’s carrying capacity κ, which was constant across t, reflecting the stable levels of pneumococcal carriage post-PCV7 (refs 15,16). The other demographic process was migration, at rate m (per month-long generation), by which isolates in the resident simulated population were replaced by genotypes randomly selected from the genomic data from the same location. Vaccine-type genotypes were subject to a fitness cost, v, representing vaccine efficacy at preventing transmission. The final term parameterized NFDS, the strength of which was determined by σ f and the exponent π i,t :
where l is an intermediate-frequency locus (gCOG or antibiotic-resistance phenotype), and g i,l is a binary variable indicating whether l is present in genotype i. Each l has an equilibrium frequency e l , its prevalence in the pre-vaccination sample, and an instantaneous frequency at generation t, f l,t . Therefore f l,t determines whether l benefits its host, when it is rare relative to e l , or has a net cost, when it is common relative to e l . Model details are described in Supplementary Fig. 4 and the Methods.
The σ f , v and m parameters were estimated for the Massachusetts population using Approximate Bayesian Computation, an inference technique for intractable simulator-based models40,41. The simulated population was compared to the sequence cluster distribution across three time points (Fig. 3a) using the Jensen–Shannon divergence (JSD) to determine similarity. Convergence of the parameter estimates found strong evidence for NFDS (σ f significantly greater than its lower bound; Table 1, Supplementary Table 1 and Supplementary Fig. 5). The precedent of other models4,6 has suggested that the fit could be improved by allowing the strength of selection to be heterogeneous across loci. Therefore, an expanded model featured a proportion, p f , of the intermediate-frequency loci experiencing NFDS at strength σ f , whereas (1 − p f ) experienced NFDS at strength σ w (see Methods). Convergence of parameter estimates found strong evidence for NFDS (σ f and p f significantly greater than their lower bounds; Table 1, Supplementary Table 1 and Supplementary Fig. 5), with a substantial improvement over the homogeneous selection model, as quantified by the significantly smaller JSD values from appropriately parameterized simulations (Wilcoxon rank-sum test on 100 simulation pairs, W = 9,902, two-sided P = 4.73 × 10−33; Supplementary Fig. 6).
In each barplot, vertical black lines split the bacterial population into sequence clusters, which are annotated at the top of the graph. Each sequence cluster is split into three timepoints: pre-vaccination, a midpoint sample and a late sample. Only sequence clusters present at a frequency greater than 2.5% in at least one of these timepoints in the genomic sample are included in the graphs; full results are shown in the Supplementary Information. The bars at each timepoint are split into red segments, for vaccine-type isolates, and blue segments, for NVT isolates. In each comparison, the top row is the genomic sample against which simulations were evaluated. The bottom row summarizes the output of 100 simulations using the heterogeneous-rate multilocus NFDS model with the point-estimate parameter values from Table 1. At the times at which samples were reported in the respective genomic collections, the same numbers of isolates were randomly selected from the simulated populations. The bars represent the median result and the error bars (orange for vaccine-type isolates and purple for NVT isolates) represent the interquartile range observed across the simulations. a, The results for Massachusetts. Isolates were split into pre-vaccination (2001; 133 isolates), midpoint (2004; 203 isolates) and late (2007; 280 isolates) samples. b, The results for Southampton. Isolates were split into pre-vaccination (up to 2007; 100 isolates), midpoint (2008–2009; 194 isolates) and late (2010–2011; 195 isolates) samples. c, The results for Nijmegen. The isolates were split into pre-vaccination (up to 2007; 209 isolates), midpoint (2008–2009; 80 isolates) and late (2010–2011; 48 isolates) samples.
At the locus level, those genes subject to stronger NFDS stabilized close to their equilibrium frequencies, whereas the frequencies of those subject to weaker NFDS showed near-neutral drift (Supplementary Fig. 6). At the lineage level, these simulations replicated three important facets of the post-vaccination population dynamics (Fig. 3a and Supplementary Fig. 7). The first was the stable post-vaccine prevalence of some NVT sequence clusters, such as SC4 and SC8. The second was serotype switching, the replacement of vaccine types by NVTs within sequence clusters that remained at stable overall frequencies, as observed in SC1, SC5, SC9 and SC15. The third was clonal replacement of vaccine types by unrelated NVTs, such as the contemporaneous disappearance of SC13, SC14, SC22 and SC24, and the expansion of SC3, SC6, SC7 and SC11. These trends were not trivial to replicate. The same framework was used to fit a neutral model (NFDS eliminated, with σ f = 0); a serotype-focused single locus NFDS model (e l applied to serotype, rather than locus, frequencies), and an ecotype model (e l applied to sequence cluster, rather than locus, frequencies). Both the neutral and serotype models poorly reproduced the stable frequency of SC8, serotype switching within SC9 and SC15, or any patterns of clonal expansion. The ecotype models better reproduced NVT sequence-cluster stability and serotype switching, but did not replicate the observed patterns of clonal replacement. All of these models resulted in significantly worse fits to the data than the heterogeneous multilocus NFDS model (Supplementary Figs. 6, 7).
The estimated vaccine selection strength, v, of 0.081 per month from the heterogenous-rate multilocus NFDS is consistent with PCV7’s halving of the rate at which vaccine types are acquired42, if pneumococci transmit at least once every six months, an interval similar to the carriage duration of vaccine serotypes43. Similarly, the estimated migration rate, m, of 0.0044 per month suggests half the resident Massachusetts pneumococcal population would be replaced by immigrant strains over approximately 13 years, which is realistic given the 50% probability that a pneumococcal lineage was detectable in different localities within Massachusetts after 3–4 years14.
Consistent evidence of NFDS in other populations
The homogeneous and heterogeneous multilocus NFDS models were also fitted to similar surveillance data from Southampton (Supplementary Figs. 5, 8). The JSD values for the heterogenous rate model were reproducible and significantly smaller than for the homogeneous rate model (Wilcoxon rank-sum test on 100 simulation pairs, W = 9,954, two-sided P = 1.01 × 10−33). The point estimates of parameter values were robust and, in the case of the three parameters determining the strength of NFDS, very similar to those for Massachusetts (Table 1 and Supplementary Table 1).
