Abstract
Activation of receptor protein kinases is prevalent in various cancers with unknown impact on ferroptosis. Here we demonstrated that AKT activated by insulin-like growth factor 1 receptor signalling phosphorylates creatine kinase B (CKB) T133, reduces metabolic activity of CKB and increases CKB binding to glutathione peroxidase 4 (GPX4). Importantly, CKB acts as a protein kinase and phosphorylates GPX4 S104. This phosphorylation prevents HSC70 binding to GPX4, thereby abrogating the GPX4 degradation regulated by chaperone-mediated autophagy, alleviating ferroptosis and promoting tumour growth in mice. In addition, the levels of GPX4 are positively correlated with the phosphorylation levels of CKB T133 and GPX4 S104 in human hepatocellular carcinoma specimens and associated with poor prognosis of patients with hepatocellular carcinoma. These findings reveal a critical mechanism by which tumour cells counteract ferroptosis by non-metabolic function of CKB-enhanced GPX4 stability and underscore the potential to target the protein kinase activity of CKB for cancer treatment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The human HCC data were derived from the TCGA Research Network. The dataset derived from this resource that supports the findings of this study is available at the following link: https://tcga.xenahubs.net/download/TCGA.LIHC.sampleMap/HiSeqV2.gz. Mass spectrometry data have been deposited in ProteomeXchange with the accession code PXD040322. UniProt protein database (EMBL-EBI) was used for protein identification. Source data are provided with this paper.
References
Stockwell, B. R. et al. Ferroptosis: a regulated cell death nexus linking metabolism, redox biology, and disease. Cell 171, 273–285 (2017).
Dixon, S. J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149, 1060–1072 (2012).
Jiang, X., Stockwell, B. R. & Conrad, M. Ferroptosis: mechanisms, biology and role in disease. Nat. Rev. Mol. Cell Biol. 22, 266–282 (2021).
Yang, W. S. et al. Regulation of ferroptotic cancer cell death by GPX4. Cell 156, 317–331 (2014).
Ingold, I. et al. Selenium utilization by GPX4 is required to prevent hydroperoxide-induced ferroptosis. Cell 172, 409–422.e21 (2018).
Yant, L. J. et al. The selenoprotein GPX4 is essential for mouse development and protects from radiation and oxidative damage insults. Free Radic. Biol. Med 34, 496–502 (2003).
Stockwell, B. R. Dawn of a new era of targeted antioxidant therapies. Cell Chem. Biol. 26, 1483–1485 (2019).
Hangauer, M. J. et al. Drug-tolerant persister cancer cells are vulnerable to GPX4 inhibition. Nature 551, 247–250 (2017).
Zou, Y. et al. A GPX4-dependent cancer cell state underlies the clear-cell morphology and confers sensitivity to ferroptosis. Nat. Commun. 10, 1617 (2019).
Li, X., Egervari, G., Wang, Y., Berger, S. L. & Lu, Z. Regulation of chromatin and gene expression by metabolic enzymes and metabolites. Nat. Rev. Mol. Cell Biol. 19, 563–578 (2018).
Li, X., Qian, X. & Lu, Z. Fructokinase A acts as a protein kinase to promote nucleotide synthesis. Cell Cycle 15, 2689–2690 (2016).
Li, X., Zheng, Y. & Lu, Z. PGK1 is a new member of the protein kinome. Cell Cycle 15, 1803–1804 (2016).
Xu, D. et al. The evolving landscape of noncanonical functions of metabolic enzymes in cancer and other pathologies. Cell Metab. 33, 33–50 (2021).
Xu, D. et al. The protein kinase activity of fructokinase A specifies the antioxidant responses of tumor cells by phosphorylating p62. Sci. Adv. 5, eaav4570 (2019).
Lu, Z. & Hunter, T. Metabolic kinases moonlighting as protein kinases. Trends Biochem. Sci. 43, 301–310 (2018).
Lu, S. & Wang, Y. Nonmetabolic functions of metabolic enzymes in cancer development. Cancer Commun. 38, 63 (2018).
