Abstract
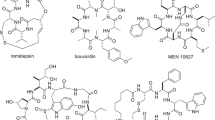
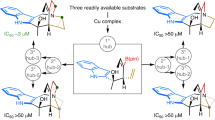
New methods capable of effecting cyclization, and forming novel three-dimensional structures while maintaining favourable physicochemical properties are needed to facilitate the development of cyclic peptide-based drugs that can engage challenging biological targets, such as protein–protein interactions. Here, we report a highly efficient and generally applicable strategy for constructing new types of peptide macrocycles using palladium-catalysed intramolecular C(sp3)–H arylation reactions. Easily accessible linear peptide precursors of simple and versatile design can be selectively cyclized at the side chains of either aromatic or modified non-aromatic amino acid units to form various cyclophane-braced peptide cycles. This strategy provides a powerful tool to address the long-standing challenge of size- and composition-dependence in peptide macrocyclization, and generates novel peptide macrocycles with uniquely buttressed backbones and distinct loop-type three-dimensional structures. Preliminary cell proliferation screening of the pilot library revealed a potent lead compound with selective cytotoxicity toward proliferative Myc-dependent cancer cell lines.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Driggers, E. M., Hale, S. P., Lee, J. & Terrett, N. K. The exploration of macrocycles for drug discovery: an underexploited structural class. Nat. Rev. Drug Discov. 7, 608–624 (2008).
Marsault, E. & Peterson, M. L. Macrocycles are great cycles: applications, opportunities, and challenges of synthetic macrocycles in drug discovery. J. Med. Chem. 54, 1961–2004 (2011).
Cardote, T. A. F. & Ciulli, A. Cyclic and macrocyclic peptides as chemical tools to recognise protein surfaces and probe protein–protein interactions. ChemMedChem 11, 787–794 (2016).
Walsh, C. T., Brien, R. V. O. & Khosla, C. Nonproteinogenic amino acid building blocks for nonribosomal peptide and hybrid polyketide scaffolds. Angew. Chem. Int. Ed. 52, 7098–7124 (2013).
White, C. J. & Yudin, A. K. Contemporary strategies for peptide macrocyclization. Nat. Chem. 3, 509–524 (2011).
Hill, T. A., Shepherd, N. E., Diness, F. & Fairlie, D. P. Constraining cyclic peptides to mimic protein structure motifs. Angew. Chem. Int. Ed. 53, 13020–13041 (2014).
Lau, Y. H., de Andrade, P., Wu, Y. & Spring, D. R. Peptide stapling techniques based on different macrocyclisation chemistries. Chem. Soc. Rev. 44, 91–102 (2015).
Veber, D. F. et al. Molecular properties that influence the oral bioavailability of drug candidates. J. Med. Chem. 45, 2615–2623 (2002).
Frost, J. R., Scully, C. C. G. & Yudin, A. K. Oxadiazole grafts in peptide macrocycles. Nat. Chem. 3, 1105–1111 (2016).
Spokoyny, A. M. et al. A perfluoroaryl-cysteine SNAr chemistry approach to unprotected peptide stapling. J. Am. Chem. Soc. 135, 5946–5949 (2013).
Osberger, T. J., Rogness, D. C., Kohrt, J. T., Stepan, A. F. & White, M. C. Oxidative diversification of amino acids and peptides by small-molecule iron catalysis. Nature 537, 214–219 (2016).
Blackwell, H. E. & Grubbs, R. H. Highly efficient synthesis of covalently cross-linked peptide helices by ring-closing metathesis. Angew. Chem. Int. Ed. 37, 3281–3284 (1998).
Kim, Y.-W., Grossmann, T. N. & Verdine, G. L. Synthesis of all-hydrocarbon stapled α-helical peptides by ring-closing olefin metathesis. Nat. Protoc. 6, 761–771 (2011).
