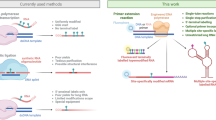
Abstract
DNA, when folded into nanostructures with a specific shape, is capable of spacing and arranging binding sites into a complex geometric pattern with nanometre precision. Here we demonstrate a designer DNA nanostructure that can act as a template to display multiple binding motifs with precise spatial pattern-recognition properties, and that this approach can confer exceptional sensing and potent viral inhibitory capabilities. A star-shaped DNA architecture, carrying five molecular beacon-like motifs, was constructed to display ten dengue envelope protein domain III (ED3)-targeting aptamers into a two-dimensional pattern precisely matching the spatial arrangement of ED3 clusters on the dengue (DENV) viral surface. The resulting multivalent interactions provide high DENV-binding avidity. We show that this structure is a potent viral inhibitor and that it can act as a sensor by including a fluorescent output to report binding. Our molecular-platform design strategy could be adapted to detect and combat other disease-causing pathogens by generating the requisite ligand patterns on customized DNA nanoarchitectures.
Similar content being viewed by others
Main
Infectious diseases, such as viruses, bacteria and toxins, present unique spatial patterns of antigens on their surfaces1,2,3. Existing weakly binding ligands (called ‘binders’ herein) that interact with these epitopes can be linked to a scaffold to improve multivalent binding avidity4,5,6,7,8. However, some viruses, such as dengue (DENV), present complex geometric patterns that cannot be addressed by existing scaffolds because they are not as precise in ligand spacing or provide limited control over the scaffold shape and ligand valency9,10,11,12. Furthermore, previous synthetic scaffolds (polymers, dendrimers, nanofibres, nanoparticles and lipid nanoemulsions) have shown some toxicity. Designer DNA nanostructures13,14,15,16,17 are able to overcome previous limitations because they can be designed into various non-toxic18,19,20,21 and biologically stable21,22,23,24,25 two-dimensional (2D) or 3D platforms to control ligand spacing, valency and spatial arrangements14,26,27. Biocompatible DNA nanostructures have exhibited blood circulation times of up to 24 h (ref. 21) but are eventually removed by the liver21 and kidney28,29.
We demonstrate our strategy by directly targeting DENV through a star-shaped DNA scaffold. This architecture was strategically designed to multivalently arrange binders in a 2D pattern mirroring the complex spatial distribution of dengue epitopes with nanometre precision. Although previous studies have shown that optimizing bivalent interactions is important in protein binding30, cell signalling31 and antibody spatial tolerance using DNA nanostructures32, to our knowledge the importance of multivalent, 2D spatial pattern matching has not yet been explored. To this end, we have included a comparison with scaffolds of different shapes and flexibilities when targeting DENV.
Gold standard diagnostic methods for DENV rely on enzyme-linked immunosorbent assay (ELISA) or quantitative polymerase chain reaction with reverse transcription (RT–qPCR)33,34,35,36. These methods are lengthy, costly and require technical laboratory training. On the therapeutic front, prevention or treatment of viral infection typically relies on neutralizing antibodies (NAbs) against target epitopes on the viral surface. Production of NAbs can be triggered by vaccines or active viruses in the host. However, NAbs may induce antibody-dependent enhancement (ADE) of infection (for example, with DENV37), where antibodies generate increased viral infectivity in vivo, or may not provide protection against new epidemics as a result of genetic drift (for example, antigenic drift in influenza38). A customizable molecular scaffold strategy capable of incorporating sensing motifs with pathogen-specific binders and patterns may address these issues on both fronts.
Results
Design and characterization of the DNA star
The envelope protein domain III (ED3) viral epitope on the DENV surface is organized into a complex polyhedral pattern39 with alternating clusters of trivalent or pentavalent ED3 sites. By connecting the clusters of ED3 sites linearly, we determined that a star shape, consisting of an interior pentagon connected to five exterior triangles, would provide an optimal multivalent scaffold (Fig. 1a). The viral diameter of DENV is 50 nm (ref. 39) and vector distances were measured using PyMOL. The distance between adjacent trivalent–trivalent clusters was 15.1 nm and the distance between adjacent trivalent–pentavalent clusters was 14.1 nm. The following equation was used to calculate the orthodromic distances needed to design the scaffold:
The orthodromic centre-to-centre distances between trivalent–trivalent and trivalent–pentavalent clusters were 15.3 nm and 14.3 nm, respectively. Based on this structural information, the star-shaped DNA scaffold (DNA star) was then designed to not only match the local, orthodromic distances between ED3 clusters but also mirror the global pattern of ED3 clusters aligning with the star’s ten vertices (Fig. 1b).