Jiang, H., Zhu, L., Xu, D. & Lu, Z. A newly discovered role of metabolic enzyme PCK1 as a protein kinase to promote cancer lipogenesis. Cancer Commun. 40, 389–394 (2020).
Jiang, H. et al. Fructose and fructose kinase in cancer and other pathologies. J. Genet. Genomics 48, 531–539 (2021).
Kazak, L. & Cohen, P. Creatine metabolism: energy homeostasis, immunity and cancer biology. Nat. Rev. Endocrinol. 16, 421–436 (2020).
Loo, J. M. et al. Extracellular metabolic energetics can promote cancer progression. Cell 160, 393–406 (2015).
Nagpal, L., Kornberg, M. D., Albacarys, L. K. & Snyder, S. H. Inositol hexakisphosphate kinase-2 determines cellular energy dynamics by regulating creatine kinase-B. Proc. Natl Acad. Sci. USA 118, e2020695118 (2021).
Rahbani, J. F. et al. Creatine kinase B controls futile creatine cycling in thermogenic fat. Nature 590, 480–485 (2021).
Papalazarou, V. et al. The creatine-phosphagen system is mechanoresponsive in pancreatic adenocarcinoma and fuels invasion and metastasis. Nat. Metab. 2, 62–80 (2020).
Kazak, L. et al. A creatine-driven substrate cycle enhances energy expenditure and thermogenesis in beige fat. Cell 163, 643–655 (2015).
Breuhahn, K. & Schirmacher, P. Reactivation of the insulin-like growth factor-II signaling pathway in human hepatocellular carcinoma. World J. Gastroenterol. 14, 1690–1698 (2008).
Xu, D. et al. The gluconeogenic enzyme PCK1 phosphorylates INSIG1/2 for lipogenesis. Nature 580, 530–535 (2020).
Zhang, Y. et al. mTORC1 couples cyst(e)ine availability with GPX4 protein synthesis and ferroptosis regulation. Nat. Commun. 12, 1589 (2021).
Liu, R. et al. Choline kinase alpha 2 acts as a protein kinase to promote lipolysis of lipid droplets. Mol. Cell 81, 2722–2735.e9 (2021).
Li, X. et al. Mitochondria-translocated PGK1 functions as a protein kinase to coordinate glycolysis and the TCA cycle in tumorigenesis. Mol. Cell 61, 705–719 (2016).
Lin, L., Perryman, M. B., Friedman, D., Roberts, R. & Ma, T. S. Determination of the catalytic site of creatine kinase by site-directed mutagenesis. Biochim. Biophys. Acta 1206, 97–104 (1994).
Wu, Z. et al. Chaperone-mediated autophagy is involved in the execution of ferroptosis. Proc. Natl Acad. Sci. USA 116, 2996–3005 (2019).
Circu, M. L. & Aw, T. Y. Redox biology of the intestine. Free Radic. Res. 45, 1245–1266 (2011).
Droge, W. Free radicals in the physiological control of cell function. Physiol. Rev. 82, 47–95 (2002).
Yang, Y. & Yee, D. IGF-I regulates redox status in breast cancer cells by activating the amino acid transport molecule xC. Cancer Res. 74, 2295–2305 (2014).
Hodgson, N., Trivedi, M., Muratore, C., Li, S. & Deth, R. Soluble oligomers of amyloid-beta cause changes in redox state, DNA methylation, and gene transcription by inhibiting EAAT3 mediated cysteine uptake. J. Alzheimers Dis. 36, 197–209 (2013).
Takahashi, S., Hisatsune, A., Kurauchi, Y., Seki, T. & Katsuki, H. Insulin-like growth factor 1 specifically up-regulates expression of modifier subunit of glutamate-cysteine ligase and enhances glutathione synthesis in SH-SY5Y cells. Eur. J. Pharmacol. 771, 99–106 (2016).
Hayes, J. D. & Dinkova-Kostova, A. T. The Nrf2 regulatory network provides an interface between redox and intermediary metabolism. Trends Biochem. Sci. 39, 199–218 (2014).
