Abstract
Unravelling the relationships between network complexity and stability under changing climate is a challenging topic in theoretical ecology that remains understudied in the field of microbial ecology. Here, we examined the effects of long-term experimental warming on the complexity and stability of molecular ecological networks in grassland soil microbial communities. Warming significantly increased network complexity, including network size, connectivity, connectance, average clustering coefficient, relative modularity and number of keystone species, as compared with the ambient control. Molecular ecological networks under warming became significantly more robust, with network stability strongly correlated with network complexity, supporting the central ecological belief that complexity begets stability. Furthermore, warming significantly strengthened the relationships of network structure to community functional potentials and key ecosystem functioning. These results indicate that preserving microbial ‘interactions’ is critical for ecosystem management and for projecting ecological consequences of future climate warming.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data accessibility
16S rRNA gene sequences were deposited to the National Center for Biotechnology Information (NCBI) under the project accession number PRJNA331185. The OTU table and OTU representative sequences, soil physical and chemical attributes, and plant biomass and richness are downloadable online at http://www.ou.edu/ieg/publications/datasets. GeoChip signal intensity data can be accessed through the URL (https://www.ou.edu/ieg/publications/datasets). Source data are provided with this paper.
Code availability
The R scripts and Python 3 scripts are publicly available on GitHub at https://github.com/Mengting-Maggie-Yuan/warming-network-complexity-stability with the identifier https://doi.org/10.5281/zenodo.4383469.
References
Montoya, J. M., Pimm, S. L. & Solé, R. V. Ecological networks and their fragility. Nature 442, 259–264 (2006).
Faust, K. & Raes, J. Microbial interactions: from networks to models. Nat. Rev. Microbiol. 10, 538–550 (2012).
Pržulj, N. & Malod-Dognin, N. Network analytics in the age of big data. Science 353, 123–124 (2016).
Berry, D. & Widder, S. Deciphering microbial interactions and detecting keystone species with co-occurrence networks. Front. Microbiol. 5, 219 (2014).
Okuyama, T. & Holland, J. N. Network structural properties mediate the stability of mutualistic communities. Ecol. Lett. 11, 208–216 (2008).
Landi, P., Minoarivelo, H. O., Brännström, Å., Hui, C. & Dieckmann, U. Complexity and stability of ecological networks: a review of the theory. Popul. Ecol. 60, 319–345 (2018).
Hillebrand, H. et al. Decomposing multiple dimensions of stability in global change experiments. Ecol. Lett. 21, 21–30 (2018).
Toju, H. et al. Species-rich networks and eco-evolutionary synthesis at the metacommunity level. Nat. Ecol. Evol. 1, 0024 (2017).
Montesinos-Navarro, A., Hiraldo, F., Tella, J. L. & Blanco, G. Network structure embracing mutualism–antagonism continuums increases community robustness. Nat. Ecol. Evol. 1, 1661–1669 (2017).
Ullah, H., Nagelkerken, I., Goldenberg, S. U. & Fordham, D. A. Climate change could drive marine food web collapse through altered trophic flows and cyanobacterial proliferation. PLoS Biol. 16, e2003446 (2018).
Dunne, J. A., Williams, R. J. & Martinez, N. D. Food-web structure and network theory: the role of connectance and size. Proc. Natl Acad. Sci. USA 99, 12917–12922 (2002).
Thébault, E. & Fontaine, C. Stability of ecological communities and the architecture of mutualistic and trophic networks. Science 329, 853–856 (2010).
García-Palacios, P., Gross, N., Gaitán, J. & Maestre, F. T. Climate mediates the biodiversity–ecosystem stability relationship globally. Proc. Natl Acad. Sci. USA 115, 8400–8405 (2018).
IPCC Climate Change 2013: The Physical Science Basis (eds Stocker, T. F. et al.) (Cambridge Univ. Press, 2013).
