Abstract
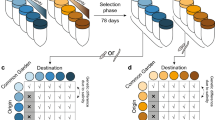
Dynamic interactions between ecological conditions and the phenotypic composition of populations likely play an important role in evolution, but the direction and strength of these feedbacks remain difficult to characterize. We investigated these dynamics across two generations of threespine sticklebacks from two evolutionary lineages undergoing secondary contact and hybridization. Independently manipulating the density and lineage of adults in experimental mesocosms led to contrasting ecosystem conditions with strong effects on total survival in a subsequent generation of juveniles. Ecosystem modifications by adults also varied the strength of selection on competing hybrid and non-hybrid juveniles. This variation in selection indicated (1) a negative eco-evolutionary feedback driven by lineage-specific resource depletion and dependence and (2) a large performance advantage of hybrid juveniles in depleted environments. This work illustrates the importance of interactions between phenotype, population density and the environment in shaping selection and evolutionary trajectories, especially in the context of range expansion with secondary contact and hybridization.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Reznick, D. Hard and soft selection revisited: how evolution by natural selection works in the real world. J. Hered. 107, 3–14 (2015).
Pelletier, F., Garant, D. & Hendry, A. P. Eco-evolutionary dynamics. Phil. Trans. R. Soc. Lond. B. 364, 1483–1489 (2009).
Matthews, B. et al. Under niche construction: an operational bridge between ecology, evolution, and ecosystem science. Ecol. Monogr. 84, 245–263 (2014).
Hendry, A. P. Eco-evolutionary Dynamics (Princeton Univ. Press, Princeton, NJ, 2016).
McPeek, M. A. The ecological dynamics of natural selection: traits and the coevolution of community structure. Am. Nat. 189, E91–E117 (2017).
Harmon, L. J. et al. Evolutionary diversification in stickleback affects ecosystem functioning. Nature 458, 1167–1170 (2009).
Des Roches, S., Shurin, J. B., Schluter, D. & Harmon, L. J. Ecological and evolutionary effects of stickleback on community structure. PLoS ONE 8, e59644 (2013).
Bassar, R. D. et al. Local adaptation in Trinidadian guppies alters ecosystem processes. Proc. Natl Acad. Sci. USA 107, 3616–3621 (2010).
Lundsgaard-Hansen, B., Matthews, B. & Seehausen, O. Ecological speciation and phenotypic plasticity affect ecosystems. Ecology 95, 2723–2735 (2014).
Bassar, R. D., Lopez-Sepulcre, A., Reznick, D. N. & Travis, J. Experimental evidence for density-dependent regulation and selection on Trinidadian guppy life histories. Am. Nat. 181, 25–38 (2013).
Matthews, B., Aebischer, T., Sullam, K. E., Lundsgaard-Hansen, B. & Seehausen, O. Experimental evidence of an eco-evolutionary feedback during adaptive divergence. Curr. Biol. 26, 483–489 (2016).
Palkovacs, E. P., Mandeville, E. G. & Post, D. M. Contemporary trait change in a classic ecological experiment: rapid decrease in alewife gill‐raker spacing following introduction to an inland lake. Freshwater Biol. 59, 1897–1901 (2014).
Fussmann, G. F., Loreau, M. & Abrams, P. A. Eco-evolutionary dynamics of communities and ecosystems. Funct. Ecol. 21, 465–477 (2007).
Becks, L., Ellner, S. P., Jones, L. E. & Hairston, N. G. The functional genomics of an eco-evolutionary feedback loop: linking gene expression, trait evolution, and community dynamics. Ecol. Lett. 15, 492–501 (2012).
Ferriere, R. & Legendre, S. Eco-evolutionary feedbacks, adaptive dynamics and evolutionary rescue theory. Phil. Trans. R. Soc. Lond. B. 368, 20120081 (2013).
Abrams, P. A. The eco-evolutionary responses of a generalist consumer to resource competition. Evolution 66, 3130–3143 (2012).
Cortez, M. H. How the magnitude of prey genetic variation alters predator–prey eco-evolutionary dynamics. Am. Nat. 188, 329–341 (2016).
Hairston, N. G., Ellner, S. P., Geber, M. A., Yoshida, T. & Fox, J. A. Rapid evolution and the convergence of ecological and evolutionary time. Ecol. Lett. 8, 1114–1127 (2005).