Ghoneum, A., Abdulfattah, A. Y., Warren, B. O., Shu, J. & Said, N. Redox homeostasis and metabolism in cancer: a complex mechanism and potential targeted therapeutics. Int. J. Mol. Sci. 21, 3100 (2020).
Canli, O. et al. Glutathione peroxidase 4 prevents necroptosis in mouse erythroid precursors. Blood 127, 139–148 (2016).
Koppula, P. et al. A targetable CoQ-FSP1 axis drives ferroptosis- and radiation-resistance in KEAP1 inactive lung cancers. Nat. Commun. 13, 2206 (2022).
Xue, J. et al. The Nrf2/GCH1/BH4 axis ameliorates radiation-induced skin injury by modulating the ROS cascade. J. Invest. Dermatol. 137, 2059–2068 (2017).
Imai, H. et al. Failure of the expression of phospholipid hydroperoxide glutathione peroxidase in the spermatozoa of human infertile males. Biol. Reprod. 64, 674–683 (2001).
Imai, H. et al. Depletion of selenoprotein GPx4 in spermatocytes causes male infertility in mice. J. Biol. Chem. 284, 32522–32532 (2009).
Dasgupta, S. et al. Metabolic enzyme PFKFB4 activates transcriptional coactivator SRC-3 to drive breast cancer. Nature 556, 249–254 (2018).
Guo, D. et al. Aerobic glycolysis promotes tumor immune evasion by hexokinase2-mediated phosphorylation of IκBα. Cell Metab. 34, 1312–1324.e6 (2022).
Li, X. et al. A splicing switch from ketohexokinase-C to ketohexokinase-A drives hepatocellular carcinoma formation. Nat. Cell Biol. 18, 561–571 (2016).
Du, L. et al. β-Catenin induces transcriptional expression of PD-L1 to promote glioblastoma immune evasion. J. Exp. Med. 217, e20191115 (2020).
Qian, X. et al. KDM3A senses oxygen availability to regulate PGC-1α-mediated mitochondrial biogenesis. Mol. Cell 76, 885–895.e7 (2019).
Qian, X. et al. PTEN suppresses glycolysis by dephosphorylating and inhibiting autophosphorylated PGK1. Mol. Cell 76, 516–527.e7 (2019).
Li, X. et al. Nucleus-translocated ACSS2 promotes gene transcription for lysosomal biogenesis and autophagy. Mol. Cell 66, 684–697.e9 (2017).
Mao, C. et al. DHODH-mediated ferroptosis defence is a targetable vulnerability in cancer. Nature 593, 586–590 (2021).
Wu, J. et al. Intercellular interaction dictates cancer cell ferroptosis via NF2-YAP signalling. Nature 572, 402–406 (2019).
Xu, D. Q. et al. PAQR3 controls autophagy by integrating AMPK signaling to enhance ATG14L-associated PI3K activity. EMBO J. 35, 496–514 (2016).
Xu, D. et al. PAQR3 modulates cholesterol homeostasis by anchoring Scap/SREBP complex to the Golgi apparatus. Nat. Commun. 6, 8100 (2015).
Yang, W. et al. Nuclear PKM2 regulates β-catenin transactivation upon EGFR activation. Nature 480, 118–122 (2011).
Lee, J. H. et al. EGFR-phosphorylated platelet isoform of phosphofructokinase 1 promotes PI3K activation. Mol. Cell 70, 197–210.e7 (2018).
Acknowledgements
This study was supported by grants from the Ministry of Science and Technology of the People’s Republic of China (2021YFA0805600, D.X.; 2020YFA0803300, Z.L.), the National Natural Science Foundation of China (82188102 and 82030074, Z.L.; 92157113 and 82072630, D.X.; 82173114, Z.W.; 82072903 and 82272872, T.L.; 82002811, M.Y.; 82030065 and 81873932, Q.Z.), the Zhejiang Natural Science Foundation Key Project (LD22H160002, D.X.; LD21H160003, Z.L.), Zhejiang Natural Science Foundation Discovery Project (LQ22H160023, Z.W.), Chinese Academy of Sciences Intramural Research Program (QTJC20220010-A, Q.X.) and the Leading Innovative and Entrepreneur Team Introduction Program of Zhejiang (2019R01001, Z.L.). Z.L. is the Kuancheng Wang Distinguished Chair. The authors received no specific funding for this work.
