Abstract
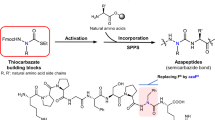
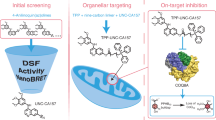
Rapamycin and FK506 are macrocyclic natural products with an extraordinary mode of action, in which they form binary complexes with FK506-binding protein (FKBP) through a shared FKBP-binding domain before forming ternary complexes with their respective targets, mechanistic target of rapamycin (mTOR) and calcineurin, respectively. Inspired by this, we sought to build a rapamycin-like macromolecule library to target new cellular proteins by replacing the effector domain of rapamycin with a combinatorial library of oligopeptides. We developed a robust macrocyclization method using ring-closing metathesis and synthesized a 45,000-compound library of hybrid macrocycles (named rapafucins) using optimized FKBP-binding domains. Screening of the rapafucin library in human cells led to the discovery of rapadocin, an inhibitor of nucleoside uptake. Rapadocin is a potent, isoform-specific and FKBP-dependent inhibitor of the equilibrative nucleoside transporter 1 and is efficacious in an animal model of kidney ischaemia reperfusion injury. Together, these results demonstrate that rapafucins are a new class of chemical probes and drug leads that can expand the repertoire of protein targets well beyond mTOR and calcineurin.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the authors upon reasonable request.
References
Sehgal, S. N., Baker, H. & Vezina, C. Rapamycin (AY-22,989), a new antifungal antibiotic. II. Fermentation, isolation and characterization. J. Antibiot. 28, 727–732 (1975).
Tanaka, H. et al. Structure of FK506, a novel immunosuppressant isolated from Streptomyces. J. Am. Chem. Soc. 109, 5031–5033 (1987).
Harding, M. W., Galat, A., Uehling, D. E. & Schreiber, S. L. A receptor for the immunosuppressant FK506 is a cis–trans peptidyl-prolyl isomerase. Nature 341, 758–760 (1989).
Siekierka, J. J., Hung, S. H., Poe, M., Lin, C. S. & Sigal, N. H. A cytosolic binding protein for the immunosuppressant FK506 has peptidyl-prolyl isomerase activity but is distinct from cyclophilin. Nature 341, 755–757 (1989).
Heitman, J., Movva, N. R. & Hall, M. N. Targets for cell cycle arrest by the immunosuppressant rapamycin in yeast. Science 253, 905–909 (1991).
Liu, J. et al. Calcineurin is a common target of cyclophilin–cyclosporin A and FKBP–FK506 complexes. Cell 66, 807–815 (1991).
Yang, H. et al. mTOR kinase structure, mechanism and regulation. Nature 497, 217–223 (2013).
Griffith, J. P. et al. X-ray structure of calcineurin inhibited by the immunophilin-immunosuppressant FKBP12–FK506 complex. Cell 82, 507–522 (1995).
Kissinger, C. R. et al. Crystal structures of human calcineurin and the human FKBP12–FK506–calcineurin complex. Nature 378, 641–644 (1995).
Marinec, P. S. et al. FK506-binding protein (FKBP) partitions a modified HIV protease inhibitor into blood cells and prolongs its lifetime in vivo. Proc. Natl Acad. Sci. USA 106, 1336–1341 (2009).
Klemm, J. D., Schreiber, S. L. & Crabtree, G. R. Dimerization as a regulatory mechanism in signal transduction. Annu. Rev. Immunol. 16, 569–592 (1998).
Bayle, J. H. et al. Rapamycin analogs with differential binding specificity permit orthogonal control of protein activity. Chem. Biol. 13, 99–107 (2006).
Guduru, S. K. R. & Arya, P. Synthesis and biological evaluation of rapamycin-derived, next generation small molecules. Med. Chem. Commun. 9, 27–43 (2018).
Chakraborty, T. K., Weber, H. P. & Nicolaou, K. C. Design and synthesis of a rapamycin-based high affinity binding FKBP12 ligand. Chem. Biol. 2, 157–161 (1995).
Wu, X. et al. Creating diverse target-binding surfaces on FKBP12: synthesis and evaluation of a rapamycin analogue library. ACS Comb. Sci. 13, 486–495 (2011).
