Abstract
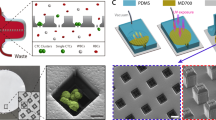
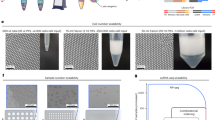
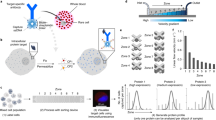
Cell-to-cell variation in gene expression creates a need for techniques that can characterize expression at the level of individual cells. This is particularly true for rare circulating tumour cells, in which subtyping and drug resistance are of intense interest. Here we describe a method for cell analysis—single-cell mRNA cytometry—that enables the isolation of rare cells from whole blood as a function of target mRNA sequences. This approach uses two classes of magnetic particles that are labelled to selectively hybridize with different regions of the target mRNA. Hybridization leads to the formation of large magnetic clusters that remain localized within the cells of interest, thereby enabling the cells to be magnetically separated. Targeting specific intracellular mRNAs enablescirculating tumour cells to be distinguished from normal haematopoietic cells. No polymerase chain reaction amplification is required to determine RNA expression levels and genotype at the single-cell level, and minimal cell manipulation is required. To demonstrate this approach we use single-cell mRNA cytometry to detect clinically important sequences in prostate cancer specimens.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Elowitz, M. B., Levine, A. J., Siggia, E. D. & Swain, P. S. Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002).
Bendall, S. C. & Nolan, G. P. From single cells to deep phenotypes in cancer. Nat. Biotechnol. 30, 639–647 (2012).
Yu, M. et al. Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science 339, 580–584 (2013).
Kalinich, M. et al. An RNA-based signature enables high specificity detection of circulating tumor cells in hepatocellular carcinoma. Proc. Natl Acad. Sci. USA 114, 1123–1128 (2017).
Clark, I. C. & Abate, A. R. Finding a helix in a haystack: nucleic acid cytometry with droplet microfluidics. Lab Chip 17, 2032–2045 (2017).
Briley, W. E., Bondy, M. H., Randeria, P. S., Dupper, T. J. & Mirkin, C. A. Quantification and real-time tracking of RNA in live cells using Sticky-flares. Proc. Natl Acad. Sci. USA 112, 9591–95955 (2015).
Geiss, G. K. et al. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nat. Biotechnol. 26, 317–325 (2008).
Deng, Q., Ramskold, D., Reinius, B. & Sandberg, R. Single-cell RNA-seq reveals dynamic, random monoallelic gene expression in mammalian cells. Science 343, 193–196 (2014).
Livak, K. J. et al. Methods for qPCR gene expression profiling applied to 1440 lymphoblastoid single cells. Methods 59, 71–79 (2013).
Lyubimova, A. et al. Single-molecule mRNA detection and counting in mammalian tissue. Nat. Protocol. 8, 1743–1758 (2013).
Itzkovitz, S. & van Oudenaarden, A. Validating transcripts with probes and imaging technology. Nat. Methods 8, S12–S19 (2011).
Halo, T. L. et al. NanoFlares for the detection, isolation, and culture of live tumor cells from human blood. Proc. Natl Acad. Sci. USA 111, 17104–17109 (2014).
Alix-Panabieres, C. & Pantel, K. Challenges in circulating tumour cell research. Nat. Rev. Cancer 14, 623–631 (2014).
Lang, J. M., Casavant, B. P. & Beebe, D. J. Circulating tumor cells: getting more from less. Sci. Transl. Med. 4, 141ps113 (2012).
Green, B. J. et al. Beyond the capture of circulating tumor cells: next-generation devices and materials. Angew. Chem. Int. Ed. 55, 1252–1265 (2016).
Zhang, J., Chen, K. & Fan, Z. H. Circulating tumor cell isolation and analysis. Adv. Clin. Chem. 75, 1–31 (2016).
Zhang, J., Sheng, W. & Fan, Z. H. An ensemble of aptamers and antibodies for multivalent capture of cancer cells. Chem. Commun. 50, 6722–6725 (2014).
Hu, X. et al. Marker-specific sorting of rare cells using dielectrophoresis. Proc. Natl Acad. Sci. USA 102, 15757–15761 (2005).
Nagrath, S. et al. Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nature 450, 1235–1239 (2007).
Adams, A. A. et al. Highly efficient circulating tumor cell isolation from whole blood and label-free enumeration using polymer-based microfluidics with an integrated conductivity sensor. J. Am. Chem. Soc. 130, 8633–8641 (2008).
Talasaz, A. H. et al. Isolating highly enriched populations of circulating epithelial cells and other rare cells from blood using a magnetic sweeper device. Proc. Natl Acad. Sci. USA 106, 3970–3975 (2009).
Stott, S. L. et al. Isolation of circulating tumor cells using a microvortex-generating herringbone-chip. Proc. Natl Acad. Sci. USA 107, 18392–18397 (2010).
Schiro, P. G. et al. Sensitive and high-throughput isolation of rare cells from peripheral blood with ensemble-decision aliquot ranking. Angew. Chem. Int. Ed. 51, 4618–4622 (2012).
Zhao, W. et al. Bioinspired multivalent DNA network for capture and release of cells. Proc. Natl Acad. Sci. USA 109, 19626–19631 (2012).
