Abstract
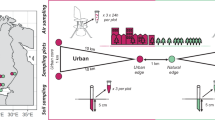
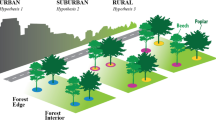
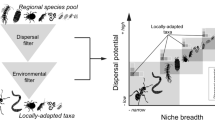
Urbanization alters the physicochemical environment, introduces non-native species and causes ecosystem characteristics to converge. It has been speculated that these alterations contribute to loss of regional and global biodiversity, but so far most urban studies have assessed macro-organisms and reported mixed evidence for biodiversity loss. We studied five cities on three continents to assess the global convergence of urban soil microbial communities. We determined the extent to which communities of bacteria, archaea and fungi are geographically distributed, and to what extent urbanization acts as a filter on species diversity. We discovered that microbial communities in general converge, but the response differed among microbial domains; soil archaeal communities showed the strongest convergence, followed by fungi, while soil bacterial communities did not converge. Our data suggest that urban soil archaeal and bacterial communities are not vulnerable to biodiversity loss, whereas urbanization may be contributing to the global diversity loss of ectomycorrhizal fungi. Ectomycorrhizae decreased in both abundance and species richness under turf and ruderal land-uses. These data add to an emerging pattern of widespread suppression of ectomycorrhizal fungi by human land-uses that involve physical disruption of the soil, management of the plant community, or nutrient enrichment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ellis, E. C. & Ramankutty, N. Putting people in the map: anthropogenic biomes of the world. Front. Ecol. Environ. 6, 439–447 (2007).
Foster, D. et al. The importance of land-use legacies to ecology and conservation. Bioscience 53, 77–88 (2003).
Rothwell, A., Ridoutt, B., Page, G. & Bellotti, W. Feeding and housing the urban population: environmental impacts at the peri-urban interface under different land-use scenarios. Land Use Policy 48, 377–388 (2015).
Scheffer, M., Carpenter, S., Foley, J. A., Folke, C. & Walker, B. Catastrophic shifts in ecosystems. Nature 413, 591–596 (2001).
Pouyat, R. V. et al. A global comparison of surface soil characteristics across five cities: a test of the urban ecosystem convergence hypothesis. Soil Sci. 180, 136–145 (2015).
Groffman, P. M. et al. Ecological homogenization of urban USA. Front. Ecol. Environ. 12, 74–81 (2014).
McKinney, M. L. & Lockwood, J. L. Biotic homogenization: a few winners replacing many losers in the next mass extinction. Trends Ecol. Evol. 14, 450–453 (1999).
Baiser, B., Olden, J. D., Record, S., Lockwood, J. L. & McKinney, M. L. Pattern and process of biotic homogenization in the New Pangaea. Proc. R. Soc. B 279, 4772–4777 (2012).
Tabarelli, M., Peres, C. A. & Melo, F. P. L. The ‘few winners and many losers’ paradigm revisited: emerging prospects for tropical forest biodiversity. Biol. Conserv. 155, 136–140 (2012).
McKinney, M. L. Urbanization as a major cause of biotic homogenization. Biol. Conserv. 127, 247–260 (2006).
Olden, J. D. & Rooney, T. P. On defining and quantifying biotic homogenization. Glob. Ecol. Biogeogr. 15, 113–120 (2006).
Fukami, T., Martijn Bezemer, T., Mortimer, S. R. & van der Putten, W. H. Species divergence and trait convergence in experimental plant community assembly. Ecol. Lett. 8, 1283–1290 (2005).
Myers, N., Mittermeier, R. A., Mittermeier, C. G., da Fonseca, G. A. B. & Kent, J. Biodiversity hotspots for conservation priorities. Nature 403, 853–858 (2000).
Boyero, L. et al. Global patterns of stream detritivore distribution: implications for biodiversity loss in changing climates. Glob. Ecol. Biogeogr. 21, 134–141 (2012).
Oksanen, J. et al. Vegan: Community Ecology Package. R package version 2.0-4 (R Foundation for Statistical Computing, 2016).
Reese, A. T. et al. Urban stress is associated with variation in microbial species composition—but not richness—in Manhattan. ISME J. 10, 751–760 (2016).
Fierer, N., Strickland, M. S., Liptzin, D., Bradford, M. A. & Cleveland, C. C. Global patterns in belowground communities. Ecol. Lett. 12, 1238–1249 (2009).
Rousk, J. et al. Soil bacterial and fungal communities across a pH gradient in an arable soil. ISME J. 4, 1340–1351 (2010).
Bru, D. et al. Determinants of the distribution of nitrogen-cycling microbial communities at the landscape scale. ISME J. 5, 532–542 (2011).
R Core Team A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2015).
Anderson, M. J. Distance-based tests for homogeneity of multivariate dispersions. Biometrics 62, 245–253 (2006).
Karpati, A. S., Handel, S. N., Dighton, J. & Horton, T. R. Quercus rubra-associated ectomycorrhizal fungal communities of disturbed urban sites and mature forests. Mycorrhiza 21, 537–547 (2011).
Jumpponen, A. & Egerton-Warburton, L. M. in The Fungal Community: Its Organization and Role in the Ecosystem 3rd edn (eds Dighton, J., White, J. F. & Oudemans, P.) Ch. 7, 130–164 (Mycology Series Vol. 23, 2005).
Tedersoo, L. et al. Towards global patterns in the diversity and community structure of ectomycorrhizal fungi. Mol. Ecol. 21, 4160–4170 (2012).
Barberán, A. et al. Continental-scale distributions of dust-associated bacteria and fungi. Proc. Natl Acad. Sci. USA 112, 5756–5761 (2015).
