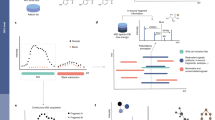
Abstract
Although metals are essential for the molecular machineries of life, systematic methods for discovering metal–small molecule complexes from biological samples are limited. Here, we describe a two-step native electrospray ionization–mass spectrometry method, in which post-column pH adjustment and metal infusion are combined with ion identity molecular networking, a rule-based data analysis workflow. This method enabled the identification of metal-binding compounds in complex samples based on defined mass (m/z) offsets of ion species with the same chromatographic profiles. As this native electrospray metabolomics approach is suited to the use of any liquid chromatography–mass spectrometry system to explore the binding of any metal, this method has the potential to become an essential strategy for elucidating metal-binding molecules in biology.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the paper and its Supplementary Information. All MS .raw and centroided .mzXML or .mzML files are publicly available in the Mass spectrometry Interactive Virtual Environment (MassIVE) under massive.ucsd.edu with project identifiers MSV000084237, MSV000085669, MSV000085206 (standards), MSV000084289 (cheese siderophores), MSV000082999 and MSV000084030 (fungal siderophores), MSV000083387 (E.coli Nissle siderophores), MSV000085554 (California current ecosystem phytoplankton bloom samples) and MSV000086744 (chelomics versus native metabolomics comparison). IIMN can be accessed through gnps.ucsd.edu under the following direct links: https://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=79d0f380b4814ff9a720836c5570036f, https://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=ad4b2665dfb744d09a9d2445f1213720, https://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=5459d22126e843a3a1449f8362cd267f, http://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=1c3e79f0ab984386bd468e2d163281e0, https://gnps.ucsd.edu/ProteoSAFe/index.jsp?task=196a29a94c2f4c788e204b9934ea4d9b, http://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=256ba734f4334c1c90f65ffbd9141d0e, https://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=042939579da64e029a8b5caef8f7f2a8.
Code availability
The modified version of MZmine 2 (2.37, corr.17.7)41,42,43 can be found at https://mzmine.github.io/iin_fbmn. GNPS44,45 can be accessed at https://gnps.ucsd.edu/ProteoSAFe/static/gnps-splash.jsp. Cytoscape63 version 3.7.1 can be accessed at https://cytoscape.org/.
References
Holden, V. I. & Bachman, M. A. Diverging roles of bacterial siderophores during infection. Metallomics 7, 986–995 (2015).
Sandy, M. & Butler, A. Microbial iron acquisition: marine and terrestrial siderophores. Chem. Rev. 109, 4580–4595 (2009).
Raymond, K. N., Allred, B. E. & Sia, A. K. Coordination chemistry of microbial iron transport. Acc. Chem. Res. 48, 2496–2505 (2015).
Vraspir, J. M. & Butler, A. Chemistry of marine ligands and siderophores. Annu. Rev. Mar. Sci 1, 43–63 (2009).
Kenney, G. E. & Rosenzweig, A. C. Chalkophores. Annu. Rev. Biochem. 87, 645–676 (2018).
Wang, L. et al. Diisonitrile natural product SF2768 functions as a chalkophore that mediates copper acquisition in Streptomyces thioluteus. ACS Chem. Biol. 12, 3067–3075 (2017).
Johnstone, T. C. & Nolan, E. M. Beyond iron: non-classical biological functions of bacterial siderophores. Dalton Trans. 44, 6320–6339 (2015).
Łobodaa, D. & Rowińska-Żyrek, M. Zinc binding sites in Pra1, a zincophore from Candida albicans. Dalton Trans. 46, 13695–13703 (2017).
Wilson, D., Citiulo, F. & Hube, B. Zinc exploitation by pathogenic fungi. PLoS Pathog. 8, e1003034 (2012).
Capdevila, D. A., Wang, J. & Giedroc, D. P. Bacterial strategies to maintain zinc metallostasis at the host–pathogen interface. J. Biol. Chem. 291, 20858–20868 (2016).