However, the vaccine selection strength was estimated to be 2.54-fold higher in Southampton than in Massachusetts. This difference is probably attributable to the substantially higher PCV7 coverage in children under 24 months of age in the years immediately after the introduction of the vaccine in the UK relative to the USA44,45, combined with the lower age range included in the Southampton study, excluding older children who are less likely to have been immunized, or in whom natural acquisition of immunity blunted the selective pressure of the vaccine16,46. Simulations using these point estimates replicated the strain dynamics observed in the genomic sample (Fig. 3b). Predominantly vaccine-type SC5 and SC18 were eliminated at realistic rates; NVT lineages SC3, SC19 and SC35 remained at stable frequencies; serotype switching occurred within SC1 and SC9, while NVT SC2 rose in prevalence at a much faster rate than same lineage did in Massachusetts.
The homogeneous and heterogeneous rate multilocus NFDS models were also fitted to a genomic dataset from cases of invasive pneumococcal disease in Nijmegen. The heterogeneous model was a significantly closer fit to the genomic data, as assessed by the JSDs (Wilcoxon rank-sum test, W = 3,988, two-sided P = 0.0135; Table 1 and Supplementary Fig. 5). Precisely replicating the observed population dynamics was difficult (Fig. 3c and Supplementary Fig. 9), owing to the sparser sampling, particularly post-PCV7, and inevitable bias towards more invasive genotypes in this dataset. While the estimated strength of NFDS was similar to both Massachusetts and Southampton, the estimated vaccine selection strength was lower than in these infant carriage surveillance projects, consistent with the Nijmegen collection being isolated in an adult population primarily protected by herd immunity18. Correspondingly, fitting the heterogenous-rate model to the Maela dataset, isolated from an entirely unvaccinated community, estimated v close to zero (Table 1).
NFDS acting on genomic islands can also affect variation in the core genome. Comparisons between pre- and post-vaccination populations, and between different locations, revealed allele frequencies of core genome single-nucleotide polymorphisms (SNPs) typically showed very similar correlations to those of accessory loci frequencies (Supplementary Fig. 10). This was not a consequence of tight linkage between SNPs in the regions flanking genomic islands (Supplementary Fig. 10). Nevertheless, simulations in which NFDS acted on only accessory loci precisely replicated the post-vaccination changes in the core SNP allele frequencies, and similar correlations to those between collections were observed in simulations where the Massachusetts population was gradually replaced with isolates from other datasets (Supplementary Fig. 10). Therefore, although it is possible that core genome loci may also be under NFDS, the observed correlations can be attributed to NFDS acting only on accessory loci.
Consequences of NFDS for the impact of vaccination
Simulations were used to investigate counterfactual scenarios. In the absence of vaccination (v = 0), the pre-PCV7 populations were stable in Massachusetts (Supplementary Fig. 7), Southampton (Supplementary Fig. 8) and Nijmegen (Supplementary Fig. 9). Eliminating migration (m = 0) significantly increased the proportion of vaccine serotypes in simulations of all three populations (Wilcoxon rank-sum tests; Massachusetts, W = 0, two-sided P = 2.56 × 10−34; Southampton, W = 0, two-sided P = 2.56 × 10−34; Nijmegen, W = 1,453, two-sided P = 4.50 × 10−18), highlighting the importance of imported or previously rare NVTs in driving out vaccine types. However, removing NFDS significantly decreased the proportion of vaccine serotypes in simulations of all three populations (Wilcoxon rank-sum tests; W = 10,000, two-sided P = 2.56 × 10−34 in Massachusetts and Southampton; W = 9,979, two-sided P = 4.81 × 10−34 in Nijmegen). This is because following vaccination, those loci enriched in vaccine-type genotypes become increasingly advantageous to their bacterial hosts as they become rarer, resulting in NFDS slowing the rate at which vaccine-type isolates are eliminated until such loci rise in frequency in NVT genotypes.
Discussion
These combined analyses of multiple population genomic datasets suggest that NFDS plays an important part both in the stable structuring of pneumococcal populations and in their dynamics following disruption by vaccine-induced immunity. According to the best-fitting model, relatively strong NFDS acts on a few hundred accessory genes, corresponding to 5.0% of the Massachusetts pangenome and 8.3% of that in Southampton. This cumulative effect across multiple loci in complex populations is predicted to maintain stable lineage compositions in the absence of disruption by vaccination, without the oscillatory dynamics associated with some single-locus NFDS processes19,30,47,48. Therefore, multiple lineages can persistently coexist within this framework despite their confinement to a niche, the human nasopharynx, that is physiochemically homogeneous compared to the varied environments inhabited by species often considered as split into ecotypes, such as Escherichia coli. Furthermore, although intraspecific recombinations are slow over the timescales simulated in this study14, horizontal DNA transfer has comprehensively reassorted genomic islands between genotypes over the species’ history. Their consequent polyclonal distribution means accessory locus frequencies can be preserved by multiple lineage combinations, thereby accounting for the diverse population structures that are observed globally, and the panoply of strains that they contain49. Although the NFDS processes represented in the multilocus model were sufficient to explain the major post-vaccination population changes, further work is required to determine whether core loci are also involved. Such continued development of quantitative models with large genomic datasets should improve our understanding of how diverse selective pressures affecting bacterial populations shape their response to public health interventions, and how best to design novel pathogen-control strategies.
Methods
Annotation of the accessory genome
The previously analysed Massachusetts population14,50 contained 1,112 COGs present in between 5% and 95% of the 616 isolates and 1,194 COGs present in a single copy in every isolate. Information on whether these were associated with capsule polysaccharide synthesis, antibiotic resistance, restriction modification systems, pneumococcal pathogenicity island 1 or MGEs was extracted from previously described analyses14,21,50. Coding sequences (CDS) associated with proteinaceous immunogenic structures were identified through the results of protein antigen array data35. Candidate bacteriocins were identified using the BAGEL3 algorithm51. The variation at the blp locus, and the other putative bacteriocin-production loci, was manually identified within de novo assemblies of the Massachusetts isolates using Artemis and ACT52. The heat map showing the distribution of the blp alleles in Supplementary Fig. 1 was generated by mapping Illumina reads for each of the Massachusetts isolates against the concatenated set of loci using BWA with default settings53. Further information on COG functional domains14 and previous automated annotations50 was additionally used to manually curate all available information into the annotation and classification in Supplementary Datasets 1, 2.
