Abstract
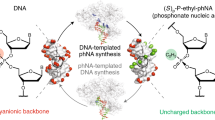
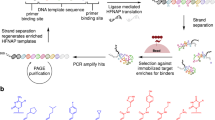
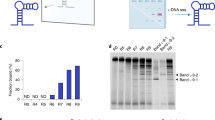
The evolution of sequence-defined synthetic polymers made of building blocks beyond those compatible with polymerase enzymes or the ribosome has the potential to generate new classes of receptors, catalysts and materials. Here we describe a ligase-mediated DNA-templated polymerization and in vitro selection system to evolve highly functionalized nucleic acid polymers (HFNAPs) made from 32 building blocks that contain eight chemically diverse side chains on a DNA backbone. Through iterated cycles of polymer translation, selection and reverse translation, we discovered HFNAPs that bind proprotein convertase subtilisin/kexin type 9 (PCSK9) and interleukin-6, two protein targets implicated in human diseases. Mutation and reselection of an active PCSK9-binding polymer yielded evolved polymers with high affinity (KD = 3 nM). This evolved polymer potently inhibited the binding between PCSK9 and the low-density lipoprotein receptor. Structure–activity relationship studies revealed that specific side chains at defined positions in the polymers are required for binding to their respective targets. Our findings expand the chemical space of evolvable polymers to include densely functionalized nucleic acids with diverse, researcher-defined chemical repertoires.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ellington, A. D. & Szostak, J. W. In vitro selection of RNA molecules that bind specific ligands. Nature 346, 818–822 (1990).
Tuerk, C. & Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249, 505–510 (1990).
Robertson, D. L. & Joyce, G. F. Selection in vitro of an RNA enzyme that specifically cleaves single-stranded DNA. Nature 344, 467–468 (1990).
Bock, L. C., Griffin, L. C., Latham, J. A., Vermaas, E. H. & Toole, J. J. Selection of single-stranded DNA molecules that bind and inhibit human thrombin. Nature 355, 564–566 (1992).
Ellington, A. D. & Szostak, J. W. Selection in vitro of single-stranded DNA molecules that fold into specific ligand-binding structures. Nature 355, 850–852 (1992).
Breaker, R. R. & Joyce, G. F. A DNA enzyme that cleaves RNA. Chem. Biol. 1, 223–229 (1994).
Mattheakis, L. C., Bhatt, R. R. & Dower, W. J. An in vitro polysome display system for identifying ligands from very large peptide libraries. Proc. Natl Acad. Sci. USA 91, 9022–9026 (1994).
Roberts, R. W. & Szostak, J. W. RNA-peptide fusions for the in vitro selection of peptides and proteins. Proc. Natl Acad. Sci. USA 94, 12297–12302 (1997).
Nemoto, N., Miyamoto-Sato, E., Husimi, Y. & Yanagawa, H. In vitro virus: bonding of mRNA bearing puromycin at the 3′-terminal end to the C-terminal end of its encoded protein on the ribosome in vitro. FEBS Lett. 414, 405–408 (1997).
Yamaguchi, J. et al. cDNA display: a novel screening method for functional disulfide-rich peptides by solid-phase synthesis and stabilization of mRNA-protein fusions. Nucleic Acids Res. 37, e108 (2009).
Brudno, Y. & Liu, D. R. Recent progress toward the templated synthesis and directed evolution of sequence-defined synthetic polymers. Chem. Biol. 16, 265–276 (2009).
Chaput, J. C., Yu, H. & Zhang, S. The emerging world of synthetic genetics. Chem. Biol. 19, 1360–1371 (2012).
Pinheiro, V. B. & Holliger, P. The XNA world: progress towards replication and evolution of synthetic genetic polymers. Curr. Opin. Chem. Biol. 16, 245–252 (2012).
Pinheiro, V. B. & Holliger, P. Towards XNA nanotechnology: new materials from synthetic genetic polymers. Trends Biotechnol. 32, 321–328 (2014).
Hollenstein, M. Nucleoside triphosphates—building blocks for the modification of nucleic acids. Molecules 17, 13569–13591 (2012).
Rogers, J. M. & Suga, H. Discovering functional, non-proteinogenic amino acid containing, peptides using genetic code reprogramming. Org. Biomol. Chem. 13, 9353–9363 (2015).
Yu, H., Zhang, S. & Chaput, J. C. Darwinian evolution of an alternative genetic system provides support for TNA as an RNA progenitor. Nat. Chem. 4, 183–187 (2012).