Hover, B. M. et al. Culture-independent discovery of the malacidins as calcium-dependent antibiotics with activity against multidrug-resistant Gram-positive pathogens. Nat. Microbiol. 3, 415–422 (2018).
Wood, T. M. & Martin, N. I. The calcium-dependent lipopeptide antibiotics: structure, mechanism and medicinal chemistry. Med. Chem. Commun. 10, 634–646 (2019).
von Eckardstein, L. et al. Total synthesis and biological assessment of novel albicidins discovered by mass spectrometric networking. Chem. Eur. J. 23, 15316–15321 (2017).
Petras, D. et al. High-resolution liquid chromatography tandem mass spectrometry enables large scale molecular characterization of dissolved organic matter. Front. Mar. Sci. https://doi.org/10.3389/fmars.2017.00405 (2017).
Gauglitz, J. M. Reference data based insights expand understanding of human metabolomes. Preprint at bioRxiv https://doi.org/10.1101/2020.07.08.194159 (2020).
Medema, M. H. & Fischbach, M. A. Computational approaches to natural product discovery. Nat. Chem. Biol. 11, 639–648 (2015).
Bachmann, B. O., Van Lanene, S. G. & Baltz, R. H. Microbial genome mining for accelerated natural products discovery: is a renaissance in the making? J. Ind. Microbiol. Biotechnol. 41, 175–184 (2014).
Kasampalidis, I. N., Pitas, I. & Lyroudia, K. Conservation of metal-coordinating residues. Proteins Struct. Funct. Bioinf. 68, 123–130 (2007).
Cvetkovic, A. et al. Microbial metalloproteomes are largely uncharacterized. Nature 466, 779–782 (2010).
Ackerman, C. M., Lee, S. & Chang, C. J. Analytical methods for imaging metals in biology: from transition metal metabolism to transition metal signaling. Anal. Chem. 89, 22–41 (2017).
Piper, K. G. & Higgins, G. Estimation of trace metals in biological material by atomic absorption spectrophotometry. Proc. Ass. Clin. Biochem. 4, 190–197 (1967).
Aschner, M. et al. Imaging metals in Caenorhabditis elegans. Metallomics 9, 357–364 (2017).
Mawji, E. et al. Production of siderophore type chelates in Atlantic Ocean waters enriched with different carbon and nitrogen sources. Mar. Chem. 124, 90–99 (2011).
Mawji, E. et al. Hydroxamate siderophores: occurrence and importance in the Atlantic Ocean. Environ. Sci. Technol. 42, 8675–8680 (2008).
Gledhill, M. et al. Production of siderophore type chelates by mixed bacterioplankton populations in nutrient enriched seawater incubations. Mar. Chem. 88, 75–83 (2004).
Velasquez, I. et al. Detection of hydroxamate siderophores in coastal and sub-Antarctic waters off the South Eastern Coast of New Zealand. Mar. Chem. 126, 97–107 (2011).
Boiteau, R. M. & Repeta, D. J. An extended siderophore suite from Synechococcus sp. PCC 7002 revealed by LC-ICPMS-ESIMS. Metallomics 7, 877–884 (2015).
Boiteau, R. M. et al. Siderophore-based microbial adaptations to iron scarcity across the eastern Pacific Ocean. Proc. Natl Acad. Sci. USA 113, 14237–14242 (2016).
Pluháček, T. et al. Characterization of microbial siderophores by mass spectrometry. Mass Spectrom. Rev. 35, 35–47 (2016).
Walker, L. R. et al. Unambiguous identification and discovery of bacterial siderophores by direct injection 21 Tesla Fourier transform ion cyclotron resonance mass spectrometry. Metallomics 9, 82–92 (2017).
Baars, O., Zhang, X., Morel, F. M. & Seyedsayamdost, M. R. The siderophore metabolome of Azotobacter vinelandii. Appl. Environ. Microbiol. 82, 27–39 (2015).
Baars, O., Morel, F. M. M. & Perlman, D. H. ChelomEx: isotope-assisted discovery of metal chelates in complex media using high-resolution LC-MS. Anal. Chem. 86, 11298–11305 (2014).