Bioinformatic analysis of genomic data
The isolate collections analysed each came from systematic sampling of defined host populations. The Massachusetts pneumococcal dataset, isolated from the nasopharynxes of children of up to six years of age during routine primary care physician visits, consisted of the 616 de novo assemblies generated with Velvet54 as described previously14,50. VelvetOptimiser55 was used to assemble data from the Maela collection38 (3,085 genomes), isolated from the nasopharynxes of infants of up to two years of age, and their mothers, in a Thai refugee camp; the Southampton collection16 (516 genomes), isolated from the nasopharynxes of children of up to four years of age during outpatient visits; and the Nijmegen collection18 (337 genomes), isolated from adults hospitalized with bacteraemic pneumonia. These were supplemented with 20 complete, publically available reference genomes (Supplementary Dataset 3). To standardize these genome collections relative to the Massachusetts dataset, assemblies were discarded if they were less than 1.98 Mb, or greater than 2.19 Mb, in length; had an N 50 less than 15 kb14,50; or necessary information was absent from the public databases. Of the 4,586 genomes, 4,462 met these criteria and were included in a preliminary analysis that identified non-pneumococcal streptococci, which were then excluded from the final analysis. Consequently, the final dataset of 4,127 genomes contained 20 reference sequences, 616 Massachusetts sequences, 491 Southampton sequences, 337 Nijmegen sequences and 2,663 Maela sequences.
Each genome was processed with RNAmmer v.1.2 to annotate rRNA56; tRNAscan-SE v.1.3.1 to annotate tRNA57; Rfam scan to annotate other non-coding RNA58; scanned for BOX, RUP and SPRITE repeats using HMM profiles59,60; and Prodigal v.2.6 (ref. 61) to annotate CDS using a model trained on the genome of S. pneumoniae ATCC 700669 (ref. 62). CDS that overlapped with the non-coding RNA or short interspersed repeat sequences were then removed from the annotation, and the remaining set was translated to allow a non-redundant set of proteins to be identified. A version of the protein sequence dataset without low-complexity regions was generated by filtering these with segmasker63 and masking choline-binding domains. All-against-all comparisons of these protein databases were then generated using BLAT v.0.34 (ref. 64). gCOGs were then generated using COGtriangles and COGcognitor65, and through linking pairs of highly similar sequences, as described previously14. The gCOG nomenclature was then applied to the full, redundant set of protein sequences.
To correct for misassemblies, particularly those reflecting differences between the methods used to assemble the genomes of the Massachusetts isolates and those from other populations, false-positive CDS were eliminated from the intermediate-frequency gCOGs. A database generated from the annotation of S. pneumoniae ATCC 700669 (ref. 62) was used to search intermediate-frequency gCOG DNA sequences using BLASTALL v.2.2.25. This identified 39 gCOGs corresponding to fragments of tRNA, oligomers of choline-binding domains, or antisense fragments of insertion sequences. This left a final set of 11,049 gCOGs, of which 1,731 were present at a frequency between 5% and 95% in the pre- or peri-vaccination samples (grouped as pre-vaccination samples in the Results) of at least one of the four study populations.
To transfer the functional annotation onto the gCOG sequences, the annotated protein sequences from Massachusetts in Supplementary Table 1 were used to identify identical proteins in the gCOG dataset. When COGs could not be matched to gCOGs through this approach, links were instead made by searching gCOGs for proteins with identity to the middle 50% of annotated protein sequences from Massachusetts. These links were then manually curated to categorize the 1,731 intermediate-frequency gCOG sequences where possible, as shown in Fig. 2.
Analysis of population structure
To analyse the overall population structure, a ‘relaxed’ core set of 1,447 gCOGs were identified that met two criteria: first, that they were present in at least 95% of the isolates; and second, that the total number of gCOG representatives was less than 105% of the number of isolates containing the gCOG, to exclude gCOGs that are present in high copy number in some, or all, genomes. A codon alignment was then generated for each gCOG using mafft v.7.221 (ref. 66), excluding any sequences from isolates containing more than one representative of the gCOG. These were concatenated, with gap sites used to pad regions where data were missing for a particular isolate, and a 293,508 bp alignment of polymorphic sites extracted using SNP-sites67. A phylogeny was generated from this alignment using FastTree2 with the ‘fastest’ option68.
Lineages were identified using hierarchical BAPS clustering69. Five independent runs of the estimation algorithm, starting from the upper bound of 200–500 clusters, all converged to the same posterior mode. Two polyphyletic primary BAPS clusters were split into their secondary level clusters, yielding 73 sequence clusters that were almost entirely congruent with the phylogeny, and SC0, which remained polyphyletic. The monophyletic sequence clusters that were most similar to those in Massachusetts14 were numbered accordingly. The plot in Fig. 1c combined cophenetic distances from the core genome phylogeny, extracted with Bioperl70, and the Jaccard distance calculated from the presence and absence matrix of gCOGs using the R package vegan71. For each isolate, 100 comparator isolates were selected at random, and this sample of pairwise comparisons was used to generate the plot.
Of the polymorphic sites in the core genome, 282,043 corresponded to a base in the S. pneumoniae ATCC 700669 reference genome. For each population, the set of sites that were both biallelic and had a non-reference allele frequency between 5% and 95% in that population were extracted with VCFtools v.0.1.14 (ref. 72); there were 27,616 of these in the Massachusetts dataset, 26,954 in the Southampton dataset, 28,396 in the Nijmegen dataset and 30,579 in the Maela dataset. The r 2 statistics between these polymorphic sites, and between the binary presence and absence information of accessory gCOGs with a representative in the S. pneumoniae ATCC 700669 genome, were then calculated with VCFtools by treating each isolate as a phased haplotype. These were used to generate the linkage analysis plots in Supplementary Fig. 10.