Xue, K. et al. Tundra soil carbon is vulnerable to rapid microbial decomposition under climate warming. Nat. Clim. Change 6, 595–600 (2016).
Brown, J. H., Gillooly, J. F., Allen, A. P., Savage, V. M. & West, G. B. Toward a metabolic theory of ecology. Ecology 85, 1771–1789 (2004).
Guo, X. et al. Climate warming leads to divergent succession of grassland microbial communities. Nat. Clim. Change 8, 813–818 (2018).
Xu, X., Sherry, R. A., Niu, S., Li, D. & Luo, Y. Net primary productivity and rain-use efficiency as affected by warming, altered precipitation, and clipping in a mixed-grass prairie. Glob. Change Biol. 19, 2753–2764 (2013).
Guo, X. et al. Climate warming accelerates temporal scaling of grassland soil microbial biodiversity. Nat. Ecol. Evol. 3, 612–619 (2019).
Zhou, J. et al. Functional molecular ecological networks. mBio 1, e00169–10 (2010).
Barabási, A.-L. & Oltvai, Z. N. Network biology: understanding the cell’s functional organization. Nat. Rev. Genet. 5, 101–113 (2004).
D’Amen, M., Mod, H. K., Gotelli, N. J. & Guisan, A. Disentangling biotic interactions, environmental filters, and dispersal limitation as drivers of species co-occurrence. Ecography 41, 1233–1244 (2018).
Barner, A. K., Coblentz, K. E., Hacker, S. D. & Menge, B. A. Fundamental contradictions among observational and experimental estimates of non-trophic species interactions. Ecology 99, 557–566 (2018).
Goberna, M. et al. Incorporating phylogenetic metrics to microbial co-occurrence networks based on amplicon sequences to discern community assembly processes. Mol. Ecol. Resour. 19, 1552–1564 (2019).
Carr, A., Diener, C., Baliga, N. S. & Gibbons, S. M. Use and abuse of correlation analyses in microbial ecology. ISME J. 13, 2647–2655 (2019).
Morton, J. T. et al. Establishing microbial composition measurement standards with reference frames. Nat. Commun. 10, 2719 (2019).
Fuhrman, J. A. Microbial community structure and its functional implications. Nature 459, 193–199 (2009).
Herren, C. M. & McMahon, K. D. Cohesion: a method for quantifying the connectivity of microbial communities. ISME J. 11, 2426–2438 (2017).
Zhou, J., Deng, Y., Luo, F., He, Z. & Yang, Y. Phylogenetic molecular ecological network of soil microbial communities in response to elevated CO2. mBio 2, e00122–11 (2011).
Banerjee, S., Schlaeppi, K. & van der Heijden, M. G. A. Keystone taxa as drivers of microbiome structure and functioning. Nat. Rev. Microbiol. 16, 567–576 (2018).
Zelikova, T. J. et al. Long-term exposure to elevated CO2 enhances plant community stability by suppressing dominant plant species in a mixed-grass prairie. Proc. Natl Acad. Sci. USA 111, 15456–15461 (2014).
Douglas, G. M. et al. PICRUSt2 for prediction of metagenome functions. Nat. Biotechnol. 38, 685–688 (2020).
MacArthur, R. Fluctuations of animal populations and a measure of community stability. Ecology 36, 533–536 (1955).
May, R. M. Stability and Complexity in Model Ecosystems (Princeton Univ. Press, 2019).
Guo, X. et al. Gene-informed decomposition model predicts lower soil carbon loss due to persistent microbial adaptation to warming. Nat. Commun. 11, 4897 (2020).
Melillo, J. M. et al. Long-term pattern and magnitude of soil carbon feedback to the climate system in a warming world. Science 358, 101–105 (2017).
Zhou, J. et al. Microbial mediation of carbon-cycle feedbacks to climate warming. Nat. Clim. Change 2, 106–110 (2012).
Galiana, N. et al. The spatial scaling of species interaction networks. Nat. Ecol. Evol. 2, 782–790 (2018).