Li, W., Bhat, S. & Liu, J. O. A simple and efficient route to the FKBP-binding domain from rapamycin. Tetrahedron Lett. 52, 5070–5072 (2011).
Furka, A., Sebestyen, F., Asgedom, M. & Dibo, G. General method for rapid synthesis of multicomponent peptide mixtures. Int. J. Pept. Protein Res. 37, 487–493 (1991).
Houghten, R. A. et al. Generation and use of synthetic peptide combinatorial libraries for basic research and drug discovery. Nature 354, 84–86 (1991).
Lam, K. S. et al. A new type of synthetic peptide library for identifying ligand-binding activity. Nature 354, 82–84 (1991).
Reichwein, J. F., Wels, B., Kruijtzer, J. A., Versluis, C. & Liskamp, R. M. Rolling loop scan: an approach featuring ring-closing metathesis for generating libraries of peptides with molecular shapes mimicking bioactive conformations or local folding of peptides and proteins. Angew. Chem. Int. Ed. 38, 3684–3687 (1999).
Liu, J. et al. Inhibition of T cell signaling by immunophilin–ligand complexes correlates with loss of calcineurin phosphatase activity. Biochemistry 31, 3896–3901 (1992).
Halt, D. A. et al. Design, synthesis and kinetic evaluation of high-affinity FKBP ligands and the X-ray crystal structures of their complexes with FKBP12. J. Am. Chem. Soc. 115, 9925–9938 (1993).
Clackson, T. et al. Redesigning an FKBP–ligand interface to generate chemical dimerizers with novel specificity. Proc. Natl Acad. Sci. USA 95, 10437–10442 (1998).
Sagan, S., Karoyan, P., Lequin, O., Chassaing, G. & Lavielle, S. N- and Calpha-methylation in biologically active peptides: synthesis, structural and functional aspects. Curr. Med. Chem. 11, 2799–2822 (2004).
Ahmed, S. A., Gogal, R. M. Jr & Walsh, J. E. A new rapid and simple non-radioactive assay to monitor and determine the proliferation of lymphocytes: an alternative to [3H]thymidine incorporation assay. J. Immunol. Methods 170, 211–224 (1994).
Young, J. D., Yao, S. Y., Baldwin, J. M., Cass, C. E. & Baldwin, S. A. The human concentrative and equilibrative nucleoside transporter families, SLC28 and SLC29. Mol. Aspects Med. 34, 529–547 (2013).
Owen, R. P. et al. Functional characterization and haplotype analysis of polymorphisms in the human equilibrative nucleoside transporter, ENT2. Drug Metab. Dispos. 34, 12–15 (2006).
Boswell-Casteel, R. C. & Hays, F. A. Equilibrative nucleoside transporters—a review. Nucleos. Nucleot. Nucl. 36, 7–30 (2017).
Xiao, J. C., Zhang, T. P. & Zhao, Y. P. Human equilibrative nucleoside transporter 1 (hENT1) predicts the Asian patient response to gemcitabine-based chemotherapy in pancreatic cancer. Hepato-Gastroenterol. 60, 258–262 (2013).
Meijer, L. L., Puik, J. R., Peters, G. J., Kazemier, G. & Giovannetti, E. hENT-1 Expression and localization predict outcome after adjuvant gemcitabine in resected cholangiocarcinoma patients. Oncologist 21, e4 (2016).
Jacobson, K. A. & Gao, Z. G. Adenosine receptors as therapeutic targets. Nat. Rev. Drug Discov. 5, 247–264 (2006).
Loffler, M., Morote-Garcia, J. C., Eltzschig, S. A., Coe, I. R. & Eltzschig, H. K. Physiological roles of vascular nucleoside transporters. Arterioscler. Thromb. Vasc. Biol. 27, 1004–1013 (2007).
Headrick, J. P. & Lasley, R. D. Adenosine receptors and reperfusion injury of the heart. Handb. Exp. Pharmacol. 2009, 189–214 (2009).
Cass, C. E. & Paterson, A. R. Inhibition by nitrobenzylthioinosine of uptake of adenosine, 2′-deoxyadenosine and 9-β-d-arabinofuranosyladenine by human and mouse erythrocytes. Biochem. Pharmacol. 24, 1989–1993 (1975).