Taddei, F. et al. Role of mutator alleles in adaptive evolution. Nature 387, 700–702 (1997).
Tenaillon, O., Toupance, B., Nagard, H. L., Taddei, F. & Godelle, B. Mutators, population size, adaptive landscape and the adaptation of asexual populations of bacteria. Genetics 152, 485–493 (1999).
Ochman, H., Lawrence, J. G. & Groisman, E. A. Lateral gene transfer and the nature of bacterial innovation. Nature 405, 299–304 (2000).
Verhamme, D. T., Prosser, J. I. & Nicol, G. W. Ammonia concentration determines differential growth of ammonia-oxidising archaea and bacteria in soil microcosms. ISME J. 5, 1067–1071 (2011).
Nicol, G. W., Leininger, S., Schleper, C. & Prosser, J. I. The influence of soil pH on the diversity, abundance and transcriptional activity of ammonia oxidizing archaea and bacteria. Environ. Microbiol. 10, 2966–2978 (2008).
Hanson, C. A., Fuhrman, J. A., Horner-Devine, M. C. & Martiny, J. B. H. Beyond biogeographic patterns: processes shaping the microbial landscape. Nat. Rev. Microbiol. 10, 497–506 (2012).
Nemergut, D. R. et al. Patterns and processes of microbial community assembly. Microbiol. Mol. Biol. Rev. 77, 342–356 (2013).
Ramirez, K. S. et al. Biogeographic patterns in below-ground diversity in New York City’s Central Park are similar to those observed globally. Proc. R. Soc. B 281, 20141988 (2014).
Högberg, M. N., Yarwood, S. A. & Myrold, D. D. Fungal but not bacterial soil communities recover after termination of decadal nitrogen additions to boreal forest. Soil Biol. Biochem. 72, 35–43 (2014).
Newbold, T. et al. Global effects of land-use on local terrestrial biodiversity. Nature 520, 45–50 (2015).
Averill, C. & Hawkes, C. V. Ectomycorrhizal fungi slow soil carbon cycling. Ecol. Lett. 19, 937–947 (2016).
Abarenkov, K. et al. The UNITE database for molecular identification of fungi – recent updates and future perspectives. New Phytol. 186, 281–285 (2010).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–1624 (2012).
Fierer, N., Jackson, J. A., Vilgalys, R. & Jackson, R. B. Assessment of soil microbial community structure by use of taxon-specific quantitative PCR assays. Appl. Environ. Microbiol. 71, 4117–4120 (2005).
Hargreaves, S. K., Roberto, A. A. & Hofmockel, K. S. Reaction- and sample-specific inhibition affect standardization of Q-PCR assays of soil bacterial communities. Soil Biol. Biochem. 59, 89–97 (2013).
Edgar, R. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Wang, Q., Garrity, G., Tiedje, J. & Cole, J. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73, 5261–5267 (2007).
DeSantis, T. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072 (2006).
Aronesty, E. ea-utils (2011); https://expressionanalysis.github.io/ea-utils/
McCune, B. & Mefford, M. PC-ORD: Multivariate Analysis of Ecological Data (MjM Software, 2011).
Dufrêne, M. & Legendre, P. Species assemblages and indicator species: the need for a flexible asymmetrical approach. Ecol. Monogr. 67, 345–366 (1997).
Gastrwirth, J. et al. lawstat v. 3.0 (2015); https://cran.r-project.org/web/packages/lawstat/index.html
Nguyen, N. H. et al. FUNGuild: an open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol. 20, 241–248 (2016).
Dlott, G., Maul, J. E., Buyer, J. & Yarwood, S. Microbial rRNA:rDNA gene ratios may be unexpectedly low due to extracellular DNA preservation in soils. J. Microbiol. Methods 115, 112–120 (2015).
Acknowledgements
This research was funded through NSF ACI 1244820; soil chemical analysis was funded through SZIE-ÁOTK: KK-UK-12007. We acknowledge G. Dlott for technical support in QIIME, and R. E. Draskovits, S. Molnar, E. Powell, M. Bernard, Z. Toth and S. Mishra assisted with field sampling.
Author information
Authors and Affiliations
Contributions
D.J.E.S. constructed the DNA and sequencing libraries, and conducted Q-PCR, bioinformatics processing and statistical analyses. K.S. is the PI of the grant, designed the study and selected the sites in Baltimore. R.V.P., H.S., D.J.K., E.H., S.C. and I.Y. designed the study, selected the sites and participated in soil sampling. M.D. participated in soil sampling and provided nutrient data on soils; S.A.Y. designed the study, and oversaw all of the lab work, bioinformatics and data analysis. All authors discussed results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1–5, Supplementary Tables 1–8, Supplementary References (PDF 1821 kb)
Rights and permissions
About this article
Cite this article
Epp Schmidt, D., Pouyat, R., Szlavecz, K. et al. Urbanization erodes ectomycorrhizal fungal diversity and may cause microbial communities to converge. Nat Ecol Evol 1, 0123 (2017). https://doi.org/10.1038/s41559-017-0123
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41559-017-0123
This article is cited by
-
Harnessing soil biodiversity to promote human health in cities
npj Urban Sustainability (2023)
-
Biochar, but not soil microbial additives, increase the resilience of urban plant species to low water availability
Urban Ecosystems (2023)
-
Quantitative Amplicon Sequencing Is Necessary to Identify Differential Taxa and Correlated Taxa Where Population Sizes Differ
Microbial Ecology (2023)
-
Soil microbiomes in lawns reveal land-use legacy impacts on urban landscapes
Oecologia (2023)
-
Rewilding in Miniature: Suburban Meadows Can Improve Soil Microbial Biodiversity and Soil Health
Microbial Ecology (2023)