Baars, O. et al. Crochelins: siderophores with an unprecedented iron‐chelating moiety from the nitrogen‐fixing bacterium Azotobacter chroococcum. Angew. Chem. Int. Ed. 130, 545–550 (2018).
Garcia-Sartal, C. et al. Two-dimensional HPLC coupled to ICP-MS and electrospray ionisation (ESI)-MS/MS for investigating the bioavailability in vitro of arsenic species from edible seaweed. Anal. Bioanal. Chem. 402, 3359–3369 (2012).
Leney, A. C. & Heck, A. J. R. Native mass spectrometry: what is in the name? J. Am. Soc. Mass. Spectrom. 28, 5–13 (2017).
Heck, A. J. R. Native mass spectrometry: a bridge between interactomics and structural biology. Nat. Methods 5, 927–933 (2008).
Schmid, R. et al. Ion identity molecular networking in the GNPS environment. Nat. Commun. 12, 3832 (2021).
Ross, A. R. S., Ikonomou, M. G. & Orians, K. J. Electrospray ionization of alkali and alkaline earth metal species. Electrochemical oxidation and pH effects. J. Mass Spectrom. 35, 981–989 (2000).
Lopes, N. P., Stark, C. B. W., Hong, H., Gates, P. J. & Stauntona, J. A study of the effect of pH, solvent system, cone potential and the addition of crown ethers on the formation of the monensin protonated parent ion in electrospray mass spectrometry. Analyst 126, 1630–1632 (2001).
Waska, H., Koschinsky, A. & Dittmar, T. Fe- and Cu-complex formation with artificial ligands investigated by ultra-high resolution Fourier-transform ion cyclotron resonance mass spectrometry (FT-ICR-MS): implications for natural metal–organic complex studies. Front. Mar. Sci. 3, 119 (2016).
Wang, M. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 34, 828–837 (2016).
Aron, A. T. Reproducible molecular networking of untargeted mass spectrometry data using GNPS. Nat. Protoc. 15, 1954–1991 (2020).
Kale, N. S. et al. MetaboLights: an open‐access database repository for metabolomics data. Curr. Protoc. 53, 14.13.11–14.13.18 (2016).
Wang, M. Mass spectrometry searches using MASST. Nat. Biotechnol. 38, 23–26 (2020).
Koh, E. et al. Metal selectivity by the virulence-associated Yersiniabactin metallophore system. Metallomics 7, 1011–1022 (2015).
Bae, W. & Mehra, R. K. Metal-binding characteristics of a phytochelatin analog (Glu–Cys)2Gly. J. Inorg. Biochem. 68, 201–210 (1997).
Raab, A., Feldmann, J. & Meharg, A. A. The nature of arsenic–phytochelatin complexes in Holcus lanatus and Pteris cretica. Plant Physiol. 134, 1113–1122 (2004).
Rellán-Álvarez, R., Abadía, J. & Álvarez-Fernández, A. Formation of metal–nicotianamine complexes as affected by pH, ligand exchange with citrate and metal exchange. A study by electrospray ionization time-of-flight mass spectrometry. Rapid Commun. Mass Spec. 22, 1553–1562 (2008).
Wang, H. & Agnes, G. R. Kinetically labile equilibrium shifts induced by the electrospray process. Anal. Chem. 71, 4166–4172 (1999).
Lee, S. W., Kim, H. S. & Beauchamp, J. L. Salt bridge chemistry applied to gas-phase peptide sequencing: selective fragmentation of sodiated gas-phase peptide ions adjacent to aspartic acid residues. J. Am. Chem. Soc. 120, 3188–3195 (1998).
Crizer, D. M., Xia, Y. & McLuckey, S. A. Metal complexes as reagents for peptide anions. J. Am. Soc. Mass. Spectrom. 20, 1718–1722 (2009).
Hu, P. & Loo, J. A. Gas-phase coordination properties of Zn2+, Cu2+, Ni2+ and Co2+ with histidine-containing peptides. J. Am. Chem. Soc. 117, 11314–11319 (1995).