a, Distribution of DENV ED3 clusters. The diameter of a virion is 50 nm. Orthodromic distances between trivalent–trivalent and trivalent–pentavalent clusters are 15.3 nm and 14.3 nm, respectively. b, The DNA star scaffold is designed to match the pattern and spacing of ED3 clusters. Each DNA strand is represented by an arrow going from the 5′ end to the 3′ end. Non-hybridizing polyT regions are for angle formation between the edges. A 3D representation of the scaffold is also shown. c, A schematic of the partial or complete star complexes (left) and characterization of the DNA star structure using non-denaturing PAGE (right). Yields of the 1-triangle, 2-triangle, 3-triangle, 4-triangle, star and unclosed star complexes are 100%, 98.2%, 81.2%, 86.3%, 88.1% and 98.3%, respectively. Experiments were repeated independently three times, with similar results. d, Native structure of the star, consisting of five ‘scaffold’ strands (S-1 to S-5) to form the internal edges, ten ‘edge’ strands to connect internal and external edges, five ‘fix’ strands to connect the external edges and one ‘close’ strand to cap all the external edges of each triangle. Single-stranded regions of the scaffold’ strands form stem-loops. AFM confirmed the formation. Experiments were repeated independently three times, with similar results.
The DNA star was constructed using the tile-based self-assembly of 21 DNA oligonucleotides (strands). Each of the ten external edges is 42 bp (base pairs) long and connected to the internal edges through four arm junctions at the inner vertices. Each of the five internal edges carries a single-stranded DNA (ssDNA) region that can form a hairpin (stem-loop) structure with a 7 bp stem and 6 bp loop. When the hairpin unzips into ssDNA, it can stretch each internal edge to a distance that fits the distance between adjacent trivalent clusters. These hairpins provide molecular beacon-like motifs for viral detection and offer local structural flexibility to ensure binding of viral targets under various solution conditions and temperatures40.
The formation of the complete DNA star (lane ‘Star’, Fig. 1c) was first characterized by 4% non-denaturing polyacrylamide gel electrophoresis (PAGE), with one of each unique component strand and partial star complex included for reference. Atomic force microscopy (AFM) imaging further confirmed formation of the star (Fig. 1d). For individual DNA nanostructure assemblies, non-denaturing gel electrophoresis is a valid and more reliable method for characterizing structural formation and yield (for example, see previous studies41,42,43). Our non-denaturing PAGE showed a distinct, major band corresponding to each of the 2D architectures, confirming that DNA star structures were formed with high efficiency (Fig. 1c). It is also worth noting that the AFM probe can damage or displace the thin DNA tile-based structures during the scan (Supplementary Fig. 1). Additionally, AFM may display the overall shape of DNA star but cannot fully resolve the details of the structure at the atomic level due to the limited resolution.
DNA star-based DENV sensing
The DNA star was functionalized for DENV detection and inhibition by hybridizing a well-characterized, ED3-binding aptamer44 at each of the 10 vertices of the DNA star to form a star–aptamer complex that geometrically matched and targeted ED3 clusters (Fig. 2a and Supplementary Fig. 2). The aptamer exhibited weak binding to DENV, as determined by surface plasmon resonance (Supplementary Fig. 3). However, the binding strength increased as aptamers were placed onto scaffolds that increasingly matched the pattern of ED3 sites on DENV with the DNA star showing the greatest binding avidity (Supplementary Fig. 3). 6-FAM (fluorophore) and BHQ-1 (quencher) carrying strands were hybridized to each inner edge, flanking the hairpin to turn the DNA star–aptamer complex into a viral sensor. In the absence of DENV, FAM and BHQ-1 molecules would be brought together by canonical Watson–Crick base pairing in the stem, much like a molecular beacon45. However, unlike a molecular beacon, which gives a fluorescent readout of target nucleic acid hybridization events, here the hairpins are pulled apart and converted to ssDNA as a result of strong aptamer binding to ED3 sites. The potent multivalent interactions, promoted by the matched geometric aptamer–ED3 pattern, separated the FAM fluorophores from BHQ-1 quenchers to afford a fluorescent readout.
a, 3′ overhangs on the DNA star allow aptamer incorporation. The 10 incorporated aptamers match the pattern and spacing of ED3 clusters. Five fluorophore–quencher pairs along the inner pentagon of the star remain in a quenching fluorescence resonance energy transfer until binding interactions between aptamers and ED3 domains unzip the hairpins into ssDNA, enabling a fluorescent readout. b, Schematic showing that the DNA star–aptamer complex will remain quenched if it does not recognize its target. In this case, the presence of another virus, the adenovirus, will not enable a fluorescent readout due to insufficient binding. c, A series of DENV concentrations were used to determine the LoD of the star sensor in human serum (1 × 102 p.f.u. ml−1) and human plasma (1 × 103 p.f.u. ml−1). Data are presented as mean ± s.d., n = 3 biologically independent samples. A two-sided t-test was performed to test significance against the background (*P < 0.1; **P < 0.01; ***P < 0.001). Individual data points below background are not shown but were involved in error calculation. d, The star sensor does not detect the presence of the adenovirus in serum. Data are presented as mean ± s.d., n = 3 biologically independent samples. A two-sided t-test was performed to test significance against the background (*P < 0.01; **P < 0.001; ***P < 0.00001). Individual data points below background are not shown but were involved in error calculation.