Lankau, R. A. & Strauss, S. Y. Community complexity drives patterns of natural selection on a chemical defense of Brassica nigra. Am. Nat. 171, 150–161 (2008).
Barrett, R. D. H. & Schluter, D. Adaptation from standing genetic variation. Trends Ecol. Evol. 23, 38–44 (2008).
Cameron, T. C., Plaistow, S., Mugabo, M., Piertney, S. B. & Benton, T. G. Chapter five—eco-evolutionary dynamics: experiments in a model system. Adv. Ecol. Res. 50, 171–206 (2014).
Turcotte, M. M., Reznick, D. N. & Hare, J. D. Experimental test of an eco-evolutionary dynamic feedback loop between evolution and population density in the green peach aphid. Am. Nat. 181, S46–S57 (2013).
Jones, A. W. & Post, D. M. Consumer interaction strength may limit the diversifying effect of intraspecific competition: a test in alewife (Alosa pseudoharengus). Am. Nat. 181, 815–826 (2013).
Svanbäck, R. & Bolnick, D. I. Intraspecific competition drives increased resource use diversity within a natural population. Proc. R. Soc. Lond. B 274, 839–844 (2007).
Schluter, D. Experimental evidence that competition promotes divergence in adaptive radiation. Science 266, 798–801 (1994).
Rundle, H. D., Vamosi, S. M. & Schluter, D. Experimental test of predation’s effect on divergent selection during character displacement in sticklebacks. Proc. Natl Acad. Sci. USA 100, 14943–14948 (2003).
Lucek, K., Roy, D., Bezault, E., Sivasundar, A. & Seehausen, O. Hybridization between distant lineages increases adaptive variation during a biological invasion: stickleback in Switzerland. Mol. Ecol. 19, 3995–4011 (2010).
Berner, D., Roesti, M., Hendry, A. P. & Salzburger, W. Constraints on speciation suggested by comparing lake-stream stickleback divergence across two continents. Mol. Ecol. 19, 4963–4978 (2010).
Lucek, K., Sivasundar, A., Roy, D. & Seehausen, O. Repeated and predictable patterns of ecotypic differentiation during a biological invasion: lake–stream divergence in parapatric Swiss stickleback. J. Evol. Biol. 26, 2691–2709 (2013).
Lucek, K., Sivasundar, A., Roy, D. & Seehausen, O. Dryad Data from: Repeated and predictable patterns of ecotypic differentiation during a biological invasion: lake–stream divergence in parapatric Swiss stickleback. (Dryad Digital Repository, 2013); http://datadryad.org/resource/doi:10.5061/dryad.0nh60.
Berner, D., Moser, D., Roesti, M., Buescher, H. & Salzburger, W. Genetic architechture of skeletal evolution in European lake and stream stickleback. Evolution 68, 1792–1805 (2014).
Schluter, D. Adaptive radiation in sticklebacks: trade-offs in feeding performance and growth. Ecology 76, 82–90 (1995).
McGee, M. D., Schluter, D. & Wainwright, P. C. Functional basis of ecological divergence in sympatric stickleback. BMC Evol. Biol. 13, 1–10 (2013).
Roy, D., Lucek, K., Walter, R. P. & Seehausen, O. Hybrid ‘superswarm’ leads to rapid divergence and establishment of populations during a biological invasion. Mol. Ecol. 24, 5394–5411 (2015).
Canestrelli, D. et al. The tangled evolutionary legacies of range expansion and hybridization. Trends Ecol. Evol. 31, 677–688 (2016).
Nosil, P. Ecological Speciation (Oxford Univ. Press, Oxford, 2012).
Hatfield, T. & Schluter, D. Ecological speciation in sticklebacks: environment-dependent hybrid fitness. Evolution 53, 866–873 (1999).
Gill, A. B. & Hart, P. J. B. Feeding behaviour and prey choice of the threespine stickleback: the interacting effects of prey size, fish size and stomach fullness. Anim. Behav. 47, 921–932 (1994).
Gergs, R. & Rothhaupt, K. O. Invasive species as driving factors for the structure of benthic communities in Lake Constance, Germany. Hydrobiologia 746, 245–254 (2015).
Janzen, D. H. Herbivores and number of tree species in tropical forests. Am. Nat. 104, 501–528 (1970).
Vamosi, S. M. & Schluter, D. Impacts of trout predation on fitness of sympatric sticklebacks and their hybrids. Proc. R. Soc. Lond. B 269, 923–930 (2002).