Pinheiro, V. B. et al. Synthetic genetic polymers capable of heredity and evolution. Science 336, 341–344 (2012).
Taylor, A. I. et al. Catalysts from synthetic genetic polymers. Nature 518, 427–430 (2014).
Dunn, M. R. & Chaput, J. C. Reverse transcription of threose nucleic acid by a naturally occurring DNA polymerase. ChemBioChem 17, 1804–1808 (2016).
Vaught, J. D. et al. Expanding the chemistry of DNA for in vitro selection. J. Am. Chem. Soc. 132, 4141–4151 (2010).
Gold, L. et al. Aptamer-based multiplexed proteomic technology for biomarker discovery. PLoS ONE 5, e15004 (2010).
Davies, D. R. et al. Unique motifs and hydrophobic interactions shape the binding of modified DNA ligands to protein targets. Proc Natl. Acad. Sci. USA 109, 19971–19976 (2012).
Gupta, S. et al. Chemically-modified DNA aptamers bind interleukin-6 with high affinity and inhibit signaling by blocking its interaction with interleukin-6 receptor. J. Biol. Chem. 289, 8706–8719 (2014).
Gelinas, A. D. et al. Crystal structure of interleukin-6 in complex with a modified nucleic acid ligand. J. Biol. Chem. 289, 8720–8734 (2014).
Imaizumi, Y. et al. Efficacy of base-modification on target binding of small molecule DNA aptamers. J. Am. Chem. Soc. 135, 9412–9419 (2013).
Gawande, B. N. et al. Selection of DNA aptamers with two modified bases. Proc. Natl Acad. Sci. USA 114, 2898–2903 (2017).
Tolle, F., Brändle, G. M., Matzner, D. & Mayer, G. A versatile approach towards nucleobase-modified aptamers. Angew. Chem. Int. Ed. 54, 10971–10974 (2015).
Santoro, S. W., Joyce, G. F., Sakthivel, K., Gramatikova, S. & Barbas, C. F. III. RNA cleavage by a DNA enzyme with extended chemical functionality. J. Am. Chem. Soc. 122, 2433–2439 (2000).
Perrin, D. M., Garestier, T. & Hélène, C. Expanding the catalytic repertoire of nucleic acid catalysts: simultaneous incorporation of two modified deoxyribonucleoside triphosphates bearing ammonium and imidazolyl functionalities. Nucleosides Nucleotides 18, 377–391 (1999).
Perrin, D. M., Garestier, T. & Hélène, C. Bridging the gap between proteins and nucleic acids: a metal-independent RNAseA mimic with two protein-like functionalities. J. Am. Chem. Soc 123, 1556–1563 (2001).
Lermer, L., Roupioz, Y., Ting, R. & Perrin, D. M. Toward an RNaseA mimic: a DNAzyme with imidazoles and cationic amines. J. Am. Chem. Soc. 124, 9960–9961 (2002).
Hollenstein, M., Hipolito, C. J., Lam, C. H. & Perrin, D. M. A self-cleaving DNA enzyme modified with amines, guanidines and imidazoles operates independently of divalent metal cations (M2+). Nucleic Acids Res. 37, 1638–1649 (2009).
Shoji, A., Kuwahara, M., Ozaki, H. & Sawai, H. Modified DNA aptamer that binds the (R)-isomer of a thalidomide derivative with high enantioselectivity. J. Am. Chem. Soc. 129, 1456–1464 (2007).
Sidorov, A. V., Grasby, J. A. & Williams, D. M. Sequence‐specific cleavage of RNA in the absence of divalent metal ions by a DNAzyme incorporating imidazolyl and amino functionalities. Nucleic Acids Res. 32, 1591–1601 (2004).
Sefah, K. et al. In vitro selection with artificial expanded genetic information systems. Proc. Natl Acad. Sci. USA 111, 1449–1454 (2014).
Zhang, L. et al. Evolution of functional six-nucleotide DNA. J. Am. Chem. Soc. 137, 6734–6737 (2015).
Kimoto, M., Yamashige, R., Matsunaga, K., Yokoyama, S. & Hirao, I. Generation of high-affinity DNA aptamers using an expanded genetic alphabet. Nat. Biotechnol. 31, 453–457 (2013).
Hili, R., Niu, J. & Liu, D. R. DNA ligase-mediated translation of DNA into densely functionalized nucleic acid polymers. J. Am. Chem. Soc. 135, 98–101 (2013).