Inference of antibiotic-resistance profiles
Genotypes of individual isolates were used to predict their antimicrobial resistance profiles. The presence of aph3′ (the gCOG CLS350021) was inferred to cause resistance to aminoglycosides; the presence of tetM (CLS03712) was inferred to cause resistance to tetracycline; the presence of cat (CLS01043) was inferred to cause resistance to chloramphenicol; and the presence of ermB (CLS01283), mef (CLS02227), or both was inferred to cause macrolide resistance62,73. These gCOGs themselves were removed from the set of loci used in the simulations, and the inferred antibiotic-resistance phenotype was used instead.
Non-susceptibility to other antibiotics is determined by core genome loci; to incorporate these into the model, resistant alleles of relevant loci were treated analogously to the presence of an accessory resistance gene. The presence of the I100L substitution in the dihydrofolate reductase protein (CLS03211) was inferred to result in resistance to trimethoprim74,75 and the presence of an insertion shortly after S61 in the dihydropteroate synthase protein (CLS01442) was inferred to result in resistance to sulphamethoxazole76. Three penicillin-binding proteins substantially contribute to β-lactam resistance. Using a similar approach to ref. 77, the population-wide protein sequences of Pbp1A (CLS01776), Pbp2X (CLS01031) and Pbp2B (CLS01093) were aligned with mafft v.7.221 (ref. 66), and the transpeptidase domain regions extracted. Following validation using the isolates from Massachusetts14, sequences exhibiting less than 97% amino acid identity with the susceptible alleles defined by in ref. 77 in the multiple sequence alignment were considered resistance-associated. These antibiotic-resistance phenotypes were included as intermediate-frequency loci if they met the criteria for a given population.
Multilocus NFDS model
The multilocus NFDS model was generated within a discrete-time Wright–Fisher framework78,79. Although such models were designed with a number of strong assumptions, the results of simulations have been found to be robust to violations of these conditions80. Each individual i had a genotype g i defined by a binary string representing the presence and absence of each gCOG or antibiotic-resistance phenotype present at an intermediate frequency in the starting population. The number of offspring arising from i at time t is a Poisson-distributed random variable X i,t . This Poisson approximation is justifiable if only a small proportion of descendants survive to the next generation78, as is likely to be the case for a nasopharyngeal colonizer with a small within-host effective population size81 that experiences a strong bottleneck at transmission. To allow for differential reproductive success between genotypes in a manner that depended on the composition of the overall population, X i,t was parameterized using the function (Supplementary Fig. 4):
The four components of the function each correspond to a different biological process. General density-dependent selection depends on κ, the carrying capacity of the environment, and N t , the total number of individuals at time t. This maintained an approximately stable population size throughout simulations. This is justifiable, as S. pneumoniae colonization levels did not substantially change in the years immediately after the introduction of PCV7.
Migration into the population occurred at rate m, subject to the limits 0 ≤ m ≤ 1, and therefore the reproductive fitness of resident individuals was reduced by a factor of (1 − m) accordingly to maintain an approximately constant population size of κ. The number of immigrating individuals at time t, N m,t , was a random variable calculated as:
Migrant individuals were selected, with replacement, from all isolates observed at any time point in the geographically specified dataset being studied. Therefore it was the only mechanism by which genotypes not present in the pre-vaccine genome samples could enter the simulated population. To prevent artefactually improving the fit of the model at high values of m through sampling all isolates in proportion to their observed frequency, the selection of an immigrating isolate was biased such that it was equally likely to come from any sequence cluster with at least one representative in the studied population, although these were present at very different frequencies within each population. Therefore, the probability of an immigrating individual being of genotype i and sequence cluster s, p m,s,i , was:
where S is the number of sequence clusters in the population, n s,i is the number of isolates in sequence cluster s of genotype i in the genome dataset, and n s is the number of isolates in the sequence cluster s in the genomic dataset.
The vaccine selection pressure to which individual i was subject, v i , depended on whether the individuals were of a vaccine serotype or not; for PCV7, the vaccine serotypes were 4, 6B, 9V, 14, 18C, 19F and 23F, as well as 6A, a vaccine-related type to which PCV7 elicited strong cross-immunity14. Consequently, v i was determined as:
where v was subject to the constraint 0 ≤ v ≤ 1.
In the homogeneous-rate multilocus model of NFDS, the magnitude of this pressure was determined by the term \({(1+{\sigma }_{f})}^{{\pi }_{i,t}}\), where σ f ≥ 0. The selection pressure depended on the genotype g i and distribution of intermediate-frequency loci at time t, as summarized by the exponent π i,t . The calculation of π i,t necessitated determining the frequency f l,t of each locus l at time t in the simulation, using the binary variables g i,l that represent the presence or absence of l in i:
These were compared to the equilibrium frequencies, e l , of the same loci, which were assumed to correspond to their frequencies in the sample of G 0 genomes from isolates sampled pre- or peri-vaccination:
The overall deviation of the L accessory genome loci included in the simulations, for individual i at time t, π i,t , was calculated as:
Therefore, if all accessory genes are at their equilibrium frequencies, then \({\left(1+{\sigma }_{f}\right)}^{{\pi }_{i,t}}\) = 1, and NFDS has no effect on an individual’s reproductive fitness. When a genotype contains many genes rarer than their equilibrium frequencies, \({\left(1+{\sigma }_{f}\right)}^{{\pi }_{i,t}}\) > 1, and NFDS increases an individual’s reproductive fitness. And when a genotype contains many genes more common than their equilibrium frequencies, \({\left(1+{\sigma }_{f}\right)}^{{\pi }_{i,t}}\) < 1, and therefore NFDS reduces an individual’s reproductive fitness. In the absence of l from an individual’s genotype, f l,t has no direct effect on its fitness.
