Abstract
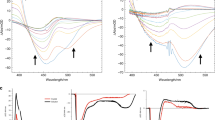
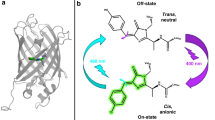
Photochromic fluorescent proteins play key roles in super-resolution microscopy and optogenetics. The light-driven structural changes that modulate the fluorescence involve both trans-to-cis isomerization and proton transfer. The mechanism, timescale and relative contribution of chromophore and protein dynamics are currently not well understood. Here, the mechanism of off-to-on-state switching in dronpa is studied using femtosecond-to-millisecond time-resolved infrared spectroscopy and isotope labelling. Chromophore and protein dynamics are shown to occur on multiple timescales, from picoseconds to hundreds of microseconds. Following excitation of the trans chromophore, a ground-state primary product is formed within picoseconds. Surprisingly, the characteristic vibrational spectrum of the neutral cis isomer appears only after several tens of nanoseconds. Further fluctuations in protein structure around the neutral cis chromophore are required to form a new intermediate, which promotes the final proton-transfer reaction. These data illustrate the interplay between chromophore dynamics and the protein environment underlying fluorescent protein photochromism.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Ando, R., Mizuno, H. & Miyawaki, A. Regulated fast nucleocytoplasmic shuttling observed by reversible protein highlighting. Science 306, 1370–1373 (2004).
Shcherbakova, D. M., Sengupta, P., Lippincott-Schwartz, J. & Verkhusha, V. V. Photocontrollable fluorescent proteins for superresolution imaging. Ann. Rev. Biophys. 43, 303–329 (2014).
Zhou, X. X. & Lin, M. Z. Photoswitchable fluorescent proteins: ten years of colorful chemistry and exciting applications. Curr. Opin. Chem. Biol. 17, 682–690 (2013).
Rodriguez, E. A. The growing and glowing toolbox of fluorescent and photoactive proteins. Trends Biochem. Sci. 42, 111–129 (2017).
Zhou, X. X., Chung, H. K., Lam, A. J. & Lin, M. Z. Optical control of protein activity by fluorescent protein domains. Science 338, 810–814 (2012).
Zhou, X. X., Fan, L. Z., Li, P., Shen, K. & Lin, M. Z. Optical control of cell signaling by single-chain photoswitchable kinases. Science 355, 836 (2017).
Korpany, K. V. et al. Conductance switching in the photoswitchable protein dronpa. J. Am. Chem. Soc. 134, 16119–16122 (2012).
Andresen, M. et al. Structural basis for reversible photoswitching in dronpa. Proc. Natl Acad. Sci. USA 104, 13005–13009 (2007).
Andresen, M. et al. Structure and mechanism of the reversible photoswitch of a fluorescent protein. Proc. Natl Acad. Sci. USA 102, 13070–13074 (2005).
Mizuno, H. et al. Light-dependent regulation of structural flexibility in a photochromic fluorescent protein. Proc. Natl Acad. Sci. USA 105, 9227–9232 (2008).
Fron, E. et al. Ultrafast excited-state dynamics of the photoswitchable protein dronpa. J. Am. Chem. Soc. 129, 4870 (2007).
Warren, M. M. et al. Ground-state proton transfer in the photoswitching reactions of the fluorescent protein dronpa. Nat. Commun. 4, 1461 (2013).
Lukacs, A. et al. Protein photochromism observed by ultrafast vibrational spectroscopy. J. Phys. Chem. B 117, 11954–11959 (2013).
Yadav, D. et al. Real-time monitoring of chromophore isomerization and deprotonation during the photoactivation of the fluorescent protein dronpa. J. Phys. Chem. B 119, 2404–2414 (2015).
Colletier, J.-P. et al. Serial femtosecond crystallography and ultrafast absorption spectroscopy of the photoswitchable fluorescent protein IrisFP. J. Phys. Chem. Lett. 7, 882–887 (2016).
Coquelle, N. et al. Chromophore twisting in the excited state of a photoswitchable fluorescent protein captured by time-resolved serial femtosecond crystallography. Nat. Chem. 10, 31–37 (2018).
Greetham, G. M. Time-resolved multiple probe spectroscopy. Rev. Sci. Instrum. 83, 103107 (2012).
Ando, R., Flors, C., Mizuno, H., Hofkens, J. & Miyawaki, A. Highlighted generation of fluorescence signals using simultaneous two-color irradiation on dronpa mutants. Biophys. J. 92, L97–L99 (2007).
