Abstract
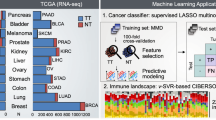
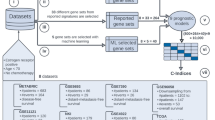
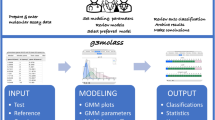
Despite its early promise as a diagnostic and prognostic tool, gene expression profiling remains cost-prohibitive and challenging to implement in a clinical setting. Here, we introduce a molecular computation strategy for analysing the information contained in complex gene expression signatures without the need for costly instrumentation. Our workflow begins by training a computational classifier on labelled gene expression data. This in silico classifier is then realized at the molecular level to enable expression analysis and classification of previously uncharacterized samples. Classification occurs through a series of molecular interactions between RNA inputs and engineered DNA probes designed to differentially weigh each input according to its importance. We validate our technology with two applications: a classifier for early cancer diagnostics and a classifier for differentiating viral and bacterial respiratory infections based on host gene expression. Together, our results demonstrate a general and modular framework for low-cost gene expression analysis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Vargas, J. D & Lima, J. A. C. A gene-expression score to predict obstructive CAD. Nat. Rev. Cardiol. 10(5), 243–244 2013).
van’t Veer, L. J. et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature 415, 530–536 (2002).
Blank, P. R. et al. Cost-effectiveness analysis of prognostic gene expression signature-based stratification of early breast cancer patients. Pharmacoeconomics 33, 179–190 (2015).
Myers, M. B. Targeted therapies with companion diagnostics in the management of breast cancer: current perspectives. Pharmgenomics Pers. Med. 9, 7–16 (2016).
Rotunno, M. et al. A gene expression signature from peripheral whole blood for stage I lung adenocarcinoma. Cancer Prev. Res 4, 1599–1608 (2011).
Lunnon, K., Sattlecker, M. & Furney, S. J. A blood gene expression marker of early Alzheimer’s disease. J. Alzheimers Dis. 33, 737–753 (2013).
Koscielny, S. Why most gene expression signatures of tumors have not been useful in the clinic. Sci. Transl. Med. 2, 14ps2 (2010).
Sotiriou, C. & Piccart, M. J. Taking gene-expression profiling to the clinic: when will molecular signatures become relevant to patient care? Nat. Rev. Cancer 7, 545–553 (2007).
Cassarino, D. S., Lewine, N., Cole, D. & Wade, B. Budget impact analysis of a novel gene expression assay for the diagnosis of malignant melanoma. J. Med. Econ. 17, 782–791 (2014).
Tsalik, E. L. et al. Host gene expression classifiers diagnose respiratory illness etiology. Sci. Transl. Med. 8, 322ra11 (2016).
Best, M. G. et al. RNA-seq of tumor-educated platelets enables blood-based pan-cancer, multiclass, and molecular pathway cancer diagnostics. Cancer Cell 28, 666–676 (2015).
Yuan, T. et al. Plasma extracellular RNA profiles in healthy and cancer patients. Sci. Rep. 6, 19413 (2016).
Dasí, F. et al. Real-time quantification in plasma of human telomerase reverse transcriptase (hTERT) mRNA: a simple blood test to monitor disease in cancer patients. Lab. Invest. 81, 767–769 (2001).
Zhang, L. et al. Salivary transcriptomic biomarkers for detection of resectable pancreatic cancer. Gastroenterology 138, 949 (2009).
Zhang, L. et al. Development of transcriptomic biomarker signature in human saliva to detect lung cancer. Cell Mol. Life Sci. 69, 3341–3350 (2012).
Kyo, S., Takakura, M., Fujiwara, T. & Inoue, M. Understanding and exploiting hTERT promoter regulation for diagnosis and treatment of human cancers. Cancer Sci. 99, 1528–1538 (2008).
Lledo et al. Real time quantification in plasma of human telomerase reverse transcriptase (hTERT) mRNA in patients with colorectal cancer. Colorectal Dis. 6, 236–242 (2004).
March-Villalba, J. A. et al. Cell-free circulating plasma hTERT mRNA is a useful marker for prostate cancer diagnosis and is associated with poor prognosis tumor characteristics. PLoS ONE 7, e43470 (2012).
Miura, N., Nakamura, H., Sato, R. & Tsukamoto, T. Clinical usefulness of serum telomerase reverse transcriptase (hTERT) mRNA and epidermal growth factor receptor (EGFR) mRNA as a novel tumor marker. Cancer Sci. 97, 1366–1373 (2006).
Terrin, L. et al. Relationship between tumor and plasma levels of hTERT mRNA in patients with colorectal cancer: implications for monitoring of neoplastic disease. Clin. Cancer Res. 14, 7444–7451 (2008).
Ramilo, O., Allman, W., Chung, W., Mejias, A. & Ardura, M. Gene expression patterns in blood leukocytes discriminate patients with acute infections. Blood 109, 2066–2077 (2007).
Chen, S. X. & Seelig, G. An engineered kinetic amplification mechanism for single nucleotide variant discrimination by DNA hybridization probes. J. Am. Chem. Soc. 138, 5076–5086 (2016).