Bastolla, U. et al. The architecture of mutualistic networks minimizes competition and increases biodiversity. Nature 458, 1018–1020 (2009).
Cardinale, B. J. et al. Biodiversity loss and its impact on humanity. Nature 486, 59–67 (2012).
Li, D., Zhou, X., Wu, L., Zhou, J. & Luo, Y. Contrasting responses of heterotrophic and autotrophic respiration to experimental warming in a winter annual-dominated prairie. Glob. Change Biol. 19, 3553–3564 (2013).
Treves, D. S., Xia, B., Zhou, J. & Tiedje, J. M. A two-species test of the hypothesis that spatial isolation influences microbial diversity in soil. Microb. Ecol. 45, 20–28 (2003).
Zhou, J., Xia, B., Huang, H., Palumbo, A. V. & Tiedje, J. M. Microbial diversity and heterogeneity in sandy subsurface soils. Appl. Environ. Microbiol. 70, 1723–1734 (2004).
Zhou, J. et al. Spatial and resource factors influencing high microbial diversity in soil. Appl. Environ. Microbiol. 68, 326–334 (2002).
O’Brien, S. L. et al. Spatial scale drives patterns in soil bacterial diversity. Environ. Microbiol. 18, 2039–2051 (2016).
Penton, C. R., Gupta, V. V. S. R., Yu, J. & Tiedje, J. M. Size matters: assessing optimum soil sample size for fungal and bacterial community structure analyses using high throughput sequencing of rRNA gene amplicons. Front. Microbiol. 7, 824 (2016).
Zhou, J., Bruns, M. A. & Tiedje, J. M. DNA recovery from soils of diverse composition. Appl. Environ. Microbiol. 62, 316–322 (1996).
Hurt, R. A. et al. Simultaneous recovery of RNA and DNA from soils and sediments. Appl. Environ. Microbiol. 67, 4495–4503 (2001).
Peiffer, J. A. et al. Diversity and heritability of the maize rhizosphere microbiome under field conditions. Proc. Natl Acad. Sci. USA 110, 6548–6553 (2013).
Wu, L. et al. Phasing amplicon sequencing on Illumina Miseq for robust environmental microbial community analysis. BMC Microbiol. 15, 125 (2015).
Wen, C. et al. Evaluation of the reproducibility of amplicon sequencing with Illumina MiSeq platform. PLoS ONE 12, e0176716 (2017).
Zhou, J. et al. High-throughput metagenomic technologies for complex microbial community analysis: open and closed formats. mBio 6, e02288–14 (2015).
Zhou, J. et al. Reproducibility and quantitation of amplicon sequencing-based detection. ISME J. 5, 1303–1313 (2011).
Luo, F. et al. Constructing gene co-expression networks and predicting functions of unknown genes by random matrix theory. BMC Bioinformatics 8, 299 (2007).
Luo, F., Zhong, J., Yang, Y., Scheuermann, R. H. & Zhou, J. Application of random matrix theory to biological networks. Phys. Lett. A 357, 420–423 (2006).
Deng, Y. et al. Molecular ecological network analyses. BMC Bioinformatics 13, 113 (2012).
Shi, S. et al. The interconnected rhizosphere: high network complexity dominates rhizosphere assemblages. Ecol. Lett. 19, 926–936 (2016).
Mehta, M. L. Random Matrices 2nd edn (Elsevier, 2004).
Plerou, V., Gopikrishnan, P., Rosenow, B., Amaral, L. A. N. & Stanley, H. E. Universal and non-universal properties of cross-correlations in financial time series. Phys. Rev. Lett. 83, 1471–1474 (1999).
Aitchison, J. The statistical analysis of compositional data. J. R. Stat. Soc. B 44, 139–160 (1982).
Gloor, G. B., Macklaim, J. M., Pawlowsky-Glahn, V. & Egozcue, J. J. Microbiome datasets are compositional: and this is not optional. Front. Microbiol. 8, 2224 (2017).