Scholtissek, C. Studies on the uptake of nucleic acid precursors into cells in tissue culture. Biochim. Biophys. Acta 158, 435–447 (1968).
Ward, J. L., Sherali, A., Mo, Z. P. & Tse, C. M. Kinetic and pharmacological properties of cloned human equilibrative nucleoside transporters, ENT1 and ENT2, stably expressed in nucleoside transporter-deficient PK15 cells. Ent2 exhibits a low affinity for guanosine and cytidine but a high affinity for inosine. J. Biol. Chem. 275, 8375–8381 (2000).
Rehan, S. & Jaakola, V. P. Expression, purification and functional characterization of human equilibrative nucleoside transporter subtype-1 (hENT1) protein from Sf9 insect cells. Protein Expr. Purif. 114, 99–107 (2015).
Rehan, S., Ashok, Y., Nanekar, R. & Jaakola, V. P. Thermodynamics and kinetics of inhibitor binding to human equilibrative nucleoside transporter subtype-1. Biochem. Pharmacol. 98, 681–689 (2015).
Hammond, J. R. Interaction of a series of draflazine analogues with equilibrative nucleoside transporters: species differences and transporter subtype selectivity. Naunyn Schmiedebergs Arch. Pharmacol. 361, 373–382 (2000).
Bierer, B. E. et al. Two distinct signal transmission pathways in T lymphocytes are inhibited by complexes formed between an immunophilin and either FK506 or rapamycin. Proc. Natl Acad. Sci. USA 87, 9231–9235 (1990).
Chresta, C. M. et al. AZD8055 is a potent, selective and orally bioavailable ATP-competitive mammalian target of rapamycin kinase inhibitor with in vitro and in vivo antitumor activity. Cancer Res. 70, 288–298 (2010).
Fischer, G., Wittmann-Liebold, B., Lang, K., Kiefhaber, T. & Schmid, F. X. Cyclophilin and peptidyl-prolyl cis–trans isomerase are probably identical proteins. Nature 337, 476–478 (1989).
Korchynskyi, O. & ten Dijke, P. Identification and functional characterization of distinct critically important bone morphogenetic protein-specific response elements in the Id1 promoter. J. Biol. Chem. 277, 4883–4891 (2002).
Kugimiya, F. et al. Mechanism of osteogenic induction by FK506 via BMP/Smad pathways. Biochem. Biophys. Res. Commun. 338, 872–879 (2005).
Spiekerkoetter, E. et al. FK506 activates BMPR2, rescues endothelial dysfunction, and reverses pulmonary hypertension. J. Clin. Invest. 123, 3600–3613 (2013).
Day, Y. J., Huang, L., Ye, H., Linden, J. & Okusa, M. D. Renal ischemia-reperfusion injury and adenosine 2A receptor-mediated tissue protection: role of macrophages. Am. J. Physiol. Renal Physiol. 288, F722–F731 (2005).
Lappas, C. M., Day, Y. J., Marshall, M. A., Engelhard, V. H. & Linden, J. Adenosine A2A receptor activation reduces hepatic ischemia reperfusion injury by inhibiting CD1d-dependent NKT cell activation. J. Exp. Med. 203, 2639–2648 (2006).
Grenz, A. et al. The reno-vascular A2B adenosine receptor protects the kidney from ischemia. PLoS Med. 5, e137 (2008).
Liu, M. et al. Acute kidney injury leads to inflammation and functional changes in the brain. J. Am. Soc. Nephrol. 19, 1360–1370 (2008).
Yan, L. & Muller, C. E. Preparation, properties, reactions and adenosine receptor affinities of sulfophenylxanthine nitrophenyl esters: toward the development of sulfonic acid prodrugs with peroral bioavailability. J. Med. Chem. 47, 1031–1043 (2004).
Arai, T., Kouama, Y., Suenaga, T. & Honda, H. Ascomycin, an antifungal antibiotic. J. Antibiot. 15, 231–232 (1962).
Hatanaka, H. et al. FR-900520 and FR-900523, novel immunosuppressants isolated from a Streptomyces. II. Fermentation, isolation and physico-chemical and biological characteristics. J. Antibiot. 41, 1592–1601 (1988).