Extension to heterogeneous frequency-dependent selection
Two further parameters were introduced when accessory genes were split into two categories, each subject to a different level of frequency-dependent selection. The σ w parameter represented the strength of weaker NFDS acting on a fraction, (1 − p f ), of the accessory genes included in the model. To facilitate inference of these two parameters, it was assumed that loci under weaker NFDS would vary in frequency to a greater extent between the initial and final genomic samples; therefore the accessory loci were ordered by the statistic Δ l :
where e l is the frequency of the gCOG or antibiotic-resistance phenotype across all pre- or peri-vaccination samples, as defined previously, and f l,t>0 is its frequency across all post-vaccination samples. The denominator is intended to emphasize the effects of gCOGs at frequencies of approximately 50%, which are likely to have a large effect on the overall population structure. The proportion p f of genes for which Δ l was smallest were considered subject to NFDS with strength (1 + σ f ), whereas the rest where subject to NFDS of strength (1 + σ w ). If the L loci were ordered by ascending values of Δ l , then l f was the highest ranking meeting the criterion, \(\frac{{l}_{f}}{L}\le {p}_{f}\). This resulted in two distinct measurements of the deviation of f l,t from e l :
and
Therefore, the modified offspring distribution was:
Simulations and parameter estimation
The model was implemented in C++using the GNU scientific library, and is available for download from https://github.com/nickjcroucher/multilocusNFDS. In each simulation, genotypes were represented by the gCOGs and antibiotic-resistance phenotypes present in between 5% and 95% of the pre- or peri-vaccination population. Therefore, L was 1,090 for Massachusetts, 1,175 for Southampton, 1,090 for Nijmegen and 1,254 for Maela. For simplicity, κ was assumed to represent the number of pneumococci likely to transmit between individuals in the sampled population. This was estimated to correspond to 25% colonization of children under ten years of age in the USA and European samples. In Massachusetts15, an under ten population of 828,129 in 2000 (ref. 82) implied a bacterial population size of 2 × 105 (105 was actually used for model fitting for computational efficiency; comparing simulations demonstrated that this had no detectable effect on the results); in Southampton (including Hampshire and Portsmouth), an under ten population of 202,404 in 2011 (ref. 83) implied a bacterial population size of 5 × 104; and in Nijmegen (including Arnhem), an under ten population of 77,753 in 2011 (ref. 84) implied a bacterial population of 2 × 104. An increased colonization rate of 50% (ref. 85) was used for Maela, where estimating that 15% of the 40,000 residents were under 10 implied a bacterial population size of 3 × 103.
Each simulation was run for a number of timesteps corresponding to the number of months spanned by the genomic collection, excluding early or late years in which sampling was sparse. The well-sampled periods were the 72 months between spring 2001 and spring 2007 for Massachusetts50; the 48 months between spring 2007 and spring 2011 for Southampton16; the 120 months between 2001 and 2011 for Nijmegen18; and the 24 months between 2007 and 2009 for Maela85. All isolates from a single winter were assigned to the year in which the season ended. In simulations of the Nijmegen population, where a substantial proportion of samples pre-dated the introduction of the vaccine, v = 0 for years up to 2007. In each case, the starting population for the simulation, of size κ, was generated by randomly resampling with replacement from the genotypes present in the pre- and peri-vaccination samples in each study; therefore, the ‘pre-vaccination’ population consisted of isolates sampled up to spring 2001 in Massachusetts, up to spring 2007 in Southampton, and up to 2007 in Nijmegen. These were the genomic samples used to calculate e l for all intermediate-frequency loci; all later samples were used to calculate f l,t>0 in the definition of Δ l .
At each time t at which a genomic sample was available, the equivalent number of genotypes was randomly sampled from the simulated population. The similarity between the simulated and genomic samples at t was then calculated as the Jensen–Shannon divergence86 (JSD t ) between the real and simulated samples:
Where f t,s,v is the simulated frequency of genotypes of sequence cluster s and vaccine-type status v at time t, and a t,s,v is the equivalent value from the genomic sample. This value was summed over all vaccine-type statuses and sequence clusters for each timepoint sampled in the genomic dataset to calculate the overall divergence of the simulation from the sampled data.
Each set of simulations was run with variation in the parameters v (range 0–0.5); m (range 0–0.2); σ f (range 10−6–0.22); σ w (range 10−6–0.15; only in the heterogeneous-rate model), and p f (range 0–1; only in the heterogeneous-rate model). Model fitting was achieved through Approximate Bayesian Computation with the BOLFI algorithm40, run for 2,000 iterations of Bayesian optimization to identify best-fitting parameter sets by minimizing the JSD (Table 1 and Supplementary Fig. 5). Point estimates of parameter values were generated based on the Gaussian process minimizers, with the distribution of the projected JSD values shown for each fit in Supplementary Fig. 5. Exploration of parameter space was performed with logarithmically transformed values to avoid discontinuity of the approximate likelihood function near the natural boundary and to enable better fit of the Gaussian process regression. The 95% posterior credible intervals for the parameters were obtained using three generations of sequential Monte Carlo sampling with the same default settings as used in ref. 40 for the pneumococcal daycare centre transmission model.
Alternative model formulations
To test whether equivalently good fits to the genomic data could be achieved using different approaches within the same framework, alternative model formulations were tested. The neutral model was fitted in the same way as the multilocus NFDS models, except that σ f was fixed at zero. The serotype NFDS model assumed all serotypes were present at equilibrium frequencies in the pre-vaccine samples, and therefore π i,t was calculated as the deviation of an isolate’s serotype from its initial frequency. This was fitted using both the homogeneous and heterogeneous selection rate models. In the latter case, Δ l was calculated by comparing the serotype e l values with their post-vaccination frequencies, as for the intermediate-frequency loci. The same parameter ranges were used as for the multilocus NFDS model, except σ f and σ w were allowed to take values within the range 10−6–25 to compensate for the single locus contributing to π i,t and ω i,t . Additionally, to avoid many lower frequency sequence clusters evolving neutrally, serotypes were considered to be at intermediate frequencies if they were between 1% and 99% prevalence in the pre- or peri-vaccination population.