Pawlowsky-Glahn, V. & Egozcue, J. J. Compositional data and their analysis: an introduction. Geol. Soc. Spec. Publ. 264, 1–10 (2006).
Friedman, J. & Alm, E. J. Inferring correlation networks from genomic survey data. PLoS Comput. Biol. 8, e1002687 (2012).
Watts, S. C., Ritchie, S. C., Inouye, M. & Holt, K. E. FastSpar: rapid and scalable correlation estimation for compositional data. Bioinformatics 35, 1064–1066 (2019).
Weiss, S. et al. Correlation detection strategies in microbial data sets vary widely in sensitivity and precision. ISME J. 10, 1669–1681 (2016).
R: a language and environment for statistical computing (R Foundation for Statistical Computing, 2019).
Goslee, S. C. & Urban, D. L. The ecodist package for dissimilarity-based analysis of ecological data. J. Stat. Softw. 22, 1–19 (2007).
Oksanen, J. et al. vegan: Community Ecology Package. Version 2.5-6 (2019).
Lima-Mendez, G. et al. Determinants of community structure in the global plankton interactome. Science 348, 1262073 (2015).
Yuan, M.M. et al. Mengting-Maggie-Yuan/warming-network-complexity-stability: warming-network-complexity-stability-v1.0. Version 1.0 (Zenodo, 2021); https://doi.org/10.5281/zenodo.4383469
He, Z. et al. GeoChip 3.0 as a high-throughput tool for analyzing microbial community composition, structure and functional activity. ISME J. 4, 1167–1179 (2010).
He, Z. et al. GeoChip: a comprehensive microarray for investigating biogeochemical, ecological and environmental processes. ISME J. 1, 67–77 (2007).
Ning, D., Deng, Y., Tiedje, J. M. & Zhou, J. A general framework for quantitatively assessing ecological stochasticity. Proc. Natl Acad. Sci. USA 116, 16892–16898 (2019).
Zhou, J. & Ning, D. Stochastic community assembly: does it matter in microbial ecology? Microbiol. Mol. Biol. Rev. 81, e00002–e00017 (2017).
Csárdi, G. & Nepusz, T. The igraph software package for complex network research. InterJ. Complex Syst. 1695, 1–9 (2006).
Maslov, S. & Sneppen, K. Specificity and stability in topology of protein networks. Science 296, 910–913 (2002).
Almeida‐Neto, M., Guimarães, P., Guimarães, P. R., Loyola, R. D. & Ulrich, W. A consistent metric for nestedness analysis in ecological systems: reconciling concept and measurement. Oikos 117, 1227–1239 (2008).
Guimerà, R. & Nunes Amaral, L. A. Functional cartography of complex metabolic networks. Nature 433, 895–900 (2005).
Olesen, J. M., Bascompte, J., Dupont, Y. L. & Jordano, P. The modularity of pollination networks. Proc. Natl Acad. Sci. USA 104, 19891–19896 (2007).
Banerjee, S., Schlaeppi, K. & van der Heijden, M. G. A. Reply to ‘can we predict microbial keystones?’. Nat. Rev. Microbiol. 17, 194 (2019).
Röttjers, L. & Faust, K. Can we predict keystones? Nat. Rev. Microbiol. 17, 193 (2019).
Langfelder, P. & Horvath, S. Eigengene networks for studying the relationships between co-expression modules. BMC Syst. Biol. 1, 54 (2007).
Hautier, Y. et al. Eutrophication weakens stabilizing effects of diversity in natural grasslands. Nature 508, 521–525 (2014).
Hui, C., McGeoch, M. A., Harrison, A. E. S. & Bronstein, E. J. L. Zeta diversity as a concept and metric that unifies incidence-based biodiversity patterns. Am. Nat. 184, 684–694 (2014).
Shi, Z. et al. Functional gene array-based ultrasensitive and quantitative detection of microbial populations in complex communities. mSystems 4, e00296–19 (2019).