Author information
Authors and Affiliations
Contributions
D.X., Z.L. and Q.Z. conceived and designed the study and wrote the paper. K.W., M.Y., T.L., Z.W., Y.D., G.J., Y.S., L.W., L.L., P.Z., B.D., X.Y., H.S., T.W., J.Z., J.Y. and Y.D. performed the experiments and analysed the data. Y.X., Q.W. and L.X. reviewed and edited the paper. X.Q., L.M. and J.F. provided the technical support in gene editing and interpretation of the data.
Corresponding authors
Ethics declarations
Competing interests
Z.L. owns shares in Signalway Biotechnology (Pearland, TX), which supplied rabbit antibodies that recognize CKB pT133 and GPX4 pS104. Z.L.’s interest in this company had no bearing on its being chosen to supply these reagents. The other authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Lawrence Kazak and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 CKB T133 phosphorylation upregulates GPX4 stability.
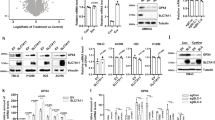
(b, d-h, l,m, r) Immunoprecipitation and immunoblotting analyses were performed with the indicated antibodies. (a, b, q-s) Data are the mean ± SD (n = 6). N.S., not significant (a, q, s); **P < 0.001 (r); ***P < 0.0001(b) by two-tailed Student’s t-test. (a, m, q-s) Huh7 and HCCLM3 were treated with IGF1 for the indicated time (a). The indicated cells expressing CKB shRNA with reconstituted expression of the indicated CKB proteins were harvested (m, s), or treated with IGF1 for 12 h (q), or treated with CHX for the indicated time (r). The mRNA expression levels of Gpx4 gene were measured using qPCR (a, q, s). (b) Huh7 and HCCLM3 cells were treated with CHX for the indicated time in the presence of IGF1. The quantification of GPX4 protein levels relative to initial protein levels is shown. (c) Selected mass spectrometry identified peptide hits are shown. (d) Huh7 cells were pretreated with or without indicated inhibitors for 30 min before treatment with or without IGF1 for 1 h. (e) Huh7 cells transfected with or without the indicated plasmid were treated with or without Tris DBA (10 μM) for 12 h. (f) Huh7 cells were treated with or without IGF1 (100 ng/ml) for 1 h. (g) A GST pull-down assay was performed as indicated. (h) In vitro kinase assays were performed in the presence of [γ-32P] ATP. Autoradiography were performed. (i) Purified GST-CKB proteins were incubated with His-AKT1 for an in vitro kinase assay. Mass-spectrometric analysis was performed. (j) Alignment of protein sequences spanning CKB T133 from different species. (k, l) Huh7 cells expressing Flag-CKB were treated with or without IGF1 for 1 h (l). IHC analyses of human HCC samples (k) or immunoblotting (l) were performed with the indicated antibodies and a CKB pT133-blocking peptide. (n-p) Genomic DNA was extracted from two individual clones of indicated cells with knock-in expression of CKB T133A. PCR products amplified from the indicated DNA fragments were shown (n, o) and sequenced (p).
Extended Data Fig. 2 GPX4 S104 phosphorylation stabilizes GPX4.
