Abstract
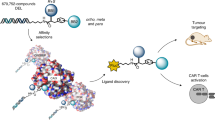
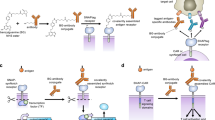
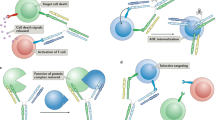
In nature, specific antibodies can be generated as a result of an adaptive selection and expansion of lymphocytes with suitable protein binding properties. We attempted to mimic antibody–antigen recognition by displaying multiple chemical diversity elements on a defined macrocyclic scaffold. Encoding of the displayed combinations was achieved using distinctive DNA tags, resulting in a library size of 35,393,112. Specific binders could be isolated against a variety of proteins, including carbonic anhydrase IX, horseradish peroxidase, tankyrase 1, human serum albumin, alpha-1 acid glycoprotein, calmodulin, prostate-specific antigen and tumour necrosis factor. Similar to antibodies, the encoded display of multiple chemical elements on a constant scaffold enabled practical applications, such as fluorescence microscopy procedures or the selective in vivo delivery of payloads to tumours. Furthermore, the versatile structure of the scaffold facilitated the generation of protein-specific chemical probes, as illustrated by photo-crosslinking.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Murphy, K., Janeway, C. A. & Mowat, A. Janeway’s Immunobiology 8th edn (Garland Science, New York, 2012).
Smith, G. P. Filamentous fusion phage: novel expression vectors that display cloned antigens on the virion surface. Science 228, 1315–1317 (1985).
Clackson, T., Hoogenboom, H. R., Griffiths, A. D. & Winter, G. Making antibody fragments using phage display libraries. Nature 352, 624–628 (1991).
Lerner, R. A., Kang, A. S., Bain, J. D., Burton, D. R. & Barbas, C. F. III Antibodies without immunization. Science 258, 1313–1314 (1992).
Winter, G., Griffiths, A. D., Hawkins, R. E. & Hoogenboom, H. R. Making antibodies by phage display technology. Annu. Rev. Immunol. 12, 433–455 (1994).
Thurber, G. M. et al. Single-cell and subcellular pharmacokinetic imaging allows insight into drug action in vivo. Nat. Commun. 4, 1504 (2013).
Heinis, C., Rutherford, T., Freund, S. & Winter, G. Phage-encoded combinatorial chemical libraries based on bicyclic peptides. Nat. Chem. Biol. 5, 502–507 (2009).
Wilson, D. S., Keefe, A. D. & Szostak, J. W. The use of mRNA display to select high-affinity protein-binding peptides. Proc. Natl Acad. Sci. USA 98, 3750–3755 (2001).
Kawakami, T. et al. Diverse backbone-cyclized peptides via codon reprogramming. Nat. Chem. Biol. 5, 888–890 (2009).
Boder, E. T. & Wittrup, K. D. Yeast surface display for screening combinatorial polypeptide libraries. Nat. Biotechnol. 15, 553–557 (1997).
Hanes, J. & Pluckthun, A. In vitro selection and evolution of functional proteins by using ribosome display. Proc. Natl Acad. Sci. USA 94, 4937–4942 (1997).
Goodnow, R. A. Jr, Dumelin, C. E. & Keefe, A. D. DNA-encoded chemistry: enabling the deeper sampling of chemical space. Nat. Rev. Drug Discov. 16, 131–147 (2017).
Lerner, R. A. & Brenner, S. DNA-encoded compound libraries as open source: a powerful pathway to new drugs. Angew. Chem. Int. Ed. 56, 1164–1165 (2017).
Brenner, S. & Lerner, R. A. Encoded combinatorial chemistry. Proc. Natl Acad. Sci. USA 89, 5381–5383 (1992).
Needels, M. C. et al. Generation and screening of an oligonucleotide-encoded synthetic peptide library. Proc. Natl Acad. Sci. USA 90, 10700–10704 (1993).
Mannocci, L. et al. High-throughput sequencing allows the identification of binding molecules isolated from DNA-encoded chemical libraries. Proc. Natl. Acad. Sci. USA 105, 17670–17675 (2008).
Clark, M. A. et al. Design, synthesis and selection of DNA-encoded small-molecule libraries. Nat. Chem. Biol. 5, 647–654 (2009).
Litovchick, A. et al. Encoded library synthesis using chemical ligation and the discovery of sEH inhibitors from a 334-million member library. Sci. Rep. 5, 10916 (2015).