Sun, S., Jones, R. B. & Fodor, A. A. Inference-based accuracy of metagenome prediction tools varies across sample types and functional categories. Microbiome 8, 46 (2020).
Acknowledgements
We thank numerous former and current members in the Institute for Environmental Genomics for their help in maintaining the long-term field experiment. This work was supported by the US Department of Energy, Office of Science, Genomic Science Program under award numbers DE-SC0004601 and DE-SC0010715, and the Office of the Vice President for Research at the University of Oklahoma. X.G. and X.Z. were generously supported by China Scholarship Council (CSC) to visit the University of Oklahoma. The statistical analyses performed by X.G. were also supported by the China Postdoctoral Science Foundation (2018M641327 and 2019T120101).
Author information
Authors and Affiliations
Contributions
All authors contributed intellectual input and assistance to this study. The original concepts were conceived by J.Z. and J.M.T. Field management was carried out by M.M.Y., X.G., Linwei W., Y.Z., Z.S., D.N. and Liyou W. Sampling collection, soil chemical and microbial characterization were carried out by M.M.Y., X.G. and X.Z. Data analyses were done by M.Y., X.G., Linwei W., Z.S. and N.X. with the assistance provided by D.N. and J.Z. All data analysis and integration were guided by J.Z. The manuscript was prepared by J.Z, M.M.Y., X.G., Linwei W. and Y.Z. with substantial input from J.M.T. and Y.Y. Considering their contributions in terms of site management, data collection, analyses and/or integration, M.Y., X.G., Linwei W. and Y.Z. were listed as co-first authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Climate Change thanks Johannes Bjork, Dongmei Xue and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
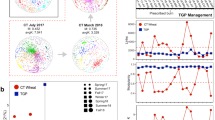
Extended Data Fig. 1 Succession and environmental drivers of the networked community structure.
a, Detrended correspondence analysis (DCA) of the structure of networked communities. The community structure is significantly different by both treatment and year. b, Canonical correspondence analysis (CCA) of the links between networked community structure and environmental drivers. The ordination plot shows the CCA model with each networked microbial community and constraining variables, including spatial distance (Distance), soil temperature (Temp.), soil moisture (Moisture), soil pH (pH), soil total N (TN), soil nitrate (NO3−N) and ammonia (NH4-N) contents, plant biomass and richness. The model is significant with p = 0.001 tested by ANOVA. c, Variation partitioning analysis (VPA) separating the variation of community structure explained by the CCA model. Soil category includes soil temperature (Temp.), soil moisture (Moisture), soil pH (pH), soil total N (TN), soil nitrate (NO3-N) and ammonia (NH4-N) contents; plant category includes plant biomass and richness.
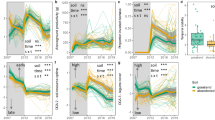
Extended Data Fig. 2 Cohesion of bacterial communities and its relationships with network complexity and stability indices.
a, Changes in positive cohesion of bactera community over time. b, Changes in negative cohesion of the commuity over time. In (a) and (b), filled red circles with solid line represent communities under warming, and open blue circles with dashed line represent communities under control condition. Each error bar corresponds to the standard deviation of cohesion in 24 plots. The slopes (b from Y=a+bX), adjusted r2 and p values of the linear model fittings are shown. c, Pearson correlations of cohesion with various network complexity and stability indices under warming (framed in red) or control (framed in blue) condition. The cells highlighted in red indicate significant positive correlations (p ≤ 0.05) and those in blue indicate significant negative correlations. Numbers inside of the cells are correlation coefficients. Correlations with p > 0.05 are in gray.