Stoner-Ma, D. et al. Proton relay reaction in green fluorescent protein (GFP): polarization-resolved ultrafast vibrational spectroscopy of isotopically edited GFP. J. Phys. Chem. B 110, 22009–22018 (2006).
van Thor, J. J., Ronayne, K. L., Towrie, M. & Sage, J. T. Balance between ultrafast parallel reactions in the green fluorescent protein has a structural origin. Biophys. J. 95, 1902–1912 (2008).
He, X., Bell, A. F. & Tonge, P. J. Isotopic labeling and normal-mode analysis of a model green fluorescent protein chromophore. J. Phys. Chem. B 106, 6056–6066 (2002).
Snellenburg, J. J., Laptenok, S. P., Seger, R., Mullen, K. M. & van Stokkum, I. H. M. Glotaran: a Java-based graphical user interface for the R package TIMP. J. Stat. Softw. 49, 1–22 (2012).
Barth, A. Infrared spectroscopy of proteins. Biochim. Biophys. Acta Bioenergetics 1767, 1073–1101 (2007).
Kaucikas, M., Tros, M. & van Thor, J. J. Photoisomerization and proton transfer in the forward and reverse photoswitching of the fast-switching M159T mutant of the dronpa fluorescent protein. J. Phys. Chem. B 119, 2350–2362(2015).
Wachter, R. M. Chromogenic cross-link formation in green fluorescent protein. Acc. Chem. Res. 40, 120–127 (2007).
Weber, W., Helms, V., McCammon, J. A. & Langhoff, P. W. Shedding light on the dark and weakly fluorescent states of green fluorescent proteins. Proc. Natl Acad. Sci. USA 96, 6177–6182 (1999).
Conyard, J. et al. A new twist in the photophysics of the GFP chromophore: a volume-conserving molecular torsion couple. Chem. Sci. 9, 1803–1812 (2018).
Voliani, V. et al. Cis–trans photoisomerization of fluorescent-protein chromophores. J. Phys. Chem. B 112, 10714–10722 (2008).
Snyder, J. W., Fales, B. S., Hohenstein, E. G., Levine, B. G. & Martínez, T. J. A direct-compatible formulation of the coupled perturbed complete active space self-consistent field equations on graphical processing units. J. Chem. Phys. 146, 174113 (2017).
Lukacs, A. et al. Photoexcitation of the blue light using FAD photoreceptor AppA results in ultrafast changes to the protein matrix. J. Am. Chem. Soc. 133, 16893–16900 (2011).
Pettersen, E. F. et al. UCSF chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Wallace, A. C., Laskowski, R. A. & Thornton, J. M. LIGPLOT—a program to generate schematic diagrams of protein ligand interactions. Protein Eng. 8, 127–134(1995).
Acknowledgements
S.R.M. acknowledges EPSRC for financial support (EP/N033647/1 and EP/M001997/1). P.J.T. acknowledges NSF for financial support (CHE-1223819). A.M. acknowledges the Japan Ministry of Education, Culture, Sports, Science and Technology Grant-in-aid for Scientific research on Innovative Areas: Resonance Bio. The authors acknowledge STFC for access to the Central Laser Facility. Calculations were performed on the High Performance Computing Cluster at the University of East Anglia.
Author information
Authors and Affiliations
Contributions
S.P.L., A.A.G., C.R.H., A.L. and J.N.I. measured and collected the data. S.P.L. and C.R.H. analysed the data. A.A.G. and J.N.I. grew and purified the samples. G.A.J. performed the DFT calculations. G.M.G. and P.D. built, developed and managed the ULTRA and LifeTime apparatus used in the measurements. A.M. designed the dronpa2 protein. P.J.T. and S.R.M. designed the experiment and wrote the paper, with discussion and editorial input from all authors
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary information
Supplementary Figures 1–9, Supplementary Tables 1–5, Supplementary Methods, Supplementary Data, Supplementary Analysis
Rights and permissions
About this article
Cite this article
Laptenok, S.P., Gil, A.A., Hall, C.R. et al. Infrared spectroscopy reveals multi-step multi-timescale photoactivation in the photoconvertible protein archetype dronpa. Nature Chem 10, 845–852 (2018). https://doi.org/10.1038/s41557-018-0073-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-018-0073-0
This article is cited by
-
Optical control of ultrafast structural dynamics in a fluorescent protein
Nature Chemistry (2023)
-
Direct structural observation of ultrafast photoisomerization dynamics in sinapate esters
Communications Chemistry (2022)
-
Large Stokes shift fluorescence activation in an RNA aptamer by intermolecular proton transfer to guanine
Nature Communications (2021)
-
Photoswitching mechanism of a fluorescent protein revealed by time-resolved crystallography and transient absorption spectroscopy
Nature Communications (2020)
-
Picosecond to millisecond tracking of a photocatalytic decarboxylation reaction provides direct mechanistic insights
Nature Communications (2019)