(a,b, h, j, k, p, t-x) Immunoprecipitation and immunoblotting analyses were performed with the indicated antibodies. (a-e, o, q-s, v-x) Data are the mean ± SD (n = 6). **P < 0.001 (v-x); ***P < 0.0001 (a-e, o, q-s, v-x); N.S., not significant (b, c, r, s) by two-tailed Student’s t-test. (a) The indicated cells treated with IGF1 for 1 h were harvested. (b-f) The indicated purified His-CKB on Ni-NTA agarose beads were incubated with active GST-AKT1 for an in vitro kinase assay. The binding affinity of CKB to creatine (b, d) and the relative CKB activity (c, e) were measured. AKT-phosphorylated CKB conjugated on Ni-NTA beads were washed and incubated with His-GPX4 proteins for an in vitro kinase assay. Mass-spectrometric analysis was performed (f). (g) Alignment of protein sequences spanning GPX4 S104. (h) Purified His-CKB were incubated with GST-GPX4 in the presence of [γ-32P] ATP for in vitro kinase assay. Autoradiography was performed. (i, j) Huh7 cells expressing Flag-GPX4 were treated with IGF1 for 1 h (j). IHC analyses of human HCC samples (i) or immunoblotting (j) were performed with the indicated antibodies and GPX4 pS104-blocking peptide. (k) Huh7 cells expressing Flag-GPX4 were transfected with HA-myr-AKT1. (l-n) Genomic DNA was extracted from the indicated cells. Amplified PCR products were shown (l, m) and sequenced (n). (o, p) Huh7 cells were pretreated with NAC (5 mM) for 30 min before IGF1 incubation for 12 h. The intercellular ROS levels were measured (o). (q-s) Relative CKB activity (q), binding affinity of the His-CKB proteins to creatine (r) and ATP (s) were measured. (t, u) Huh7 or HCCLM3 cells expressing CKB shRNA with reconstituted expression of the indicated CKB proteins were treated with or without IGF1 for 1 h . (v-x) The indicated cells expressing CKB shRNA with reconstituted expression of the indicated CKB proteins (v) or with knock-in expression of GPX4 mutants (w, x) were treated with CHX for the indicated time. The quantification of GPX4 protein levels relative to initial protein levels is shown.
Extended Data Fig. 3 CKB-mediated GPX4 S104 phosphorylation stabilizes GPX4.
(a, b) Alignment of protein sequences spanning GPX4 S104 and the adjacent CMA-target motif from different species. The structure of GPX4 exhibiting CMA-target motif and S104. (c) Purified His-CKB WT or T133D proteins on Ni-NTA beads were incubated with the indicated GST-GPX4 proteins for an in vitro kinase assay. Then the beads were removed and the remaining GST-GPX4 proteins were incubated with His-HSC70 followed by a GST pulldown assay as indicated. (d, e) Huh7 and HCCLM3 cells expressing GPX4 shRNA with reconstituted expression of the indicated GPX4 proteins were harvested for immunoblotting and mRNA detection using quantitative PCR as indicated. Data are the mean ± SD (n = 6). N.S., not significant (two-tailed Student’s t-test).
Extended Data Fig. 4 GPX4 S104 phosphorylation suppresses Lipid peroxidation.
(a-d) Parental Huh7 and HCCLM3 cells and the indicated clones with knock-in expression of CKB T133A mutants (upper) or GPX4 S104A (lower) were treated with or without cystine deprivation (a, c) or 20 μM Erastin (b, d) and IGF1 for 24 h. Reduced GSH (a, b) and GSSG (c, d) were measured respectively. Data are the mean ± SD (n = 6), ^P < 0.05; *P < 0.01; **P < 0.001; ***P < 0.0001 by two-tailed Student’s t-test. (e) Parental Huh7 and HCCLM3 cells and the indicated clones with knock-in expression of CKB T133A mutants (left) or GPX4 S104A (right) were treated with or without cystine deprivation or 20 μM Erastin in the presence or absence of IGF1 (100 ng/ml) for 24 h. The cell lysates were harvested for immunoblotting analyses as indicated. (f-k) Parental HCCLM3 (f, g, j,k) or Huh7 (h, i) cells and the indicated clones with knock-in expression of CKB T133A or GPX4 S104A mutants were treated with or without cystine deprivation (f, h, j) or 20 μM Erastin (g, i, k) combined with Fer-1 (2 μM) in the absence or presence of IGF1 for 24 h. Lipid ROS-positive cells were measured. Data are the mean ± SD (n = 3), *P < 0.01 by two-tailed Student’s t-test (f, g). Lipid peroxidation was assessed using BODIPY 581/591 C11 staining followed by FACS analysis.