Hasko, G., Linden, J., Cronstein, B. & Pacher, P. Adenosine receptors: therapeutic aspects for inflammatory and immune diseases. Nat. Rev. Drug Discov. 7, 759–770 (2008).
Vaswani, M., Linda, F. K. & Ramesh, S. Role of selective serotonin reuptake inhibitors in psychiatric disorders: a comprehensive review. Prog. Neuropsychopharmacol. Biol. Psychiatry 27, 85–102 (2003).
Chen, J. F., Eltzschig, H. K. & Fredholm, B. B. Adenosine receptors as drug targets—what are the challenges? Nat. Rev. Drug Discov. 12, 265–286 (2013).
Laplante, M. & Sabatini, D. M. mTOR signaling in growth control and disease. Cell 149, 274–293 (2012).
Dazert, E. & Hall, M. N. mTOR signaling in disease. Curr. Opin. Cell Biol. 23, 744–755 (2011).
Liu, J. O. Calmodulin-dependent phosphatase, kinases and transcriptional corepressors involved in T-cell activation. Immunol. Rev. 228, 184–198 (2009).
Schreiber, S. L. & Crabtree, G. R. The mechanism of action of cyclosporin A and FK506. Immunol. Today 13, 136–142 (1992).
Rao, A., Luo, C. & Hogan, P. G. Transcription factors of the NFAT family: regulation and function. Annu. Rev. Immunol. 15, 707–747 (1997).
Acknowledgements
This work was made possible by the NIH Director’s Pioneer Award, the Flight Attendant Medical Research Institute and a generous gift from S. Yan and H. Mao (J.O.L.), a Damon Runyon Postdoctoral Fellowship (H.P.) and an NIH Postdoctoral Training Award (M.D.). V.O.P. is supported by the Academy of Finland (grant no. 289737) and the Sigrid Juselius Foundation. The authors thank S.A. Head for critical comments on the manuscript. Correspondence and requests for materials should be addressed to J.O.L.
Author information
Authors and Affiliations
Contributions
J.O.L. conceived the original idea. Z.G., S.Y.H., J.W., S.R., W.L., H.P., M.D., W.L., S.B., B.P., B.R.U., Z.T., C.S.-F., V.O.P. and Z.S. designed and conducted the experiments. Z.G., S.Y.H., J.W., S.R., W.L., H.P., M.D., W.L., S.B., B.P., B.R.U., Z.T., C.S.-F., C.-M.T., G.F., I.C., V.O.P., Z.S. and J.O.L. analysed the results. J.O.L., J.W., Z.G., S.Y.H. and V.O.P. co-wrote the manuscript. Z.G., S.Y.H. and J.W. contributed equally to this work. All authors reviewed and provided input into the revision of the manuscript.
Corresponding author
Ethics declarations
Competing interests
Patent applications covering the rapafucin library and rapadocin have been filed by Johns Hopkins University and licensed to Rapafusyn Pharmaceuticals, Inc. J.O.L. is a co-founder of, as well as a Scientific Advisory Board Member for, Rapafusyn Pharmaceuticals, Inc. This arrangement has been reviewed and approved by the Johns Hopkins University in accordance with its conflict of interest policies.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1-16, Supplementary Tables 1-5, and Materials and Methods
Rights and permissions
About this article
Cite this article
Guo, Z., Hong, S.Y., Wang, J. et al. Rapamycin-inspired macrocycles with new target specificity. Nature Chem 11, 254–263 (2019). https://doi.org/10.1038/s41557-018-0187-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-018-0187-4
This article is cited by
-
Concurrent inhibition of oncogenic and wild-type RAS-GTP for cancer therapy
Nature (2024)
-
Design, synthesis and bioactive properties of a class of macrocycles with tunable functional groups and ring size
Scientific Reports (2022)
-
Selection for constrained peptides that bind to a single target protein
Nature Communications (2021)
-
Near quantitative synthesis of urea macrocycles enabled by bulky N-substituent
Nature Communications (2021)
-
A modular biomimetic strategy for the synthesis of macrolide P-glycoprotein inhibitors via Rh-catalyzed C-H activation
Nature Communications (2020)