The ecotype NFDS model assumed each sequence cluster was adapted to a specific ecological niche, and therefore was present at an equilibrium frequency in the pre-vaccine samples. Therefore, π i,t and ω i,t were calculated as the deviation of an isolate’s sequence cluster from its initial frequency. This was fitted using both the homogeneous and heterogeneous selection rate models using the same parameter ranges and intermediate-frequency range as for the serotype NFDS model, as well as the same approach to the calculation of Δ l . For both the serotype and ecotype models, fitting was conducted with BOLFI40 as for the multilocus NFDS model, using JSDs to quantify the differences between the simulated and sampled populations. Results are shown in Table 1. These implementations are not intended to represent the optimal versions of each model, but instead demonstrate that the fits of the multilocus NFDS models cannot be trivially replicated by changing the genetic basis of NFDS.
Simulations in which isolates from two populations were combined used the pre-vaccination population from Massachusetts and post-vaccine isolates from one of the alternative populations. The initial population was drawn only from Massachusetts; both these isolates, and those from the alternative dataset, could enter the simulated population through migration. When the alternative population corresponded to Southampton or Nijmegen, the population size, number of generations, parameter point estimates, Δ l and e l values were those of the alternative population. When the alternative population was Maela, the e l and Δ l values were those of the alternative population, but the simulations were otherwise parameterized for the Massachusetts population, due to the difficulty of obtaining robust point estimates for parameters from the Maela population as a consequence of the lack of vaccine introduction in this location.
Statistical analyses
Statistical analyses, including calculation of Pearson’s R 2, Wilcoxon tests, interquartile ranges and Fisher’s exact tests, were performed using R87. Estimation of parameter values and credibility intervals through model fitting were performed with BOLFI40. All reported P values are two-sided.
Code availability
The model code used in this analysis is freely available from the GitHub repository, https://github.com/nickjcroucher/multilocusNFDS.
Data availability
The sequence datasets analysed in the current study are available in the public sequence databases with the accession codes listed in Supplementary Dataset 3. The epidemiological and phylogenetic data analysed in the current study are available from https://microreact.org/project/multilocusNFDS.
References
Haegeman, B. & Weitz, J. S. A neutral theory of genome evolution and the frequency distribution of genes. BMC Genomics 13, 196 (2012).
Baumdicker, F., Hess, W. R. & Pfaffelhuber, P. The infinitely many genes model for the distributed genome of bacteria. Genome Biol. Evol. 4, 443–456 (2012).
Marttinen, P., Croucher, N. J., Gutmann, M. U., Corander, J. & Hanage W. P. Recombination produces coherent bacterial species clusters in both core and accessory genomes. Microb. Genom. 1, e000038 (2015).
Hogg, J. S. et al. Characterization and modeling of the Haemophilus influenzae core and supragenomes based on the complete genomic sequences of Rd and 12 clinical nontypeable strains. Genome Biol. 8, R103 (2007).
Collins, R. E. & Higgs, P. G. Testing the infinitely many genes model for the evolution of the bacterial core genome and pangenome. Mol. Biol. Evol. 29, 3413–3425 (2012).
Lobkovsky, A. E., Wolf, Y. I. & Koonin, E. V. Gene frequency distributions reject a neutral model of genome evolution. Genome Biol. Evol. 5, 233–242 (2013).
McInerney, J. O., McNally, A. & O’Connell, M. J. Why prokaryotes have pangenomes. Nat. Microbiol. 2, 17040 (2017).
Shapiro, B. J. et al. Population genomics of early events in the ecological differentiation of bacteria. Science 336, 48–51 (2012).
Cohan, F. M. Bacterial species and speciation. Syst. Biol. 50, 513–524 (2001).
Cohan, F. M. What are bacterial species? Annu. Rev. Microbiol. 56, 457–487 (2002).
Watkins, E. R. et al. Vaccination drives changes in metabolic and virulence profiles of Streptococcus pneumoniae. PLoS Pathog. 11, e1005034 (2015).
Regev-Yochay, G. et al. Re-emergence of the type 1 pilus among Streptococcus pneumoniae isolates in Massachusetts, USA. Vaccine 28, 4842–4846 (2010).
Cobey, S. & Lipsitch, M. Niche and neutral effects of acquired immunity permit coexistence of pneumococcal serotypes. Science 335, 1376–1380 (2012).
Croucher, N. J. et al. Population genomics of post-vaccine changes in pneumococcal epidemiology. Nat. Genet. 45, 656–663 (2013).
Huang, S. S. et al. Continued impact of pneumococcal conjugate vaccine on carriage in young children. Pediatrics 124, e1–11 (2009).
Gladstone, R. A. et al. Five winters of pneumococcal serotype replacement in UK carriage following PCV introduction. Vaccine 33, 2015–2021 (2015).
Gladstone, R. A. et al. Pre-vaccine serotype composition within a lineage signposts its serotype replacement — a carriage study over 7 years following pneumococcal conjugate vaccine use in the UK. Microb. Genom. 3, 119 (2017).
Cremers, A. J. H. et al. The post-vaccine microevolution of invasive Streptococcus pneumoniae. Sci. Rep. 5, 14952 (2015).
Levin, B. R. Frequency-dependent selection in bacterial populations. Phil. Trans. R. Soc. Lond. B 319, 459–472 (1988).
Maynard Smith, J. Evolutionary Genetics 2nd edn (Oxford Univ. Press, Oxford, 1998).
Croucher, N. J. et al. Diversification of bacterial genome content through distinct mechanisms over different timescales. Nat. Commun. 5, 5471 (2014).
Croucher, N. J. et al. Horizontal DNA transfer mechanisms of bacteria as weapons of intragenomic conflict. PLoS Biol. 14, e1002394 (2016).
Takeuchi, N., Cordero, O. X., Koonin, E. V. & Kaneko, K. Gene-specific selective sweeps in bacteria and archaea caused by negative frequency-dependent selection. BMC Biol. 13, 20 (2015).
Cordero, O. X. & Polz, M. F. Explaining microbial genomic diversity in light of evolutionary ecology. Nat. Rev. Microbiol. 12, 263–273 (2014).
Dawid, S., Roche, A. M. & Weiser, J. N. The blp bacteriocins of Streptococcus pneumoniae mediate intraspecies competition both in vitro and in vivo. Infect. Immun. 75, 443–451 (2007).