Lei, Y., Kong, D. & Hili, R. A high-fidelity codon set for the T4 DNA ligase-catalyzed polymerization of modified oligonucleotides. ACS Comb. Sci. 17, 716–721 (2015).
Cohen, J. C., Boerwinkle, E., Mosley, T. H. Jr & Hobbs, H. H. Sequence variations in PCSK9, low LDL, and protection against coronary heart disease. N. Engl. J. Med. 354, 1264–1272 (2006).
Lambert, G., Charlton, F., Rye, K.-A. & Piper, D. E. Molecular basis of PCSK9 function. Atherosclerosis 203, 1–7 (2009).
Stein, E. A. et al. Effect of a monoclonal antibody to PCSK9 on LDL cholesterol. N. Engl. J. Med. 366, 1108–1118 (2012).
Brudno, Y., Birnbaum, M. E., Kleiner, R. E. & Liu, D. R. An in vitro translation, selection and amplification system for peptide nucleic acids. Nat. Chem. Biol. 6, 148–155 (2010).
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003).
Ortigão, J. F. R. et al. Antisense effect of oligodeoxynucleotides with inverted terminal internucleotidic linkages: a minimal modification protecting against nucleolytic degradation. Antisense Res. Dev. 2, 129–146 (1992).
Lo Surdo, P. et al. Mechanistic implications for LDL receptor degradation from the PCSK9/LDLR structure at neutral pH. EMBO Rep. 12, 1300–1305 (2011).
Hunter, C. A. & Jones, S. A. IL-6 as a keystone cytokine in health and disease. Nat. Immunol. 16, 448–457 (2015).
Gibbs, J. P. et al. Impact of target-mediated elimination on the dose and regimen of evolocumab, a human monoclonal antibody against proprotein convertase subtilisin/kexin type 9 (PCSK9). J. Clin. Pharmacol. 57, 616–626 (2017).
Kühnast, S. et al. Alirocumab inhibits atherosclerosis, improves the plaque morphology, and enhances the effects of a statin. J. Lipid Res. 55, 2103–2112 (2014).
Guo, C., Watkins, C. P. & Hili, R. Sequence-defined scaffolding of peptides on nucleic acid polymers. J. Am. Chem. Soc. 137, 11191–11196 (2015).
Acknowledgements
This work was supported by the DARPA Fold Fx program (N66001-14-2-4053), the NIH R01 EB022376 (formerly R01 GM065400) and R35 GM118062, and the Howard Hughes Medical Institute. Z.C. was partially supported by the Y. Kishi Graduate Prize in Chemistry and Chemical Biology sponsored by the Eisai Corporation. The authors thank J. Niu, R. Hili, J. P. Maianti, D. L. Usanov and A. Chan for helpful discussions. We thank the Center for Macromolecular Interactions in the Department of Biological Chemistry and Molecular Pharmacology at Harvard Medical School for access to the Biacore T200 SPR instrument, and M. Blome, B. Lang and K. Arnett for technical assistance with the SPR experiments. We thank the Harvard FAS Small Molecule Mass Spectrometry facility for access to ESI-MS instruments and S. A. Trauger for technical assistance.
Author information
Authors and Affiliations
Contributions
Z.C. and D.R.L. conceived and designed the study. Z.C., P.A.L., A.P.B. and J.C.C. performed the experiments. Z.C., P.A.L. and D.R.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
Z.C., D.R.L., and Harvard University have filed patent applications on DNA-templated polymerization.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary information
Supplementary Figs. 1–13, Supplementary Materials and Methods
Rights and permissions
About this article
Cite this article
Chen, Z., Lichtor, P.A., Berliner, A.P. et al. Evolution of sequence-defined highly functionalized nucleic acid polymers. Nature Chem 10, 420–427 (2018). https://doi.org/10.1038/s41557-018-0008-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-018-0008-9
This article is cited by
-
Sequence structure controllable polymerization-induced self-assembly
Science China Chemistry (2024)
-
Nonviral delivery systems for antisense oligonucleotide therapeutics
Biomaterials Research (2022)
-
A mating mechanism to generate diversity for the Darwinian selection of DNA-encoded synthetic molecules
Nature Chemistry (2022)
-
Nonenzymatic polymerase-like template-directed synthesis of acyclic l-threoninol nucleic acid
Nature Communications (2021)
-
Reconstruction of evolving gene variants and fitness from short sequencing reads
Nature Chemical Biology (2021)