Extended Data Fig. 3 Modules preserved across time and treatments.
a, Large modules (that is, those with ≥ 5 nodes) shown in circular layout for the 11 networks. Colors of nodes indicate major taxa. Red links indicate positive correlations between nodes. Blue links indicate negative correlations between nodes. The bar underneath each network shows the proportions of positive and negative links. The label nearby each module represents its ID. b, Preserved module pairs highlighted and connected in the same module layout as (a). Modules are in the same color if they are in the same module cluster (that is, a cluster of modules consisting all the directly paired and indirectly linked modules). Note that two clusters of modules (the red and the blue clusters) were preserved over time consistently between Year 1 (2010) and Year 5 (2014). More details are in Supplementary text C.
Extended Data Fig. 4 Keystone taxa and their relative abundances in bacterial communities.
a,b, Putative keystone taxa identified based on the node topological roles in networks under control (a) and warming (b). Each symbol represents a node in one of the networks. A node was identified as a module hub if its Zi ≥ 2.5, as a connector if its Pi ≥ 0.62, and as a network hub if it had Zi ≥ 2.5 and Pi ≥ 0.62. Detailed taxonomic information for module hubs, connectors and network hubs is listed in Supplementary Table 6. c–f, The relative abundances of module hubs (c, d) and connectors (e, f) in the networks under control (c, e) and warming (d, f). The relative abundance of an OTU was estimated as the percentage of its number of sequences in the total number of squences detected for the community. g, A maximum likelihood phylogenetic tree of keystone nodes in all networks. Green, red, and blue dots represent the taxa of keystone nodes that occurred in both warming and control networks, only under warming, and only in control, respectively. Branches are colored based on bacterial phyla identified using RDP classifier.
Extended Data Fig. 5 Temporal variations (that is, constancy) of network nodes and links.
a, Network node constancy. Each box shows the constancy distribution of all the nodes, averaged across experimental plots, present in the networks under warming (n = 603) or control (n = 601). Mann-Whitney U test results are shown. b, The number of overlapping nodes under warming and control among different numbers of networks (that is, orders). For example, for order=2, the overlapping nodes were between any two pairs of networks; for order=3, they were among any three networks. The nodes consistently present in all the time points are listed in Supplementary Table 3. c, The number of overlapping nodes among multiple networks from different gap times. The datapoints include orders 2 to 6. Linear regression results are shown. d, Unweighted network link constancy. Each box shows the constancy distribution of the links in the networks under warming (n = 4,661) or control (n = 3,526). Mann-Whitney U test results are shown here.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10, text and references.
Supplementary Data 1
Supplementary Tables 1–9.
Source data
Source Data Fig. 1
Source data for overall network.
Source Data Fig. 2
Source data for correlations.
Source Data Fig. 3
Source data for stability.
Source Data Fig. 4
Source data for ecosystem processes and functioning.
Source Data Extended Data Fig. 1
Statistical source data for DCA, CCA and VPA.
Source Data Extended Data Fig. 2
Statistical source data for cohesion.
Source Data Extended Data Fig. 3
Source data for module preservation.
Source Data Extended Data Fig. 4
Source data for node types.
Source Data Extended Data Fig. 5
Source data for node overlap and constancy.
Rights and permissions
About this article
Cite this article
Yuan, M.M., Guo, X., Wu, L. et al. Climate warming enhances microbial network complexity and stability. Nat. Clim. Chang. 11, 343–348 (2021). https://doi.org/10.1038/s41558-021-00989-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41558-021-00989-9
This article is cited by
-
Host genotype-specific rhizosphere fungus enhances drought resistance in wheat
Microbiome (2024)
-
New insights into bioaugmented removal of sulfamethoxazole in sediment microcosms: degradation efficiency, ecological risk and microbial mechanisms
Microbiome (2024)
-
Metagenome-assembled genome reveals species and functional composition of Jianghan chicken gut microbiota and isolation of Pediococcus acidilactic with probiotic properties
Microbiome (2024)
-
Comparative analysis of microbial composition and functional characteristics in dental plaque and saliva of oral cancer patients
BMC Oral Health (2024)
-
Effect of stress urinary incontinence on vaginal microbial communities
BMC Microbiology (2024)