Miller, E. L., Abrudan, M. I., Roberts, I. S. & Rozen, D. E. Diverse ecological strategies are encoded by Streptococcus pneumoniae bacteriocin-like peptides. Genome Biol. Evol. 8, 1072–1090 (2016).
Bogaardt, C., van Tonder, A. J. & Brueggemann, A. B. Genomic analyses of pneumococci reveal a wide diversity of bacteriocins — including pneumocyclicin, a novel circular bacteriocin. BMC Genomics 16, 554 (2015).
Maricic, N., Anderson, E. S., Opipari, A. M. E., Yu, E. A. & Dawid, S. Characterization of a multipeptide lantibiotic locus in Streptococcus pneumoniae. mBio 7, e01656-15 (2016).
Hoover, S. E. et al. A new quorum-sensing system (TprA/PhrA) for Streptococcus pneumoniae D39 that regulates a lantibiotic biosynthesis gene cluster. Mol. Microbiol. 97, 229–243 (2015).
Kerr, B., Riley, M. A., Feldman, M. W. & Bohannan, B. J. M. Local dispersal promotes biodiversity in a real-life game of rock–paper–scissors. Nature 418, 171–174 (2002).
Stewart, F. M. & Levin, B. R. Partitioning of resources and the outcome of interspecific competition: a model and some general considerations. Am. Nat. 107, 171–198 (1973).
Levin, B. R. Coexistence of two asexual strains on a single resource. Science 175, 1272–1274 (1972).
Colijn, C. & Cohen, T. How competition governs whether moderate or aggressive treatment minimizes antibiotic resistance. eLife 4, e10559 (2015).
Lehtinen, S. et al. Evolution of antibiotic resistance is linked to any genetic mechanism affecting bacterial duration of carriage. Proc. Natl Acad. Sci. USA 114, 1075–1080 (2017).
Croucher, N. J. et al. Diverse evolutionary patterns of pneumococcal antigens identified by pangenome-wide immunological screening. Proc. Natl Acad. Sci. USA 114, E357–E366 (2017).
Croucher, N. J. et al. Selective and genetic constraints on pneumococcal serotype switching. PLoS Genet. 11, e1005095 (2015).
Bagnoli, F. et al. A second pilus type in Streptococcus pneumoniae is prevalent in emerging serotypes and mediates adhesion to host cells. J. Bacteriol. 190, 5480–5492 (2008).
Chewapreecha, C. et al. Dense genomic sampling identifies highways of pneumococcal recombination. Nat. Genet. 46, 305–309 (2014).
Goossens, H. et al. Outpatient antibiotic use in Europe and association with resistance: a cross-national database study. Lancet 365, 579–587 (2005).
Gutmann, M. U. & Corander, J. Bayesian optimization for likelihood-free inference of simulator-based statistical models. J. Mach. Learn. Res. 17, 1–47 (2016).
Lintusaari, J., Gutmann, M. U., Dutta, R., Kaski, S. & Corander, J. Fundamentals and recent developments in Approximate Bayesian Computation. Syst. Biol. 66, e66–e82 (2017).
Rinta-Kokko, H., Dagan, R., Givon-Lavi, N. & Auranen, K. Estimation of vaccine efficacy against acquisition of pneumococcal carriage. Vaccine 27, 3831–3837 (2009).
Lipsitch, M. et al. Estimating rates of carriage acquisition and clearance and competitive ability for pneumococcal serotypes in Kenya with a Markov transition model. Epidemiology 23, 510–519 (2012).
Health Protection Agency COVER programme. October to December 2008. Quarterly vaccination coverage statistics for children aged up to five years in the United Kingdom. Heal. Prot. Rep. 3, 8–15 (2009).
Nuorti, J. P., Martin, S. W., Smith, P. J., Moran, J. S. & Schwartz, B. Uptake of pneumococcal conjugate vaccine among children in the 1998–2002 United States birth cohorts. Am. J. Prev. Med. 34, 46–53 (2008).
Huang, S. S., Finkelstein, J. A., Rifas-Shiman, S. L., Kleinman, K. & Platt, R. Community-level predictors of pneumococcal carriage and resistance in young children. Am. J. Epidemiol. 159, 645–654 (2004).
Durrett, R. & Levin, S. Allelopathy in spatially distributed populations. J. Theor. Biol. 185, 165–171 (1997).
Gupta, S., Ferguson, N. & Anderson, R. Chaos, persistence, and evolution of strain structure in antigenically diverse infectious agents. Science 280, 912–915 (1998).
Henriques-Normark, B., Blomberg, C., Dagerhamn, J., Bättig, P. & Normark, S.The rise and fall of bacterial clones: Streptococcus pneumoniae. Nat. Rev. Microbiol. 6, 827–837 (2008).
Croucher, N. J. et al. Population genomic datasets describing the post-vaccine evolutionary epidemiology of Streptococcus pneumoniae. Sci. Data 2, 150058 (2015).
van Heel, A. J., de Jong, A., Montalbán-López, M., Kok, J. & Kuipers, O. P. BAGEL3: automated identification of genes encoding bacteriocins and (non-)bactericidal posttranslationally modified peptides. Nucleic Acids Res. 41, W448–W453 (2013).
Carver, T. et al. Artemis and ACT: viewing, annotating and comparing sequences stored in a relational database. Bioinformatics 24, 2672–2676 (2008).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Zerbino, D. R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Gladman, S. VelvetOptimiser (Victorian Bioinformatics Consortium, 2010); http://www.vicbioinformatics.com/software.velvetoptimiser.shtml
Lagesen, K. et al. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res. 35, 3100–3108 (2007).
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 (1997).
Burge, S. W. et al. Rfam 11.0: 10 years of RNA families. Nucleic Acids Res. 41, D226–D232 (2013).
Croucher, N. J., Vernikos, G. S., Parkhill, J. & Bentley, S. D. Identification, variation and transcription of pneumococcal repeat sequences. BMC Genomics 12, 120 (2011).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11, 119 (2010).