Extended Data Fig. 5 GPX4 S104 phosphorylation suppresses ferroptosis.
(a, b, d-k) Data are the mean ± SD, *P < 0.01; **P < 0.001;***P < 0.0001 by two-tailed Student’s t-test. (a, b, d-g) Parental Huh7 cells and the indicated clones with knock-in expression of CKB T133A or GPX4 S104A mutants were treated with or without cystine deprivation (a, d, f) or 20 μM Erastin (b, e, g) in the presence or absence of IGF1 (100 ng/ml) for 24 h. The indicated mRNA expression levels were measured using quantitative PCR. GSSG (d, e) and reduced GSH (f, g) were measured respectively (n = 6). (c) Huh7 cells stably transfected with GLRX and GSTA1 shRNA were harvested for immunoblotting analyses as indicated. (h, i) Parental HCCLM3 cells and the indicated clones with knock-in expression of CKB T133A or GPX4 S104A mutants were treated with or without cystine deprivation (h) or 20 μM Erastin (i) combined with Fer-1 (2 μM) in the absence or presence of IGF1 for 24 h. Cell death were measured respectively (n = 3). (j, k) Parental HCCLM3 cells and the indicated clones with knock-in expression of CKB T133A mutants or GPX4 S104A were treated with different doses of sulfasalazine (SAS) (j) or Erastin (k) for 24 h, cell viability were measured (n = 3). (l) Parental Huh7 and HCCLM3 cells and the indicated clones with knock-in expression of CKB T133A mutants were treated with or without IGF1 for 12 h. Immunoblotting analyses were performed with the indicated antibodies.
Extended Data Fig. 6 CKB suppresses ferroptosis in a GPX4 pS104 dependent manner.
(a-l) Data are the mean ± SD (n = 3), *P < 0.01; **P < 0.001; N.S., not significant by two-tailed Student’s t-test. (a-d) Huh7 and HCCLM3 cells expressing GPX4 shRNA with reconstituted expression of the indicated GPX4 proteins were treated with or without cystine deprivation (a, c) or 20 μM Erastin (b, d) for 24 h. Lipid ROS-positive cells (a, b) and cell death (c, d) were measured respectively. (e, f) Huh7 and HCCLM3 cells expressing GPX4 shRNA with reconstituted expression of the indicated GPX4 proteins were treated with different doses of sulfasalazine (SAS) (e) or Erastin (f) for 24 h, cell viability were measured. (g-l) Parental Huh7 cells and the indicated clones with knock-in expression of GPX4 S104A mutants were stably transfected with CKB shRNA and reconstituted with indicated CKB proteins. The cells were treated with or without cystine deprivation (g, i) or 20 μM Erastin (h, j) for 24 h. Lipid ROS-positive cells (g, h) and cell death (i, j) were measured respectively. The cells were treated with different doses of sulfasalazine (SAS) (k) or Erastin (l) for 24 h, cell viability were measured.
Extended Data Fig. 7 CKB suppresses ferroptosis through its protein kinase activity.
(a-l) Data are the mean ± SD (n = 3), ^ P < 0.05; *P < 0.01; **P < 0.001; N.S., not significant by two-tailed Student’s t-test. (a-d) Huh7 and HCCLM3 cells expressing IGF1R-CA and CKB shRNA with reconstituted expression of the indicated CKB proteins were treated with or without cystine deprivation (a, c) or 20 μM Erastin (b, d) for 24 h. Lipid ROS-positive cells (a, b) and cell death (c, d) were measured respectively. (e, f) Huh7 and HCCLM3 cells expressing IGF1R-CA and CKB shRNA with reconstituted expression of the indicated CKB proteins were treated with different doses of sulfasalazine (SAS) (e) or Erastin (f) for 24 h, cell viability were measured. (g-j) Parental Huh7 and HCCLM3 cells and the indicated clones with knock-in expression of CKB T133A (g, i) or GPX4 S104A mutants (h, j) were treated with or without 2 μM RSL3 for 8 h. Lipid ROS-positive cells (g, h) and cell death (i, j) were measured respectively. (k, l) Parental Huh7 and HCCLM3 cells and the indicated clones with knock-in expression of CKB T133A (k) and GPX4 S104A mutants (l) were treated with different doses of RSL3 for 8 h, cell viability were measured.