The DNA star–aptamer sensor was able to directly detect DENV virions with high sensitivity, affording a limit of detection (LoD) of 1 × 102 p.f.u. ml−1 and 1 × 103 p.f.u. ml−1, respectively, in human serum and plasma (Fig. 2c). To exclude the possibility that the series of DENV-mixed DNA star sensing signals obtained within 1–2 min resulted from an automatic (non-target-triggered) separation of the fluorophores from quenchers, we measured fluorescent readouts over time at different temperatures. Fluorescent measurements over time showed that the FAM fluorophores remained quenched during the duration required for sensing and functioned at different temperatures (Supplementary Fig. 4). To examine the specificity of our sensor, we tested it against another virus available to us, the adenovirus. If the star–aptamer complex does not recognize the virus through proper binding, the complex will remain quenched (Fig. 2b). The experimental results verified that the star–aptamer complex designed for DENV detection is not able to sense the adenovirus, even at high concentrations (Fig. 2d).
To make a comparison between the DNA star-based and gold standard methods under similar conditions, we carried out RT–qPCR and ELISA assays to respectively detect DENV RNA and NS1 antigen (Supplementary Fig. 5). The more sensitive method, RT–qPCR, showed a LoD of 1 × 103 p.f.u. ml−1 after we processed different concentrations of DENV-containing human serum (Supplementary Fig. 5), demonstrating the superior sensitivity of our new approach.
In summary, the LoD of our sensor is well below the viral concentration (>1 × 105 p.f.u. ml−1, Supplementary Table 1) in patients on day 0, or the onset of illness, when fever and a variety of symptoms start to occur and the virus begins to become very pathogenic46. Thus, detection before this date is highly beneficial to the patient and to the timely screening and control of pandemic outbreaks within surveillance and diagnostic networks.
Bivalent and linear 1D sensors
To investigate if 2D spatial pattern matching is critical for sensing, we constructed two 1D scaffolds (Fig. 3a). The first was functionalized with two aptamers (bivalent) while the second was functionalized with six aptamers templated by a flexible linear DNA scaffold (abbreviated as linear). In both structures, the strand(s) used to create the DNA star were reused to maintain the same length, sequence, initial aptamer spacing and flexibility wherever possible. Scaffold formation was characterized through 10% non-denaturing PAGE (Supplementary Fig. 7). The flexible linear sensor began detecting DENV with a better sensitivity than the bivalent sensor only at a serum sample containing 1 × 105 p.f.u. ml−1 of the virus. However, both sensors failed to provide comparable sensitivity to the DNA star when standardized under the same fluorophore–quencher pair concentration (Fig. 3c). These control experiments confirm that 2D spatial pattern recognition is important for detection.
a, Schematic showing the design of bivalent, flexible, linear, hexagon-centred and heptagon-centred control sensors. AFM imaging verified formation of the 2D scaffolds. AFM experiments were repeated independently three times, with similar results. b, Schematic demonstrating that ED3 clusters show a hexagonal-shaped pattern when a trivalent ED3 cluster is centred. The pattern-matching hexagon scaffold allows aptamers to strongly bind the ED3 domains, enabling a fluorescent readout. c, At a DENV concentration of 1 × 105 p.f.u. ml−1, the DNA star and hexagon sensors exhibit good detection ability. The linear sensor exhibits poor detection ability. Heptagon and bivalent sensors show little to no sensing. Data are presented as mean ± s.d., n = 3 biologically independent samples. A two-sided t-test was performed to test significance against the background (***P < 0.0001). Individual data points below background are not shown but involved in error calculation. d, A comparison of the DNA star and hexagon sensors in DENV sensing capabilities. The hexagon sensor exhibits a LoD of 103 p.f.u. ml−1. Data are presented as mean ± s.d., n = 3 biologically independent samples. A two-sided t-test was performed to test significance against the background (*P < 0.05; **P < 0.005; ***P < 0.0005). Individual data points below background are not shown but were involved in error calculation.