Croucher, N. J. et al. Role of conjugative elements in the evolution of the multidrug-resistant pandemic clone Streptococcus pneumoniae Spain23F ST81. J. Bacteriol. 191, 1480–1489 (2009).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Kent, W. J. BLAT—the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Kristensen, D. M. et al. A low-polynomial algorithm for assembling clusters of orthologous groups from intergenomic symmetric best matches. Bioinformatics 26, 1481–1487 (2010).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Page, A. J. et al. SNP-sites: rapid efficient extraction of SNPs from multi-FASTA alignments. Microb. Genom. 2, e000056 (2016).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2–approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Cheng, L., Connor, T. R., Sirén, J., Aanensen, D. M. & Corander, J. Hierarchical and spatially explicit clustering of DNA sequences with BAPS software. Mol. Biol. Evol. 30, 1224–1228 (2013).
Stajich, J. E. et al. The Bioperl toolkit: Perl modules for the life sciences. Genome Res. 12, 1611–1618 (2002).
Dixon, P. VEGAN, a package of R functions for community ecology. J. Veg. Sci. 14, 927–930 (2003).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Croucher, N. J. et al. Rapid pneumococcal evolution in response to clinical interventions. Science 331, 430–434 (2011).
Pikis, A., Donkersloot, J. A., Rodriguez, W. J. & Keith, J. M. A conservative amino acid mutation in the chromosome-encoded dihydrofolate reductase confers trimethoprim resistance in Streptococcus pneumoniae. J. Infect. Dis. 178, 700–706 (1998).
Maskell, J. P., Sefton, A. M. & Hall, L. M. C. Multiple mutations modulate the function of dihydrofolate reductase in trimethoprim-resistant Streptococcus pneumoniae. Antimicrob. Agents Chemother. 45, 1104–1108 (2001).
Haasum, Y. et al. Amino acid repetitions in the dihydropteroate synthase of Streptococcus pneumoniae lead to sulfonamide resistance with limited effects on substrate Km. Antimicrob. Agents Chemother. 45, 805–809 (2001).
Li, Y. et al. Penicillin-binding protein transpeptidase signatures for tracking and predicting β-lactam resistance levels in Streptococcus pneumoniae. mBio 7, e00756-16 (2016).
Fisher, R. A. The Genetical Theory of Natural Selection (Oxford Univ. Press, Oxford, 1930).
Wright, S. Evolution in Mendelian populations. Genetics 16, 97–159 (1931).
Der, R., Epstein, C. & Plotkin, J. B. Dynamics of neutral and selected alleles when the offspring distribution is skewed. Genetics 191, 1331–1344 (2012).
Li, Y., Thompson, C. M., Trzciński, K. & Lipsitch, M. Within-host selection is limited by an effective population of Streptococcus pneumoniae during nasopharyngeal colonization. Infect. Immun. 81, 4534–4543 (2013).
Census 2000 (US Census Bureau, 2000); https://www.census.gov/census2000/states/ma.html
Census 2011 (Office for National Statistics, 2011); http://www.ons.gov.uk/ons/guide-method/census/2011/index.html
Dutch Census 2011 (European Statistical System, 2011); https://ec.europa.eu/CensusHub2/
Turner, P. et al. A longitudinal study of Streptococcus pneumoniae carriage in a cohort of infants and their mothers on the Thailand–Myanmar border. PLoS ONE 7, e38271 (2012).
Wong, A. K. C. & You, M. Entropy and distance of random graphs with application to structural pattern recognition. IEEE Trans. Pattern Anal. Mach. Intell. 7, 599–609 (1985).
R Core Development Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, 2011); http://www.R-project.org/
Acknowledgements
We thank R. Gladstone, J. Jefferies, S. Faust and S. Clarke for sharing epidemiological data on the Southampton isolates. N.J.C. was funded by a Sir Henry Dale fellowship, and jointly funded by the Wellcome Trust and Royal Society (Grant Number 104169/Z/14/Z). J.C. was funded by the COIN Centre of Excellence. M.L. was funded by NIH grant R01 AI048935 and W.P.H. by NIH grant R01 AI106786.
Author information
Authors and Affiliations
Contributions
J.C., C.F., B.A., W.P.H., M.L. and N.J.C. designed the model; J.C., M.U.G. and N.J.C. fitted the model; W.P.H., S.D.B. and N.J.C. analysed the genomic data; J.C. and N.J.C. initially drafted the manuscript, with all authors contributing to the final version.
Corresponding author
Ethics declarations
Competing interests
M.L. has consulted for Pfizer, Affinivax and Merck and has received grant support not related to this paper from Pfizer and PATH Vaccine Solutions. W.P.H., M.L. and N.J.C. have consulted for Antigen Discovery Inc.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Information
Supplementary Figures 1–10; Supplementary Table 1; legends for Supplementary Datasets 1–3
Supplementary Dataset 1
Annotation of the intermediate frequency genes in the Massachusetts pneumococcal population
Supplementary Dataset 2
Annotation of the core genes in the Massachusetts pneumococcal population
Supplementary Dataset 3
Samples used in the analyses, associated epidemiological characteristics, and accession codes
Rights and permissions
About this article
Cite this article
Corander, J., Fraser, C., Gutmann, M.U. et al. Frequency-dependent selection in vaccine-associated pneumococcal population dynamics. Nat Ecol Evol 1, 1950–1960 (2017). https://doi.org/10.1038/s41559-017-0337-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-017-0337-x
This article is cited by
-
Genomic and panproteomic analysis of the development of infant immune responses to antigenically-diverse pneumococci
Nature Communications (2024)
-
Mathematical Modelling of Parasite Dynamics: A Stochastic Simulation-Based Approach and Parameter Estimation via Modified Sequential-Type Approximate Bayesian Computation
Bulletin of Mathematical Biology (2024)
-
The metabolic, virulence and antimicrobial resistance profiles of colonising Streptococcus pneumoniae shift after PCV13 introduction in urban Malawi
Nature Communications (2023)
-
Post-vaccine epidemiology of serotype 3 pneumococci identifies transformation inhibition through prophage-driven alteration of a non-coding RNA
Genome Medicine (2022)
-
Pneumococcal within-host diversity during colonization, transmission and treatment
Nature Microbiology (2022)