Extended Data Fig. 8 CKB phosphorylates GPX4 to inhibit ferroptosis.
(a, b) MDA-MB-231, A549 and HCT116 cells expressing CKB shRNA with reconstituted expression of the indicated CKB proteins were treated with or without IGF1 for 12 h. Immunoblotting analyses were performed with the indicated antibodies. (c, d) MDA-MB-231, A549 and HCT116 cells expressing CKB shRNA with reconstituted expression of the indicated CKB proteins were treated with or without 20 μM Erastin in the absence or presence of IGF1 for 24 h. Lipid ROS-positive cells (c) and cell death (d) were measured respectively. Data are the mean ± SD (n = 3), ^ P < 0.05; *P < 0.01; **P < 0.001 by two-tailed Student’s t-test. (e-g) Genomic DNA was extracted from parental MDA-MB-231, A549 and HCT116 cells and the indicated clones with knock-in expression of GPX4 S104A. PCR products amplified from the indicated DNA fragments were shown (e, f) and sequenced (g). (h) Parental MDA-MB-231, A549 and HCT116 cells and the indicated clones with knock-in expression of GPX4 S104A mutants were treated with or without IGF1 for 12 h. Immunoblotting analyses were performed with the indicated antibodies. (i, j) Parental MDA-MB-231, A549 and HCT116 cells and the indicated clones with knock-in expression of GPX4 S104A mutants were treated with or without 20 μM Erastin for 24 h. Lipid ROS-positive cells (i) and cell death (j) were measured respectively. Data are the mean ± SD (n = 3), *P < 0.01; **P < 0.001 by two-tailed Student’s t-test.
Extended Data Fig. 9 AKT-CKB-GPX4 axis suppresses ferroptosis.
(a) The whole cell lysates of LN18, LN229, U87 and U251 were harvested for immunoblotting analyses as indicated. (b, c) LN18, LN229, U87 and U251 cells were treated with or without 20 μM Erastin combined with Fer-1 (2 μM) for 24 h. Lipid ROS-positive cells (b) and cell death (c) were measured respectively. Data are the mean ± SD (n = 3), *P < 0.01; **P < 0.001; *** P < 0.0001 by two-tailed Student’s t-test. (d) U251 and U87 cells were treated with or without MK-2206 (5 μM) for 12 h. Immunoblotting analyses were performed with the indicated antibodies. (e, f) Parental U251 and U87 cells were treated with or without 20 μM Erastin in the absence or presence of MK-2206 (5 μM) for 24 h. Lipid ROS-positive cells (e) and cell death (f) were measured respectively. Data are the mean ± SD (n = 3), *P < 0.01; **P < 0.001 by two-tailed Student’s t-test.
Extended Data Fig. 10 CKB-mediated GPX4 phosphorylation promotes tumour growth.