Hexagon-centred and heptagon-centred 2D sensors
Next, we determined if a correctly matched 2D pattern was required for proper sensing. For that purpose, two additional 2D DNA nanostructures, hexagon- and heptagon-centred star complexes, were constructed and then characterized using AFM and a 4% non-denaturing PAGE (Fig. 3a and Supplementary Fig. 7). The hexagon-centred, six-point star (hexagon) was made by adding an additional, equivalent triangle to the original five-point star design. Similarly, a heptagon-centred, seven-point star (heptagon) was made by adding another equivalent triangle to the hexagon design. We observed that the hexagon provided good sensing capabilities (LoD of 1 × 103 p.f.u. ml−1) while the heptagon could not detect the virus even at a DENV serum sample concentration of 1 × 105 p.f.u. ml−1 when standardized with the same fluorophore–quencher pair concentrations.
The positive sensing provided by the hexagon was expected, because when we analysed DENV surface ED3 patterns, we also recognized a hexagon-shaped distribution of ED3 clusters when centring the DENV virion on a trivalent cluster (Fig. 3b); this additional pattern differed from the pentavalent cluster-centred star shape (Fig. 1a). The fact that the hexagon can serve as a good positive control sensor proves the need for pattern matching in detection. The sensitivity of the hexagon sensor was tested, providing a LoD of 1 × 103 p.f.u. ml−1 (Fig. 3d). Although the heptagon carried more aptamers than the hexagon, the sensor resulted in poor to no detection. This comparison is particularly interesting as it demonstrates that a small mismatch of 2D geometry, under the same bivalently correct spacing and with higher ligand valency and density (as compared to the hexagon), leads to dramatically reduced sensing capabilities.
In vitro DENV inhibition
Encouraged by the previous results, we investigated the in vitro inhibition of DENV through standard antiviral, plaque-forming half-maximum effective concentration (EC50) assays. Briefly, the DENV viral particles were incubated with different concentrations of each inhibitor in human serum and the remaining infectivity was determined by a plaque reduction assay (Fig. 4a). The dose-dependent inhibition of DENV for the monovalent aptamer and each of the scaffold–aptamer complexes was examined (Fig. 4b). The EC50 value of the DNA star–aptamer complex for DENV infection inhibition was 2 nM, whereas the EC50 value for the monovalent aptamer was 15 µM (Fig. 4c,d). These results demonstrate that the DNA star–aptamer multivalent inhibitor was 7.5 × 103-fold more effective than the monovalent aptamer for the in vitro inhibition of DENV infection in human serum.
a, Representative plaque assays corresponding to the aptamer concentration and scaffold. The no-inhibitor treated control well is on the left, then the other wells are arranged from highest reduction to lowest reduction from left to right, regardless of 2D or 1D scaffolding. The experiments were repeated independently three times, with similar results. b, Dose-dependent, plaque-reducing inhibition curves for the monovalent aptamer (abbreviated as aptamer), heptagon, bivalent, flexible linear (abbreviated as linear), hexagon and star inhibitors. Inhibitor concentration was standardized through aptamer concentration. Data are presented as mean ± s.d., n = 3 biologically independent experiments. c, A comparison of each inhibitor’s fold change in EC50. Data are presented as mean ± s.d., n = 3 biologically independent experiments. A two-sided t-test was performed to test significance against the DNA star (**P < 0.005; ***P ≤ 0.001; ****P < 0.0001). d, A schematic representing the inhibitory nanostructures with their corresponding mean EC50 values ± s.d., n = 3 biologically independent experiments. The schematic for the star and hexagon show an unzipped hairpin region (as part of the dark blue strands), because potent pattern matching occurs. Other scaffolds represent the hairpins as stem-loop structures to indicate a lack of potent pattern matching.
We also examined the EC50 values of the other control scaffolds, where a lower EC50 value demonstrates more potent inhibition (Fig. 4b–d). In correlation with our DENV sensing data, the hexagon scaffold provided the second most potent inhibitory effect, with an EC50 value of 10 nM, while the heptagon scaffold exhibited poor inhibition, with an EC50 value of 440 nM. The flexible linear scaffold provided better inhibition, at an EC50 of 90 nM, than the bivalent scaffold, at an EC50 of 130 nM. Additionally, as previous studies predicted, potentially matching the spacing could increase the inhibitory effect as all the structures proved to be more potent than the monovalent aptamer.
These results point out that stronger inhibition was achieved when the spatial pattern was properly matched. Interestingly, the 2D heptagon scaffold exhibited worse inhibitory potency than the 1D bivalent and flexible linear scaffolds. A mismatch of a fixed geometric pattern, while keeping the same initial spacing, leads to reduced inhibition potency when compared to flexibly matching the spacing. In summary, the more the binders match the 2D pattern of binding sites, the stronger the inhibition as aptamers bound to the star scaffold show superior performance compared to the other scaffold shapes.