Lawson, K. V., Rose, T. E. & Harran, P. G. Template-constrained macrocyclic peptides prepared from native, unprotected precursors. Proc. Natl Acad. Sci. USA 110, E3753–E3760 (2013).
Beckmann, H. S. G. et al. A strategy for the diversity-oriented synthesis of macrocyclic scaffolds using multidimensional coupling. Nat. Chem. 5, 861–867 (2013).
Rogdan, A. R., Jerome, S. V., Houk, K. N. & James, K. Strained cyclophane macrocycles: impact of progressive ring size reduction on synthesis and structure. J. Am. Chem. Soc. 132, 2127–2138 (2010).
Bockus, A. T., McEwen, C. M. & Lokey, R. S. Form and function in cyclic peptide natural products: a pharmacokinetic perspective. Curr. Top. Med. Chem. 13, 821–836 (2013).
Booker, S. J. Anaerobic functionalization of unactivated C–H bonds. Curr. Opin. Chem. Biol. 13, (58–73 (2009).
Sydor, P. K. et al. Regio- and stereodivergent antibiotic oxidative carbocyclizations catalysed by Rieske oxygenase-like enzymes. Nat. Chem. 3, 388–392 (2011).
Schramma, K. R., Bushin, L. B. & Seyedsayamdost, M. R. Structure and biosynthesis of a macrocyclic peptide containing an unprecedented lysine-to-tryptophan crosslink. Nat. Chem. 7, 431–437 (2015).
Feng, Y. & Chen, G. Total synthesis of celogentin C by stereoselective C–H activation. Angew. Chem. Int. Ed. 49, 958–961 (2010).
Cram, D. J. & Cram, J. M. Cyclophane chemistry: bent and battered benzene rings. Acc. Chem. Res. 4, 204–213 (1970).
Gulder, T. & Baran, P. S. Strained cyclophane natural products: macrocyclization at its limits. Nat. Prod. Rep. 29, 899–934 (2012).
Dong, H., Limberakis, C., Liras, S., Price, D. & James, K. Peptidic macrocyclization via palladium-catalyzed chemoselective indole C2 arylation. Chem. Commun. 48, 11644–11646 (2012).
Mendive-Tapia, L. et al. New peptide architectures through C–H activation stapling between tryptophan-phenylalanine/tyrosine residues. Nat. Commun. 6, 7160 (2015).
Gong, W., Zhang, G., Liu, T., Giri, R. & Yu, J.-Q. Site-selective C(sp 3)−H functionalization of di-, tri-, and tetrapeptides at the N-terminus. J. Am. Chem. Soc. 136, 16940–16946 (2014).
Noisier, A. F. M., García, J., Ionut, I. A. & Albericio, F. Stapled peptides by late-stage C(sp 3)–H activation. Angew. Chem., Int. Ed. 56, 314–318 (2017).
Tang, J., He, Y., Chen, H., Sheng, W. & Wang, H. Synthesis of bioactive and stabilized cyclic peptides by macrocyclization using C(sp 3)–H activation. Chem. Sci. 8, 4565–4570 (2017).
Godula, K. & Sames, D. C–H Bond functionalization in complex organic synthesis. Science 312, 67–72 (2006).
Yamaguchi, J., Yamaguchi, A. D. & Itami, K. C–H Bond functionalization: emerging synthetic tools for natural products and pharmaceuticals. Angew. Chem. Int. Ed. 51, 8960–9009 (2012).
McMurray, L., O’Hara, F. & Gaunt, M. J. Recent developments in natural product synthesis using metal-catalysed C–H bond functionalisation. Chem. Soc. Rev. 40, 1885–1898 (2011).
Chen, X., Engle, K. M., Wang, D.-H. & Yu, J.-Q. Palladium(II)-catalyzed C–H activation/C–C cross-coupling reactions: versatility and practicality. Angew. Chem. Int. Ed. 48, 5094–5115 (2009).
Lyons, T. W. & Sanford, M. S. Palladium-catalyzed ligand-directed C–H functionalization reactions. Chem. Rev. 110, 1147–1169 (2010).