Arnegard, M. E. et al. Genetics of ecological divergence during speciation. Nature 511, 307–311 (2014).
Taylor, E. B. et al. Speciation in reverse: morphological and genetic evidence of the collapse of a three-spined stickleback (Gasterosteus aculeatus) species pair. Mol. Ecol. 15, 343–355 (2006).
Rieseberg, L. H. et al. Hybridization and the colonization of novel habitats by annual sunflowers. Genetica 129, 149–165 (2007).
Hochholdinger, F. & Hoecker, N. Towards the molecular basis of heterosis. Trends Plant Sci. 12, 427–432 (2007).
Stelkens, R. & Seehausen, O. Genetic distance between species predicts novel trait expression in their hybrids. Evolution 63, 884–897 (2009).
Blanchet, S., Bernatchez, L. & Dodson, J. J. Does interspecific competition influence relationships between heterozygosity and fitness-related behaviors in juvenile Atlantic salmon (Salmo salar)? Behav. Ecol. Sociobiol. 63, 605–615 (2009).
Brock, P. M., Goodman, S. J., Hall, A. J., Cruz, M. & Acevedo-Whitehouse, K. Context-dependent associations between heterozygosity and immune variation in a wild carnivore. BMC Evol. Biol. 15, 242 (2015).
Domínguez, A. & Albornoz, J. Environment-dependent heterosis in Drosophila melanogaster. Genet. Sel. Evol. 19, 37–48 (1987).
Blum, A. Heterosis, stress, and the environment: a possible road map towards the general improvement of crop yield. J. Exp. Bot. 64, 4829–4837 (2013).
Anaya-Rojas, J. M. Host–Parasite Interactions and the Eco-Evolutionary Dynamics of Aquatic Ecosystems PhD thesis, Univ. Bern (2017).
Lucek, K., Sivasundar, A., Kristjánsson, B., Skúlason, S. & Seehausen, O. Quick divergence but slow convergence during ecotype formation in lake and stream stickleback pairs of variable age. J. Evol. Biol. 27, 1878–1892 (2014).
Jones, O. R. & Wang, J. COLONY: a program for parentage and sibship inference from multilocus genotype data. Mol. Ecol. Res. 10, 551–555 (2010).
R Development Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, 2012).
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. http://dx.doi.org/10.18637/jss.v067.i01/ (2015).
Fox, J. & Weisberg, S. An R Companion to Applied Regression (Sage, Thousand Oaks, CA, 2011).
Burnham, K. P. & Anderson, D. R. Model Selection and Multimodel Inference (Springer, Berlin, 2002).
Schmid, D. About the Role of Resource Environment in the Population Divergence of the Lake Constance Threespine Stickleback MSc thesis, Univ. Zurich (2016).
Acknowledgements
We thank M. Lürig, G. Antoniazza, E. Birnstiel, L. Catalano, K. Müller, A. Taverna, E. Schäffer, F. Brunner, D. Hohmann and D. Steiner for major contributions to fish breeding and care, as well as experimental set-up, maintenance and sampling. We thank K. Lucek, S. Mwaiko and C. Schmid for assistance with microsatellite genotyping, and D. Marques and M. McGee for input on the study system and experimental design. S. Robert, P. Kathriner and B. Kienholz provided laboratory facilities and infrastructure support. We thank M. Rheinhof for access to sampling sites on Lake Constance.
Author information
Authors and Affiliations
Contributions
B.M., J.M.A.-R., O.S. and R.J.B. designed the core experiment. R.J.B. and J.M.A.-R. carried out the experiment. M.C.L. designed and carried out the isotopic analysis, and D.W.S. provided supplementary data on feeding efficiency. R.J.B. analysed the data and wrote the manuscript with substantial contributions and revisions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing finacial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Information
Supplementary Figures 1 and 2, Supplementary Tables 1–3
Rights and permissions
About this article
Cite this article
Best, R.J., Anaya-Rojas, J.M., Leal, M.C. et al. Transgenerational selection driven by divergent ecological impacts of hybridizing lineages. Nat Ecol Evol 1, 1757–1765 (2017). https://doi.org/10.1038/s41559-017-0308-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-017-0308-2
This article is cited by
-
Horizontal gene transfer is predicted to overcome the diversity limit of competing microbial species
Nature Communications (2024)
-
How warp-speed evolution is transforming ecology
Nature (2018)