Pardee, K., Green, A. A., Ferrante, T. & Cameron, D. E. Paper-based synthetic gene networks. Cell 159, 940–954 (2014).
Pardee, K. et al. Rapid, low-cost detection of Zika virus using programmable biomolecular components. Cell 165, 1255–1266 (2016).
Jung, C. & Ellington, A. D. Diagnostic applications of nucleic acid circuits. Acc. Chem. Res 47, 1825–1835 (2014).
Gootenberg, J. S. et al. Nucleic acid detection with CRISPR-Cas13a/C2c2. Science 356, 438–442 (2017).
Qian, L. & Winfree, E. Scaling up digital circuit computation with DNA strand displacement cascades. Science 332, 1196–1201 (2011).
Qian, L., Winfree, E. & Bruck, J. Neural network computation with DNA strand displacement cascades. Nature 475, 368–372 (2011).
Seelig, G., Soloveichik, D., Zhang, D. & Winfree, E. Enzyme-free nucleic acid logic circuits. Science 314, 1585–1588 (2006).
Chen, Y.-J. et al. Programmable chemical controllers made from DNA. Nat. Nanotech. 8, 755–762 (2013).
Genot, A. J., Fujii, T. & Rondelez, Y. Scaling down DNA circuits with competitive neural networks. J. R. Soc. Interface 10, 20130212 (2013).
Franco, E. et al. Timing molecular motion and production with a synthetic transcriptional clock. Proc. Natl Acad. Sci. USA 108, E784–E793 (2011).
Mills, A. P. Gene expression profiling diagnosis through DNA molecular computation. Trends Biotechnol. 20, 137–140 (2002).
Green, A. A. et al. Complex cellular logic computation using ribocomputing devices. Nature 548, 117–121 (2017).
Brown, M. P. S., Grundy, W. N. & Lin, D. Knowledge-based analysis of microarray gene expression data by using support vector machines. Proc. Natl Acad. Sci. USA 97, 262–267 (2000).
Abusamra, H. A comparative study of feature selection and classification methods for gene expression data of glioma. Procedia Comput. Sci. 23, 5–14 (2013).
Liu, H., Li, J. & Wong, L. A comparative study on feature selection and classification methods using gene expression profiles and proteomic patterns. Genome Inform. 13, 51–60 (2002).
Shelton, V. M., Sosnick, T. R. & Pan, T. Applicability of urea in the thermodynamic analysis of secondary and tertiary RNA folding. Biochemistry 38, 16831–16839 (1999).
Zhang, D. Y. & Seelig, G. in DNA Computing and Molecular Programming (eds Sakakibara, Y. & Mi, Y.) Vol. 6518, 176–186 (Springer, Berlin, 2010).
Zhang, D. Cooperative hybridization of oligonucleotides. J. Am. Chem. Soc. 133, 1077–1086 (2011).
Dasí, F. et al. Real-time quantification of human telomerase reverse transcriptase mRNA in the plasma of patients with prostate cancer. Ann. NY Acad. Sci. 1075, 204–210 (2006).
Yang, Y. J., Chen, H., Huang, P., Li, C. H. & Dong, Z. Quantification of plasma hTERT DNA in hepatocellular carcinoma patients by quantitative fluorescent polymerase chain reaction. Clin. Invest. 34, E238 (2011).
Lizardi, P. M., Huang, X., Zhu, Z. & Bray-Ward, P. Mutation detection and single-molecule counting using isothermal rolling-circle amplification. Nat. Genet 19, 225–232 (1998).
Zhao, W., Ali, M. M., Brook, M. A. & Li, Y. Rolling circle amplification: applications in nanotechnology and biodetection with functional nucleic acids. Angew. Chem. Int Ed. 47, 6330–6337 (2008).
Notomi, T., Okayama, H. & Masubuchi, H. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 28, e63 (2000).
Tomita, N., Mori, Y., Kanda, H. & Notomi, T. Loop-mediated isothermal amplification (LAMP) of gene sequences and simple visual detection of products. Nat. Protoc. 3, 877–882 (2008).
Acknowledgements
The authors thank Y.-J. Chen, S. Chen, G. Chatterjee and D.Y. Zhang for their support and discussions. This work was supported by NSF grants CCF-171449 and CCF-1317653.
Author information
Authors and Affiliations
Contributions
R.L.B. and G.S. designed the experiments and wrote the paper. R.L.B. and R.W. performed the experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Text, Supplementary Figures, and Supplementary Tables
Rights and permissions
About this article
Cite this article
Lopez, R., Wang, R. & Seelig, G. A molecular multi-gene classifier for disease diagnostics. Nature Chem 10, 746–754 (2018). https://doi.org/10.1038/s41557-018-0056-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-018-0056-1
This article is cited by
-
DNA-based molecular classifiers for the profiling of gene expression signatures
Journal of Nanobiotechnology (2024)
-
DNA as a universal chemical substrate for computing and data storage
Nature Reviews Chemistry (2024)
-
Employing toehold-mediated DNA strand displacement reactions for biomedical applications
Med-X (2024)
-
DNA-framework-based multidimensional molecular classifiers for cancer diagnosis
Nature Nanotechnology (2023)
-
Nano scale instance-based learning using non-specific hybridization of DNA sequences
Communications Engineering (2023)