Confocal imaging
Time-lapsed, live confocal imaging demonstrated that DENV virions lose their cell internalization ability after being bound by the DNA star–aptamer complex (Fig. 5). For the assay, the HepG2 line was chosen as it is derived from the liver, an organ DENV infects, and was cultured in medium containing human serum. Membrane-bound structures, including the cell membrane, were stained with Dil (yellow) while cell nuclei (blue) were stained with Hoechst. DENV virions were labelled with DiD (red) and the DNA star–aptamer complex was labelled with the 6-FAM fluorophore (green), quenched in the absence of DENV. When the green DNA stars and the red virions co-localize, the merged signals appear yellow. The Dil stain and 6-FAM dye also exhibit fluorescence spillover due to slightly overlapping excitation and emission spectra. The finer details between labelled structures can be resolved when viewing the merged image as separate fluorescence channels (Supplementary Fig. 8).
a, DENV virions (red), membrane bound structures (yellow) and cell nuclei (blue) were labelled with DiD, Dil and Hoescht stains, respectively. Due to the combined effects of fluorescence spillover and the merged red and green signals, co-localized DNA star-bound virions appear yellow. Representative viral particles are indicated with white arrows. Top row: viral accumulation over time for the no-inhibitor treated condition. Bottom row: viral entry inhibition over time during DNA star treatment. Each panel is accompanied by a cross-section of the same image (along the dotted lines in the main images). An eye symbol orients the viewing direction for the cross-sections reconstructed from Z stacks, with images taken at different focal planes. b, A volume reconstruction of a close-up confocal view shows unbound DENV particles (red spheres) accumulating in the cell (top panels) and a DNA star-bound DENV particle (green sphere) inhibited from cell entry (bottom panels). Each confocal experiment was performed in singlicate.
Unbound or DNA star-bound virions were introduced to the labelled cells (Fig. 5, Supplementary Fig. 9 and Supplementary Videos 1 and 2). In the unbound condition, DENV accumulated into the cells over time. In the DNA star-bound condition, DENV accumulation was drastically reduced and virions were generally electrostatically repelled from cells. Even if some DNA star-bound virions reached the cell membrane, they were unable to enter cells over the time course of an hour.
A cross-section (indicated by the white dotted line in Fig. 5a) accompanies each panel, with an eye symbol denoting the viewing direction. The view was reconstructed by images taken from different planes (Z stacks) during imaging. Volume reconstruction using Imaris software confirmed viral accumulation in the unbound condition and viral entry inhibition when bound to the DNA star (Fig. 5b).
Discussion
By targeting a challenging exemplar platform, the DENV flavivirus, we have demonstrated the importance of integrating a structurally defined DNA nanoarchitecture with precise, multivalent spatial pattern-recognizing properties. Sensors and inhibitors recognizing the DENV pattern identity allow them to work effectively. When aptamers are arranged into an optimal 2D shape, they can be induced to fit the correct DENV pattern. In contrast, linear complexes, such as the bivalent–aptamer complex, initially have neither optimal shape nor optimal spacing, so they have weak affinity to ED3 sites. Moreover, for those that bind transiently, they have trouble staying on because the hairpin region also has affinity to the base pair. Without the optimal shape identity to keep these hairpin interactions from forming, sensing and inhibitory abilities suffer.
We also observed that an incorrect shape will be more detrimental than having no shape at all, as observed with the heptagon scaffold. We speculate that the heptagon–aptamer complex can bind, in the best-case scenario, bivalently to two ED3 clusters, but that leaves certain sites unbound and capable of cell internalization (Supplementary Fig. 7). In addition, a bound heptagon–aptamer complex would sterically prevent more heptagon–aptamer complexes from binding while linear scaffolds do not experience a steric block to such a degree.
Our DNA star sensor shows superiority over current gold standard DENV detection methods. The reduced sensitivity of RT–qPCR can result from the low amount of starting material (one genome copy per viral particle), RNA extraction process and instability of the extracted RNA. The star sensor, on the other hand, has direct access to the unprocessed viral sample. Up to two stars can specifically bind to a single viral particle, translating to a 10-fluorophore label (five per star). Additionally, the immunoglobulin-M (IgM) ELISA or IgM rapid test cannot provide early viral sensing as antibody production in the body requires several days after initial infection. So, no gold standard method for DENV detection can achieve the same sensing capacity in terms of cost, ease, sensitivity and speed (see Supplementary Fig. 6, Supplementary Table 1, and Note for Supplementary Table 1 for a detailed comparison). As is standard with viral infection screening, secondary confirmation of infection by some of the standard methods (for example, viral isolation or nucleic acid sensing) should still be employed after the initial precautionary screening in order to make a full documentation and evaluation of an epidemic outbreak.