Bowden, J. A., Albert, C. J., Barnaby, O. S. & Ford, D. A. Analysis of cholesteryl esters and diacylglycerols using lithiated adducts and electrospray ionization-tandem mass spectrometry. Anal. Biochem. 417, 202–210 (2011).
Billo, E. J., Brito, K. K. & Wilkins, R. G. Kinetics of formation and dissociation of metallocarboxypeptidases. Bioinorg. Chem. 8, 461–475 (1978).
Magzoub, M., Padmawar, P., Dix, J. A. & Verkman, A. S. Millisecond association kinetics of K+ with triazacryptand-based K+ indicators measured by fluorescence correlation spectroscopy. J. Phys. Chem. B 110, 21216–21221 (2006).
Chock, P. B. Relaxation study of complex formation between monovalent cations and cyclic polyethers. Proc. Natl Acad. Sci. USA 69, 1939–1942 (1972).
Pasternack, R. F., Gipp, L. & Sigel, H. Thermodynamics and kinetics of complex formation between cobalt(II), nickel(II) and copper (II) with glycyl-l-leucine and l-leucylglycine. J. Am. Chem. Soc. 94, 8031–8038 (1972).
Konermann, L. Addressing a common misconception: ammonium acetate as neutral pH ‘buffer’ for native electrospray mass spectrometry. J. Am. Soc. Mass. Spectrom. 28, 1827–1835 (2017).
Kuhl, C., Tautenhahn, R., Böttcherr, C., Larson, T. R. & Neumann, S. CAMERA: an integrated strategy for compound spectra extraction and annotation of LC/MS data sets. Anal. Chem. 84, 283–289 (2012).
Broeckling, C. D., Afsar, F. A., Neumann, S., Ben-Hur, A. & Prenni, J. E. RAMClust: a novel feature clustering method enables spectral matching-based annotation for metabolomics data. Anal. Chem. 86, 6812–6817 (2014).
Moree, W. et al. Interkingdom metabolic transformations captured by microbial imaging mass spectrometry. Proc. Natl Acad. Sci. USA 109, 13811–13816 (2012).
Fischbach, M. A. & Clardy, J. One pathway, many products. Nat. Chem. Biol. 3, 353–355 (2007).
Tang, C. Y. & Allen, H. C. Ionic binding of Na+ versus K+ to the carboxylic acid headgroup of palmitic acid monolayers studied by vibrational sum frequency generation spectroscopy. J. Phys. Chem. A 113, 7383–7393 (2009).
Aggerholm, T., Nanita, S. C., Koch, K. J. & Cooks, R. G. Clustering of nucleosides in the presence of alkali metals: biologically relevant quartets of guanosine, deoxyguanosine and uridine observed by ESI-MS/MS. J. Mass Spec. 38, 87–97 (2003).
Moons, R. et al. Metal ions shape α-synuclein. Sci. Rep. 10, 16293 (2020).
Gledhill, M. Electrospray ionisation-mass spectrometry of hydroxamate siderophores. Analyst 126, 1359–1362 (2001).
Harris, W. R., Carrano, C. J. & Raymond, K. N. Coordination chemistry of microbial iron transport compounds. 16. Isolation, characterization and formation constants of ferric aerobactin. J. Am. Chem. Soc. 101, 2722–2728 (1979).
Leib, R. D., Flick, T. G. & Williams, E. R. Direct quantitation of peptide mixtures without standards using clusters formed by electrospray ionization mass spectrometry. Anal. Chem. 81, 3965–3972 (2009).
DeMuth, J. C. & McLuckey, S. A. Electrospray droplet exposure to organic vapors: metal ion removal from proteins and protein complexes. Anal. Chem. 87, 1210–1218 (2015).
Crizer, D. M., Xia, Y. & McLuckey, S. A. Transition metal complex cations as reagents for gas-phase transformation of multiply deprotonated polypeptides. J. Am. Soc. Mass. Spectrom. 20, 1718–1722 (2011).
Huang, T. & McLuckey, S. A. Gas-phase chemistry of multiply charged bioions in analytical mass spectrometry. Annu. Rev. Anal. Chem. 3, 365–385 (2011).