Franzini, R. M. & Randolph, C. Chemical space of DNA-encoded libraries. J. Med. Chem. 59, 6629–6644 (2016).
Halpin, D. R. & Harbury, P. B. DNA display I. Sequence-encoded routing of DNA populations. PLoS Biol. 2, e173 (2004).
Gartner, Z. J. et al. DNA-templated organic synthesis and selection of a library of macrocycles. Science 305, 1601–1605 (2004).
Hansen, M. H. et al. A yoctoliter-scale DNA reactor for small-molecule evolution. J. Am. Chem. Soc. 131, 1322–1327 (2009).
Li, Y., Zhao, P., Zhang, M., Zhao, X. & Li, X. Multistep DNA-templated synthesis using a universal template. J. Am. Chem. Soc. 135, 17727–17730 (2013).
Wichert, M. et al. Dual-display of small molecules enables the discovery of ligand pairs and facilitates affinity maturation. Nat. Chem. 7, 241–249 (2015).
Reddavide, F. V., Lin, W., Lehnert, S. & Zhang, Y. DNA-encoded dynamic combinatorial chemical libraries. Angew. Chem. Int. Ed. 54, 7924–7928 (2015).
Li, G. et al. Design, preparation, and selection of DNA-encoded dynamic libraries. Chem. Sci. 6, 7097–7104 (2015).
Daguer, J. P. et al. DNA display of fragment pairs as a tool for the discovery of novel biologically active small molecules. Chem. Sci. 6, 739–744 (2015).
Deng, H. et al. Discovery of highly potent and selective small molecule ADAMTS-5 inhibitors that inhibit human cartilage degradation via encoded library technology (ELT). J. Med. Chem. 55, 7061–7079 (2012).
Barluenga, S. et al. Novel PTP1B inhibitors identified by DNA display of fragment pairs. Bioorg. Med. Chem. Lett. 26, 1080–1085 (2016).
Maianti, J. P. et al. Anti-diabetic activity of insulin-degrading enzyme inhibitors mediated by multiple hormones. Nature 511, 94–98 (2014).
Mutter, M., Hersperger, R., Gubernator, K. & Muller, K. The construction of new proteins. V. A template-assembled synthetic protein (TASP) containing both a 4-helix bundle and beta-barrel-like structure. Proteins 5, 13–21 (1989).
Dumy, P., Eggleston, I. M., Esposito, G., Nicula, S. & Mutter, M. Solution structure of regioselectively addressable functionalized templates: an NMR and restrained molecular dynamics investigation. Biopolymers 39, 297–308 (1996).
Peluso, S. et al. Crystal structure of a synthetic cyclodecapeptide for template-assembled synthetic protein design. ChemBioChem 2, 432–437 (2001).
Salamon, H., Klika Škopić, M., Jung, K., Bugain, O. & Brunschweiger, A. Chemical biology probes from advanced DNA-encoded libraries. ACS Chem. Biol. 11, 296–307 (2016).
Franzini, R. M. et al. Systematic evaluation and optimization of modification reactions of oligonucleotides with amines and carboxylic acids for the synthesis of DNA-encoded chemical libraries. Bioconjug. Chem. 25, 1453–1461 (2014).
Li, Y. et al. Optimized reaction conditions for amide bond formation in DNA-encoded combinatorial libraries. ACS Comb. Sci. 18, 438–443 (2016).
Decurtins, W. et al. Automated screening for small organic ligands using DNA-encoded chemical libraries. Nat. Protoc. 11, 764–780 (2016).
Krall, N. et al. A small-molecule drug conjugate for the treatment of carbonic anhydrase IX expressing tumors. Angew. Chem. Int. Ed. 53, 4231–4235 (2014).
Gilabert, M. A. A. et al. Differential substrate behaviour of phenol and aniline derivatives during oxidation by horseradish peroxidase: kinetic evidence for a two-step mechanism. Biochim. Biophys. Acta 1699, 235–243 (2004).
Franzini, R. M. et al. Identification of structure–activity relationships from screening a structurally compact DNA-encoded chemical library. Angew. Chem. Int. Ed. 54, 3927–3931 (2015).
Malone, M. L. & Paegel, B. M. What is a ‘DNA-compatible’ reaction? ACS Comb. Sci. 18, (182–187 (2016).
Deng, H. et al. Discovery, SAR, and X-ray binding mode study of BCATm inhibitors from a novel DNA-encoded library. ACS Med. Chem. Lett. 6, 919–924 (2015).