The performance of the DNA star during in vitro DENV inhibition shows promising therapeutic potential. As is the case for various infectious diseases (for example, the malaria parasite47, DENV48, Zika49 and influenza A8), interactions with negatively charged glycosaminoglycans (GAGs) are required for host cell invasion. Thus, the DNA architecture not only relies on specific multivalent interactions for binding DENV but also electrostatically traps and isolates virions from the host cell plasma membrane and GAGs via the negative charges of the scaffold. Moreover, other monovalent aptamer combinations have been used as antiviral agents for human viruses50. Thus, our scaffold-based strategy can be used to arrange these same aptamers into arrangements that increase specific, on-target binding while reducing off-target binding based on pattern identity.
As binders specific to other binding sites can be evolved or synthesized, our strategy could be widely applicable, and such scaffolds could be used to efficiently detect and inhibit a wide-range of pathogens (Fig. 6), modulate cell signalling by analysing the distribution of cellular receptors, or organize previous drugs into more potent configurations. Although DNA nanostructures have shown certain stability in vivo21,29, the component DNA strands can be chemically modified to further improve its stability.
a, Selecting, synthesizing or evolving a specific binder. Binders can include, but are not limited to, peptides, aptamers, oligosaccharides or small molecules. b, Analysis of the spatial surface pattern of the binding sites. c, Designing a DNA nanostructure that mirrors the spatial surface pattern. d, Incorporating binders at appropriate locations onto the DNA. e, Adapting the potent, multivalent DNA nanostructure–binder complex for the intended application. Two more simple examples, where curvature is not accounted for, are used for illustration. b–e, Top: a heptagon-shaped DNA structure, containing a seven-arm junction, matches heptavalent binding sites on anthrax. Bottom: a triangle-shaped DNA structure matches haemagglutinin trimers on influenza virus.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Online content
Any Nature Research reporting summaries, source data, extended data, supplementary information, acknowledgements, peer review information; details of author contributions and competing interests; and statements of data and code availability are available at https://doi.org/10.1038/s41557-019-0369-8.
Data availability
All data supporting the findings of this work are available within this paper (figures, videos and description) and its Supplementary Information. All other data are available from the corresponding authors upon request.
Code availability
The SEQUIN program used in this study is available from the corresponding authors upon reasonable request.
References
Harrison, S. C. Virology: looking inside adenovirus. Science 329, 1026–1027 (2010).
Krantz, B. A. et al. A phenylalanine clamp catalyzes protein translocation through the anthrax toxin pore. Science 309, 777–781 (2005).
Bandlow, V. et al. Spatial screening of hemagglutinin on influenza A virus particles: sialyl-LacNAc displays on DNA and PEG scaffolds reveal the requirements for bivalency enhanced interactions with weak monovalent binders. J. Am. Chem. Soc. 139, 16389–16397 (2017).
Mourez, M. et al. Designing a polyvalent inhibitor of anthrax toxin. Nat. Biotechnol. 19, 958–961 (2001).
Rai, P. et al. Statistical pattern matching facilitates the design of polyvalent inhibitors of anthrax and cholera toxins. Nat. Biotechnol. 24, 582–586 (2006).
Joshi, A. et al. Nanotube-assisted protein deactivation. Nat. Nanotechnol. 3, 41–45 (2008).
Kitov, P. I. et al. Shiga-like toxins are neutralized by tailored multivalent carbohydrate ligands. Nature 403, 669–672 (2000).
Kwon, S. J. et al. Nanostructured glycan architecture is important in the inhibition of influenza A virus infection. Nat. Nanotechnol. 12, 48–54 (2017).
Ahmad, K. M., Xiao, Y. & Soh, H. T. Selection is more intelligent than design: improving the affinity of a bivalent ligand through directed evolution. Nucleic Acids Res. 40, 11777–11783 (2012).
King, D. J. & Noss, R. R. Toxicity of polyacrylamide and acrylamide monomer. Rev. Environ. Health 8, 3–16 (1989).
Malik, N. et al. Dendrimers: relationship between structure and biocompatibility in vitro, and preliminary studies on the biodistribution of 125i-labelled polyamidoamine dendrimers in vivo. J. Control. Release 65, 133–148 (2000).
Strauch, E. M. et al. Computational design of trimeric influenza-neutralizing proteins targeting the hemagglutinin receptor binding site. Nat. Biotechnol. 35, 667–671 (2017).
Lin, C., Liu, Y., Rinker, S. & Yan, H. DNA tile based self-assembly: building complex nanoarchitectures. ChemPhysChem 7, 1641–1647 (2006).