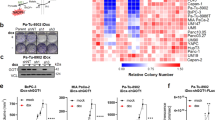
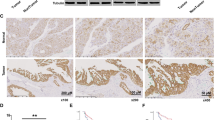
(a, b, e-i, k, l, n, o, q) Data are the mean ± SD, ^P < 0.05; *P < 0.01; **P < 0.001; ***P < 0.0001; N.S., not significant by two-tailed Student’s t-test (a, e, g, h, i, l, o) and by two-tailed Mann-Whitney U test (b, f, k, n, q). (b, f, j, m, p) IHC analyses of the indicated xenograft tumors from nude mice were performed with the indicated antibodies. Representative staining images and staining scores are shown. (a, b) Huh7 cells were subcutaneously injected into 6-week-old male athymic nude mice. When the tumor reached 50 mm3, the mice were assigned randomly into different treatment groups. Sulfasalazine (SAS) and Liproxstatin-1 (Lip-1) were intraperitoneally injected daily at a dose of 100 mg/kg and 10 mg/kg respectively until the endpoint at Day 28. Tumor volume and weight were analyzed (n = 7) (a). (c, r) The whole cell lysates of WT and GPX4 knockout Huh7 cells (c) and L02, THLE-2, Huh7, HCCLM3 (r) were harvested for immunoblotting analyses as indicated. (d-f) WT and GPX4 knockout Huh7 cells were subcutaneously injected into the left or right flanks of 6-week-old male athymic nude mice with and without daily Lip-1 intraperitoneal injection from the 4th day. The resulting tumors were resected 28 days after injection (d). Tumor volume and weight were analyzed (n = 6) (e). (g-q) IGF1R CA-expressing parental Huh7 cells and the indicated clones with knock-in expression of CKB T133A (g) and GPX4 S104A (h) or Huh7 cells expressing CKB shRNA with reconstituted expression of the indicated CKB proteins (i-k) or Parental Huh7 cells and the indicated clones with knock-in expression of GPX4 S104D mutants stably transfected with CKB shRNA and reconstituted with indicated CKB proteins (l-q) were intrahepatically (g, h) or subcutaneously (i-q) injected into 6-week-old male athymic nude mice (n = 7). When the tumor reached 50 mm3, the mice were assigned randomly into different treatment groups. SAS was intraperitoneally injected daily at a dose of 100 mg/kg until the endpoint at Day 28. Tumor volume and weight were analyzed.
Supplementary information
Supplementary Information
Supplementary Information
Source data
Source Data Fig. 1
Unprocessed WBs.
Source Data Fig. 2
Unprocessed WBs.
Source Data Fig. 3
Unprocessed WBs.
Source Data Extended Data Fig. 1
Unprocessed WBs.
Source Data Extended Data Fig. 2
Unprocessed WBs.
Source Data Extended Data Fig. 3
Unprocessed WBs.
Source Data Extended Data Fig. 4
Unprocessed WBs.
Source Data Extended Data Fig. 5
Unprocessed WBs.
Source Data Extended Data Fig. 8
Unprocessed WBs.
Source Data Extended Data Fig. 9
Unprocessed WBs.
Source Data Extended Data Fig. 10
Unprocessed WBs.
Source Data Fig. 1
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data for Extended Data Fig. 1.
Source Data Extended Data Fig. 2
Statistical source data for Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Statistical source data for Extended Data Fig. 3.
Source Data Extended Data Fig. 4
Statistical source data for Extended Data Fig. 4.
Source Data Extended Data Fig. 5
Statistical source data for Extended Data Fig. 5.
Source Data Extended Data Fig. 6
Statistical source data for Extended Data Fig. 6.
Source Data Extended Data Fig. 7
Statistical source data for Extended Data Fig. 7.
Source Data Extended Data Fig. 8
Statistical source data for Extended Data Fig. 8.
Source Data Extended Data Fig. 9
Statistical source data for Extended Data Fig. 9.
Source Data Extended Data Fig. 10
Statistical source data for Extended Data Fig. 10.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, K., Yan, M., Liu, T. et al. Creatine kinase B suppresses ferroptosis by phosphorylating GPX4 through a moonlighting function. Nat Cell Biol 25, 714–725 (2023). https://doi.org/10.1038/s41556-023-01133-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-023-01133-9
This article is cited by
-
Ferroptosis in cancer: From molecular mechanisms to therapeutic strategies
Signal Transduction and Targeted Therapy (2024)
-
The cell biology of ferroptosis
Nature Reviews Molecular Cell Biology (2024)
-
Modulation of Cellular Levels of Adenosine Phosphates and Creatine Phosphate in Cultured Primary Astrocytes
Neurochemical Research (2024)
-
A guideline on the molecular ecosystem regulating ferroptosis
Nature Cell Biology (2024)
-
The mechanism of ferroptosis and its related diseases
Molecular Biomedicine (2023)