Ozkumur, E. et al. Inertial focusing for tumor antigen-dependent and -independent sorting of rare circulating tumor cells. Sci. Transl. Med. 5, 179ra147 (2013).
Zhang, Y., Wu, M., Han, X., Wang, P. & Qin, L. High-throughput, label-free isolation of cancer stem cells on the basis of cell adhesion capacity. Angew. Chem. Int. Ed. 54, 10838–10842 (2015).
Zhang, Y., Zhou, L. & Qin, L. High-throughput 3D cell invasion chip enables accurate cancer metastatic assays. J. Am. Chem. Soc. 136, 15257–15262 (2014).
Yoon, H. J. et al. Sensitive capture of circulating tumour cells by functionalized graphene oxide nanosheets. Nat. Nanotechnol. 8, 735–741 (2013).
Poudineh, M. et al. Tracking the dynamics of circulating tumour cell phenotypes using nanoparticle-mediated magnetic ranking. Nat. Nanotechnol. 12, 274–281 (2017).
Altieri, D. C. Validating survivin as a cancer therapeutic target. Nat. Rev. Cancer 3, 46–54 (2003).
Fulda, S. & Vucic, D. Targeting IAP proteins for therapeutic intervention in cancer. Nat. Rev. Drug Discov. 11, 109–124 (2012).
Carrasco, R. A. et al. Antisense inhibition of survivin expression as a cancer therapeutic. Mol. Cancer Ther. 10, 221–232 (2011).
Watson, P. A., Arora, V. K. & Sawyers, C. L. Emerging mechanisms of resistance to androgen receptor inhibitors in prostate cancer. Nat. Rev. Cancer 15, 701–711 (2015).
Robinson, D. et al. Integrative clinical genomics of advanced prostate cancer. Cell 161, 1215–1228 (2015).
Antonarakis, E. S. et al. AR-V7 and resistance to enzalutamide and abiraterone in prostate cancer. N. Engl. J. Med. 371, 1028–1038 (2014).
Tomlins, S. A. et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science 310, 644–648 (2005).
Tomlins, S. A. et al. Urine TMPRSS2:ERG fusion transcript stratifies prostate cancer risk in men with elevated serum PSA. Sci. Transl. Med. 3, 94ra72 (2011).
Edwardson, T. G., Lau, K. L., Bousmail, D., Serpell, C. J. & Sleiman, H. F. Transfer of molecular recognition information from DNA nanostructures to gold nanoparticles. Nat. Chem. 8, 162–170 (2016).
Park, S. Y. et al. DNA-programmable nanoparticle crystallization. Nature 451, 553–556 (2008).
Aldaye, F. A. & Sleiman, H. F. Dynamic DNA templates for discrete gold nanoparticle assemblies: control of geometry, modularity, write/erase and structural switching. J. Am. Chem. Soc. 129, 4130–4131 (2007).
Chuah, K. et al. Ultrasensitive electrochemical detection of prostate-specific antigen (PSA) using gold-coated magnetic nanoparticles as ‘dispersible electrodes’. Chem. Commun. 48, 3503–3505 (2012).
Farlow, J. et al. Formation of targeted monovalent quantum dots by steric exclusion. Nat. Methods 10, 1203–1205 (2013).
Acknowledgements
Research reported in this publication was supported by the Canadian Institutes of Health Research (grant no. FDN-148415), the Natural Sciences and Engineering Research Council of Canada (grant no. 2016-06090), the Province of Ontario through the Ministry of Research, Innovation and Science (grant no. RE05-009), and the National Cancer Institute of the National Institutes of Health (grant no. 1R33CA204574). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health or the other funding agencies. We also thank A. Joshua at the Princess Margaret Hospital for supplying clinical specimens.
Author information
Authors and Affiliations
Contributions
M.L., S.O.K. and E.H.S. conceived and designed the experiments; M.L., R.M.M., M.P., S.U.A., I.I., C.-L.H. and M.M. performed the experiments and analysed the data. All authors discussed the results and contributed to the preparation and editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods, Supplementary Tables, Supplementary Figs. and Supplementary Simulations
Rights and permissions
About this article
Cite this article
Labib, M., Mohamadi, R.M., Poudineh, M. et al. Single-cell mRNA cytometry via sequence-specific nanoparticle clustering and trapping. Nature Chem 10, 489–495 (2018). https://doi.org/10.1038/s41557-018-0025-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-018-0025-8
This article is cited by
-
Identification of druggable regulators of cell secretion via a kinome-wide screen and high-throughput immunomagnetic cell sorting
Nature Biomedical Engineering (2023)
-
A magneto-activated nanoscale cytometry platform for molecular profiling of small extracellular vesicles
Nature Communications (2023)
-
Multiplexed RNA profiling by regenerative catalysis enables blood-based subtyping of brain tumors
Nature Communications (2023)
-
A high-dimensional microfluidic approach for selection of aptamers with programmable binding affinities
Nature Chemistry (2023)
-
Manipulation of single cells via a Stereo Acoustic Streaming Tunnel (SteAST)
Microsystems & Nanoengineering (2022)