Chandrasekaran, A. R., Anderson, N., Kizer, M., Halvorsen, K. & Wang, X. Beyond the fold: emerging biological applications of DNA origami. ChemBioChem 17, 1081–1089 (2016).
Hong, F., Zhang, F., Liu, Y. & Yan, H. DNA origami: scaffolds for creating higher order structures. Chem. Rev. 117, 12584–12640 (2017).
Hu, Q., Li, H., Wang, L., Gu, H. & Fan, C. DNA nanotechnology-enabled drug delivery systems. Chem. Rev. 119, 6459–6506 (2019).
Seeman, N. C. Nucleic acid junctions and lattices. J. Theor. Biol. 99, 237–247 (1982).
Zhang, Q. et al. DNA origami as an in vivo drug delivery vehicle for cancer therapy. ACS Nano 8, 6633–6643 (2014).
Lee, D. S., Qian, H., Tay, C. Y. & Leong, D. T. Cellular processing and destinies of artificial DNA nanostructures. Chem. Soc. Rev. 45, 4199–4225 (2016).
Vinther, M. & Kjems, J. Interfacing DNA nanodevices with biology: challenges, solutions and perspectives. New J. Phys. 18, 085005 (2016).
Li, S. et al. A DNA nanorobot functions as a cancer therapeutic in response to a molecular trigger in vivo. Nat. Biotechnol. 36, 258–264 (2018).
Mei, Q. A. et al. Stability of DNA origami nanoarrays in cell lysate. Nano Lett. 11, 1477–1482 (2011).
Hahn, J., Wickham, S. F. J., Shih, W. M. & Perrault, S. D. Addressing the instability of DNA nanostructures in tissue culture. ACS Nano 8, 8765–8775 (2014).
Agarwal, N. P., Matthies, M., Gur, F. N., Osada, K. & Schmidt, T. L. Block copolymer micellization as a protection strategy for DNA origami. Angew. Chem. Int. Ed. 56, 5460–5464 (2017).
Perrault, S. D. & Shih, W. M. Virus-inspired membrane encapsulation of DNA nanostructures to achieve in vivo stability. ACS Nano 8, 5132–5140 (2014).
Rothemund, P. W. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Wilner, O. I. & Willner, I. Functionalized DNA nanostructures. Chem. Rev. 112, 2528–2556 (2012).
Lee, H. et al. Molecularly self-assembled nucleic acid nanoparticles for targeted in vivo siRNA delivery. Nat. Nanotechnol. 7, 389–393 (2012).
Jiang, D. et al. DNA origami nanostructures can exhibit preferential renal uptake and alleviate acute kidney injury. Nat. Biomed. Eng. 2, 865–877 (2018).
Rinker, S. et al. Nanostructures for distance-dependent multivalent ligand–protein binding. Nat. Nanotechnol. 3, 418–422 (2008).
Shaw, A. et al. Spatial control of membrane receptor function using ligand nanocalipers. Nat. Methods. 11, 841–846 (2014).
Shaw, A. et al. Binding to nanopatterned antigens is dominated by the spatial tolerance of antibodies. Nat. Nanotechnol. 14, 184–190 (2019).
Storch, G. A. Diagnostic virology. Clin. Infect. Dis. 31, 739–751 (2000).
Read, S. J., Burnett, D. & Fink, C. G. Molecular techniques for clinical diagnostic virology. J. Clin. Pathol. 53, 502–506 (2000).
Scof, S. Recent advances in diagnostic testing for viral infections. Biosci. Horizons 9, 1–11 (2016).
Dengue: Guidelines for Diagnosis, Treatment, Prevention and Control: New Edition Ch. 4 (World Health Organization, 2009).
Whitehead, S. S., Blaney, J. E., Durbin, A. P. & Murphy, B. R. Prospects for a dengue virus vaccine. Nat. Rev. Microbiol. 5, 518–528 (2007).
Boni, M. F. Vaccination and antigenic drift in influenza. Vaccine 26, C8–C14 (2008).
Zhang, X. et al. Cryo-EM structure of the mature dengue virus at 3.5-Å resolution. Nat. Struct. Mol. Biol. 20, 105–110 (2013).
Fibriansah, G. et al. Structural changes in dengue virus when exposed to a temperature of 37 °C. J. Virol. 87, 7585–7592 (2013).
Chen, J. H. & Seeman, N. C. Synthesis from DNA of a molecule with the connectivity of a cube. Nature 350, 631–633 (1991).
Wang, X. & Seeman, N. C. Assembly and characterization of 8-arm and 12-arm DNA branched junctions. J. Am. Chem. Soc. 129, 8169–8176 (2007).