Noisier, F. M. & Brimble, M. A. C–H functionalization in the synthesis of amino acids and peptides. Chem. Rev. 114, 8775–8806 (2014).
Zaitsev, V. G., Shabashov, D. & Daugulis, O. Highly regioselective arylation of sp 3 C–H bonds catalyzed by palladium acetate. J. Am. Chem. Soc. 127, 13154–13155 (2005).
Shabashov, M. & Daugulis, O. Auxiliary-assisted palladium-catalyzed arylation and alkylation of sp 2 and sp 3 carbon–hydrogen bonds. J. Am. Chem. Soc. 132, 3965–3972 (2010).
Daugulis, O., Do, H. & Shabashov, D. Palladium- and copper-catalyzed arylation of carbon-hydrogen bonds. Acc. Chem. Res. 42, 1074–1086 (2009).
Reddy, B. V. S., Reddy, L. R. & Corey, E. J. Novel acetoxylation and C−C Coupling reactions at unactivated positions in α-amino acid derivatives. Org. Lett. 8, 3391–3394 (2006).
He, G., Wang, B., Nack, W. A. & Chen, G. Syntheses and transformations of α‐amino acids via palladium-catalyzed auxiliary-directed sp 3 C−H functionalization. Acc. Chem. Res. 49, 635–645 (2016).
Feng, Y., Wang, Y., Landgraf, B., Liu, S. & Chen, G. Facile benzo-ring construction via palladium-catalyzed functionalization of unactivated sp 3 C−H bonds under mild reaction conditions. Org. Lett. 12, 3414–3417 (2010).
He, G., Zhang, S., Nack, W. A. & Chen, G. Use of a readily removable auxiliary group for the synthesis of pyrrolidones by the palladium-catalyzed intramolecular amination of unactivated γ C(sp 3)−H bonds. Angew. Chem., Int. Ed. 52, 11124–11128 (2013).
Frisch, M. J. et al. Gaussian 09, Revision D.01 (Gaussian, 2009).
Lapointe, D. & Fagnou, K. Overview of the mechanistic work on the concerted metallationdeprotonation pathway. Chem. Lett. 39, 1118–1126 (2010).
Wang, B., Nack, W. A., He, G., Zhang, S.-Y. & Chen, G. Palladium-catalyzed trifluoroacetate-promoted mono-arylation of the methyl group of alanine at room temperature: synthesis of β-arylated α-amino acids through sequential C–H functionalization. Chem. Sci. 5, 3952–3957 (2014).
Dang, Y. et al. The mechanism of a ligand-promoted C(sp 3)–H activation and arylation reaction via palladium catalysis: theoretical demonstration of a Pd(ii)/Pd(iv) redox manifold. J. Am. Chem. Soc. 137, 2006–2014 (2015).
Hickman, A. J. & Sanford, M. S. High-valent organometallic copper and palladium in catalysis. Nature 484, 177–185 (2012).
Dang, C. V. MYC on the path to cancer. Cell 149, 22–35 (2012).
Jain, M. et al. Sustained loss of a neoplastic phenotype by brief inactivation of MYC. Science 297, 102–104 (2002).
McKeown, M. R. & Bradner, J. E. MYC activation is a hallmark of cancer initiation and maintenance. Cold Spring Harb. Persp. Med. 4, a014241 (2014).
Ottaviani, G., Martel, S. & Carrupt, P.-A. Parallel artificial membrane permeability assay: a new membrane for the fast prediction of passive human skin permeability. J. Med. Chem. 49, 3948–3954 (2006).