Di Marco, V. B. & Bombi, G. G. Electrospray mass spectrometry (ESI-MS) in the study of metal–ligand solution equilibria. Mass Spectrom. Rev. 25, 347–379 (2006).
Chaturvedi, K. S., Hung, C. S., Crowley, J. R., Stapleton, A. E. & Henderson, J. P. The siderophore yersiniabactin binds copper to protect pathogens during infection. Nat. Chem. Biol. 8, 731–736 (2012).
Sheppard, L. N. & Kontoghiorghes, G. J. Competition between deferiprone, desferrioxamine and other chelators for iron and the effect of other metals. Arzneimittelforschung 43, 659–663 (1993).
Bonham, K. S., Wolfe, B. E. & Dutton, R. J. Extensive horizontal gene transfer in cheese-associated bacteria. eLife 6, e22144 (2017).
Monnet, C., Back, A. & Irlinger, F. Growth of aerobic ripening bacteria at the cheese surface is limited by the availability of iron. Appl. Environ. Microbiol. 78, 3185–3192 (2012).
Blin, K. et al. antiSMASH 4.0—improvements in chemistry prediction and gene cluster boundary identification. Nucleic Acids Res. 45, W36–W41 (2017).
Moscatello, N. J. & Pfeifer, B. A. Yersiniabactin metal binding characterization and removal of nickel from industrial wastewater. Biotech. Prog. 33, 1548–1554 (2017).
Wilks, J. C. & Slonczewski, J. L. pH of the cytoplasm and periplasm of Escherichia coli: rapid measurement by green fluorescent protein fluorimetry. J. Bacteriol. 189, 5601–5607 (2007).
Slonczewski, J. L., Rosen, B. P., Alger, J. R. & Macnab, R. M. pH homeostasis in Escherichia coli: measurement by 31P nuclear magnetic resonance of methylphosphonate and phosphate. Proc. Natl Acad. Sci. USA 78, 6271–6275 (1981).
Xu, G., Guo, H. & Lv, H. Metabolomics assay identified a novel virulence-associated siderophore encoded by the high-pathogenicity island in uropathogenic Escherichia coli. J. Proteome Res. 18, 2331–2336 (2019).
Dührkop, K. et al. SIRIUS 4: a rapid tool for turning tandem mass spectra into metabolite structure information. Nat. Methods 16, 299–302 (2019).
Zhi, H. et al. Siderophore-mediated zinc acquisition enhances enterobacterial colonization of the inflamed gut. Preprint at bioRxiv https://doi.org/10.1101/2020.07.20.212498 (2020).
Chandrangsu, P., Rensing, C. & Helmann, J. D. Metal homeostasis and resistance in bacteria. Nat. Rev. Microbiol. 15, 338–350 (2017).
Keyer, K. & Imlay, J. A. Inactivation of dehydratase [4Fe-4S] clusters and disruption of iron homeostasis upon cell exposure to peroxynitrite. J. Biol. Chem. 272, 27652–27659 (1997).
Gianelli, L., Amendola, V., Fabbrizzi, L., Pallavicini, P. & Mellerio, G. G. Investigation of reduction of Cu(II) complexes in positive‐ion mode electrospray mass spectrometry. Rapid Commun. Mass Spec. 15, 2347–2353 (2001).
Blanco-Ulate, B., Rolshausen, P. E. & Cantu, D. Draft genome sequence of the grapevine dieback fungus Eutypa lata UCR-EL1. Genome Announcement 1, e00228–e00213 (2013).
Reistera, M. et al. Complete genome sequence of the Gram-negative probiotic Escherichia coli strain Nissle 1917. J. Biotechnol. 187, 106–107 (2014).
Hof, C., Eisfeld, K., Antelo, L., Foster, A. J. & Anke, H. Siderophore synthesis in Magnaporthe grisea is essential for vegetative growth, conidiation and resistance to oxidative stress. Fungal Genet. Biol. 46, 321–332 (2009).
Antelo, L. et al. Siderophores produced by Magnaporthe grisea in the presence and absence of iron. Z. Naturforsch C 61, 461–464 (2006).