Ke, Y. et al. Scaffolded DNA origami of a DNA tetrahedron molecular container. Nano Lett. 9, 2445–2447 (2009).
Chen, H. L., Hsiao, W. H., Lee, H. C., Wu, S. C. & Cheng, J. W. Selection and characterization of DNA aptamers targeting all four serotypes of dengue viruses. PLoS One 10, e0131240 (2015).
Tyagi, S. & Kramer, F. R. Molecular beacons: probes that fluoresce upon hybridization. Nat. Biotechnol. 14, 303–308 (1996).
Gubler, D. J. Dengue and dengue hemorrhagic fever. Clin. Microbiol. Rev. 11, 480–496 (1998).
Beaudet, J. M. et al. Characterization of human placental glycosaminoglycans and regional binding to VAR2CSA in malaria infected erythrocytes. Glycoconj. J. 31, 109–116 (2014).
Chen, Y. et al. Dengue virus infectivity depends on envelope protein binding to target cell heparan sulfate. Nat. Med. 3, 866–871 (1997).
Kim, S. Y. et al. Interaction of Zika virus envelope protein with glycosaminoglycans. Biochemistry 56, 1151–1162 (2017).
Gonzalez, V. M., Martin, M. E., Fernandez, G. & Garcia-Sacristan, A. Use of aptamers as diagnostics tools and antiviral agents for human viruses. Pharmaceuticals 9, 78 (2016).
Acknowledgements
We thank the core facilities at RPI-CBIS, Wadsworth Center of New York State Department of Health, Organic Electronics and Information Displays and Jiangsu Key Laboratory and Institute of Advanced Materials of Nanjing University of Posts and Telecommunications (China). This work was funded by an RPI-CBIS start-up fund and award, a gift fund from HT Materials Corporation to X.W., the Ministry of Science and Technology of China (2017YFA0205302), the National Science Foundation of China (21922408 and 61771253), the Natural Science Foundation of Jiangsu Province for Distinguished Young Scholars (BK20190038) to J.C., the Global Research Laboratory Program through the National Research Foundation of Korea (2014K1A1A2043032) to J.S.D. and S.-J.K., and the National Institute of Health (DK111958) to R.J.L. The microscopy for time-lapsed analysis is supported by the National Science Foundation (NSF-MRI-1725984).
Author information
Authors and Affiliations
Contributions
P.S.K., S.-J.K., J.S.D., R.J.L. and X.W. conceived the original 2D spatial pattern-matching strategy. P.S.K., S.-J.K., R.J.L., J.C. and X.W. conceived and designed the experiments. S.R. and J.C. designed the sequences of the DNA scaffolds and performed non-denaturing PAGE. S.R., M.X., D.Z., J.C., F. Zhou and N.C.S. performed the AFM experiments. P.S.K., S.R. and M.E.K. purified and folded the DNA. L.K. and L.D.K. prepared viruses, conducted RT–qPCR detection and performed the plaque forming assays. F. Zhang performed SPR analysis. P.S.K., S.-J.K. and D.K. performed confocal microscopy. P.S.K. performed all other experiments. K.F. analysed the aptamer binding pattern of DENV. P.S.K., S.-J.K. and X.W. performed data analysis. P.S.K., S.-J.K., R.J.L. and X.W. led the preparation of the manuscript with contributions from all authors. P.S.K., S.R., S.-J.K., M.E.K. and L.K. contributed equally to this work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary materials, methods, Figs. 1–9, Tables 1–11, notes and references.
Supplementary Video 1 (unbound condition)
Time-lapsed, live confocal imaging of dengue cell internalization.
Supplementary Video 2 (DNA star-bound condition)
Time-lapsed, live confocal imaging of DNA star-bound dengue losing cell internalization ability.
Rights and permissions
About this article
Cite this article
Kwon, P.S., Ren, S., Kwon, SJ. et al. Designer DNA architecture offers precise and multivalent spatial pattern-recognition for viral sensing and inhibition. Nat. Chem. 12, 26–35 (2020). https://doi.org/10.1038/s41557-019-0369-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-019-0369-8
This article is cited by
-
A DNA origami device spatially controls CD95 signalling to induce immune tolerance in rheumatoid arthritis
Nature Materials (2024)
-
Nanotechnology-based theranostic and prophylactic approaches against SARS-CoV-2
Immunologic Research (2024)
-
Fine tuning of CpG spatial distribution with DNA origami for improved cancer vaccination
Nature Nanotechnology (2024)
-
Exploring the diverse biomedical applications of programmable and multifunctional DNA nanomaterials
Journal of Nanobiotechnology (2023)
-
A temporally resolved DNA framework state machine in living cells
Nature Machine Intelligence (2023)