Acknowledgements
G.C. thanks the State Key Laboratory of Elemento-Organic Chemistry at Nankai University, NSFC-21672105, NSFC-21421062, the ‘111’ project (B06005) of the Ministry of Education of China, and programme 973 (2014CB849603 to X.Q.) for financial support of the experimental part of this work. P.L. thanks the University of Pittsburgh for financial support for the computational part of the work. Calculations were performed at the Center for Simulation and Modeling at the University of Pittsburgh and the Extreme Science and Engineering Discovery Environment (XSEDE) supported by the National Science Foundation. W.S. and M.M. thank M. Hull, M. Wogan, H. Nguyen and E. Chen of Calibr for technical support and help. G.C. dedicates this work to Q. Zhou on the occasion of his 60th birthday.
Author information
Authors and Affiliations
Contributions
X.Z. carried out most of the reaction optimization and structural determination of products, and prepared the Supplementary Information. Y.M. developed peptide macrocyclization at non-aromatic amino acid units. M.Z., W.H., Y.H. and Q.W. prepared some amino acid building blocks and peptide substrates. J.C. conducted all the X-ray crystallography experiments. G.L. conducted the computations. M.M. carried out the cell proliferation assays. W.S. supervised the biological activity studies. X.Q. advised the macrocycles druggability especially the permeability optimization and directed the PAMPA assay. M.S. carried out the PAMPA assays and analysed the PAMPA data. G.H. supervised experimental studies. P.L. directed the computational studies. P.L. and G.L. prepared the computational sections of the manuscript. G.C. formulated the initial ideas of this work, supervised the project, coordinated with P.L. on computational studies, coordinated with W.S. on biological studies, and prepared most of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supporting Information
Supplementary Experimental Details, Supplementary Data and Supplementary Figures.
Crystallographic data
Crystallographic data for compound 3a; CCDC reference: 1526698
Crystallographic data
Structure factors file for compound 3a; CCDC reference: 1526698
Crystallographic data
Crystallographic data for compound 3b; CCDC reference: 1526699
Crystallographic data
Structure factors file for compound 3b; CCDC reference: 1526699
Crystallographic data
Crystallographic data for compound 11a; CCDC reference: 1526702
Crystallographic data
Structure factors file for compound 11a; CCDC reference: 1526702
Crystallographic data
Crystallographic data for compound 17; CCDC reference: 1526701
Crystallographic data
Structure factors file for compound 17; CCDC reference: 1526701
Crystallographic data
Crystallographic data for compound 29a; CCDC reference: 1526700
Crystallographic data
Structure factors file for compound 29a; CCDC reference: 1526700
Crystallographic data
Crystallographic data for compound 29b; CCDC reference: 1526703
Crystallographic data
Structure factors file for compound 29b; CCDC reference: 1526703
Crystallographic data
Crystallographic data for compound 31a; CCDC reference: 1526704
Crystallographic data
Structure factors file for compound 31; CCDC reference: 1526704
Crystallographic data
Crystallographic data for compound 32; CCDC reference: 1526705
Crystallographic data
Structure factors file for compound 32; CCDC reference: 1526705
Crystallographic data
Crystallographic data for compound 34a; CCDC reference: 1526707
Crystallographic data
Structure factors file for compound 34a; CCDC reference: 1526707
Rights and permissions
About this article
Cite this article
Zhang, X., Lu, G., Sun, M. et al. A general strategy for synthesis of cyclophane-braced peptide macrocycles via palladium-catalysed intramolecular sp3 C−H arylation. Nature Chem 10, 540–548 (2018). https://doi.org/10.1038/s41557-018-0006-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-018-0006-y
This article is cited by
-
Recent advances in chemical protein synthesis: method developments and biological applications
Science China Chemistry (2024)
-
Modular synthesis of clickable peptides via late-stage maleimidation on C(7)-H tryptophan
Nature Communications (2023)
-
Photochemical single-step synthesis of β-amino acid derivatives from alkenes and (hetero)arenes
Nature Chemistry (2022)
-
Late-stage C–H functionalization offers new opportunities in drug discovery
Nature Reviews Chemistry (2021)
-
Chemodivergent manganese-catalyzed C–H activation: modular synthesis of fluorogenic probes
Nature Communications (2021)