Satz, A. L. Simulated screens of DNA encoded libraries: the potential influence of chemical synthesis fidelity on interpretation of structure–activity relationships. ACS Comb. Sci. 18, 415–424 (2016).
Griffiths, A. D. et al. Isolation of high affinity human antibodies directly from large synthetic repertoires. EMBO J. 13, 3245–3260 (1994).
Nissim, A. et al. Antibody fragments from a ‘single pot’ phage display library as immunochemical reagents. EMBO J. 13, 692–698 (1994).
Vaughan, T. J. et al. Human antibodies with sub-nanomolar affinities isolated from a large non-immunized phage display library. Nat. Biotechnol. 14, 309–314 (1996).
Gram, H. et al. In vitro selection and affinity maturation of antibodies from a naive combinatorial immunoglobulin library. Proc. Natl. Acad. Sci. USA 89, 3576–3580 (1992).
Jespers, L. S., Roberts, A., Mahler, S. M., Winter, G. & Hoogenboom, H. R. Guiding the selection of human antibodies from phage display repertoires to a single epitope of an antigen. Biotechnology 12, 899–903 (1994).
Pini, A. et al. Design and use of a phage display library. Human antibodies with subnanomolar affinity against a marker of angiogenesis eluted from a two-dimensional gel. J. Biol. Chem. 273, 21769–21776 (1998).
Fellouse, F. A., Wiesmann, C. & Sidhu, S. S. Synthetic antibodies from a four-amino-acid code: a dominant role for tyrosine in antigen recognition. Proc. Natl. Acad. Sci. USA 101, 12467–12472 (2004).
Li, B. et al. Minimization of a polypeptide hormone. Science 270, 1657–1660 (1995).
Driggers, E. M., Hale, S. P., Lee, J. & Terrett, N. K. The exploration of macrocycles for drug discovery—an underexploited structural class. Nat. Rev. Drug Discov. 7, 608–624 (2008).
Acknowledgements
This work was supported by ETH Zürich, the Swiss National Science Foundation (310030B_163479/1 grant, 31003A_144242 grant and CRSII2_160699/1 Sinergia grant), the ERC Advanced Grant ‘Zauberkugel’ (Grant agreement 670603), Philochem AG and through Start-up Funding for the ‘Hundred Young-Talent Scheme’ Professorship provided by Chongqing University in China (0247001104406). The authors thank G. Annuziato for help with synthesis, C. Aquino and L. Optiz for help with high-throughput DNA sequencing, and M. Wichert, R. Franzini, G. Zimmerman, A. Dal Corso, W. Decurtins, A. Schmid, L. Sun, M. Xi and J. Zhang for discussions. Instant JChem (ChemAxon) was used for structure and data management (http://www.chemaxon.com).
Author information
Authors and Affiliations
Contributions
Y.L., J.S. and D.N. designed the project. Y.L. constructed the library. R.D.L. provided target proteins. Y.L. designed and performed the selections. Y.L. and J.S. analysed high-throughput DNA screening data. Y.L. performed synthesis and hit validation experiments and performed the photo-crosslinking experiments. Y.L. and F.P. performed the immunofluorescence experiments. Y.L. and S.C. performed in vivo experiments. D.B. performed the biotinylation of target proteins. Y.L., J.S. and D.N. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
D.N. is a co-founder and shareholder of Philochem AG (Otelfingen, Switerland) and J.S. is a board member of Philochem AG.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods, Supplementary Data, and Supplementary Figures 1–77
Rights and permissions
About this article
Cite this article
Li, Y., De Luca, R., Cazzamalli, S. et al. Versatile protein recognition by the encoded display of multiple chemical elements on a constant macrocyclic scaffold. Nature Chem 10, 441–448 (2018). https://doi.org/10.1038/s41557-018-0017-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-018-0017-8
This article is cited by
-
Trio-pharmacophore DNA-encoded chemical library for simultaneous selection of fragments and linkers
Nature Communications (2023)
-
A DNA-encoded chemical library based on chiral 4-amino-proline enables stereospecific isozyme-selective protein recognition
Nature Chemistry (2023)
-
Genetically encoded discovery of perfluoroaryl macrocycles that bind to albumin and exhibit extended circulation in vivo
Nature Communications (2023)
-
Synthesis and direct assay of large macrocycle diversities by combinatorial late-stage modification at picomole scale
Nature Communications (2022)
-
A mating mechanism to generate diversity for the Darwinian selection of DNA-encoded synthetic molecules
Nature Chemistry (2022)