McRose, D. L., Seyedsayamdost, M. R. & Morel, F. M. M. Multiple siderophores: bug or feature? J. Biol. Inorg. Chem. 23, 983–993 (2018).
Jiang, L. Q., Carter, B. R., Feely, R. A., Lauvset, S. K. & Olsen, A. Surface ocean pH and buffer capacity: past, present and future. Sci. Rep. 9, 18624 (2019).
Rue, E. & Bruland, K. Domoic acid binds iron and copper: a possible role for the toxin produced by the marine diatom Pseudo-nitzschia. Mar. Chem. 76, 127–134 (2001).
Geuer, J. K. et al. Dissolved domoic acid does not improve growth rates and iron content in iron-stressed Pseudo-nitzschia subcurvata. Front. Mar. Sci. 7, 478 (2020).
Sumner, L. W. Proposed minimum reporting standards for chemical analysis Chemical Analysis Working Group (CAWG) Metabolomics Standards Initiative (MSI). Metabolomics. 3, 211–221 (2007).
Acknowledgements
Work in the P.C.D. laboratory was supported by grants P41-GM103484 and GMS10RR029121 and the Betty and Gordon Moore Foundation (A.T.A.). D.P. was supported through the Deutsche Forschungsgemeinschaft with grant PE 2600/1. Work in the M.R. laboratory is supported by Public Health Service grants AI126277, AI114625 and AI145325, by the Chiba University–UCSD Center for Mucosal Immunology, Allergy and Vaccines, and by the UCSD Department of Pediatrics (H.Z. and S.P.N.). M.R. also holds an Investigator in the Pathogenesis of Infectious Disease Award from the Burroughs Wellcome Fund. Work in the R.J.D. laboratory is supported by NIH grant 1 DP2 AT010401-01 (K.P.M. and C.C.S.). We also thank A. Jarmusch and S. McLuckey for helpful discussions and B. Duggan for helpful discussions and assistance with NMR experiments.
Author information
Authors and Affiliations
Contributions
A.T.A., D.P., R.S. and P.C.D. developed the concept. D.P., J.M.G., I.B., L.A., E.T., H.Z., S.P.N., C.C.S., K.P.M., R.J.D. and M.R. cultured organisms and prepared samples. A.T.A., D.P. and J.M.G. ran MS experiments. R.S. wrote code and provided feedback. A.T.A. and D.P. analysed the data. E.T., R.J.D., L.I.A., M.R. and P.C.D. provided supervision and materials. A.T.A., D.P. and P.C.D. wrote the manuscript. All authors edited and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
P.C.D. is a scientific adviser to Sirenas, Galileo and Cybele, and co-founder and scientific advisor to Ometa and Enveda with approval by the University of California, San Diego. M.R. is also on the scientific advisory board of Sirenas.
Additional information
Peer review information Nature Chemistry thanks Corey Broeckling and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary methods, figures, tables and references.
Rights and permissions
About this article
Cite this article
Aron, A.T., Petras, D., Schmid, R. et al. Native mass spectrometry-based metabolomics identifies metal-binding compounds. Nat. Chem. 14, 100–109 (2022). https://doi.org/10.1038/s41557-021-00803-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-021-00803-1
This article is cited by
-
Identification and validation of serum metabolite biomarkers for endometrial cancer diagnosis
EMBO Molecular Medicine (2024)
-
Accessing the specialized metabolome of actinobacteria from the bulk soil of Paullinia cupana Mart. on the Brazilian Amazon: a promising source of bioactive compounds against soybean phytopathogens
Brazilian Journal of Microbiology (2024)
-
Surface manipulation for prevention of migratory viscous crude oil fouling in superhydrophilic membranes
Nature Communications (2023)
-
Gas phase multicomponent detection and analysis combining broadband dual-frequency comb absorption spectroscopy and deep learning
Communications Engineering (2023)
-
Small molecule metabolites: discovery of biomarkers and therapeutic targets
Signal Transduction and Targeted Therapy (2023)