Abstract
Vascular endothelial growth factor receptor type 2 (VEGFR2, also known as KDR and FLK1) signalling in endothelial cells (ECs) is essential for developmental and reparative angiogenesis. Reactive oxygen species and copper (Cu) are also involved in these processes. However, their inter-relationship is poorly understood. Evidence of the role of the endothelial Cu importer CTR1 (also known as SLC31A1) in VEGFR2 signalling and angiogenesis in vivo is lacking. Here, we show that CTR1 functions as a redox sensor to promote angiogenesis in ECs. CTR1-depleted ECs showed reduced VEGF-induced VEGFR2 signalling and angiogenic responses. Mechanistically, CTR1 was rapidly sulfenylated at Cys189 at its cytosolic C terminus after stimulation with VEGF, which induced CTR1–VEGFR2 disulfide bond formation and their co-internalization to early endosomes, driving sustained VEGFR2 signalling. In vivo, EC-specific Ctr1-deficient mice or CRISPR–Cas9-generated redox-dead Ctr1(C187A)-knockin mutant mice had impaired developmental and reparative angiogenesis. Thus, oxidation of CTR1 at Cys189 promotes VEGFR2 internalization and signalling to enhance angiogenesis. Our study uncovers an important mechanism for sensing reactive oxygen species through CTR1 to drive neovascularization.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All data supporting the findings of this study are available from the corresponding author on reasonable request. Source data are provided with this paper.
References
Simons, M., Gordon, E. & Claesson-Welsh, L. Mechanisms and regulation of endothelial VEGF receptor signalling. Nat. Rev. Mol. Cell Biol. 17, 611–625 (2016).
Simons, M. An inside view: VEGF receptor trafficking and signaling. Physiology 27, 213–222 (2012).
Eichmann, A. & Simons, M. VEGF signaling inside vascular endothelial cells and beyond. Curr. Opin. Cell Biol. 24, 188–193 (2012).
Lanahan, A. et al. The neuropilin 1 cytoplasmic domain is required for VEGF-A-dependent arteriogenesis. Dev. Cell 25, 156–168 (2013).
Sawamiphak, S. et al. Ephrin-B2 regulates VEGFR2 function in developmental and tumour angiogenesis. Nature 465, 487–491 (2010).
Genet, G. et al. Endophilin-A2 dependent VEGFR2 endocytosis promotes sprouting angiogenesis. Nat. Commun. 10, 2350 (2019).
Tang, X. et al. Endothelium-specific deletion of Nox4 delays retinal vascular development and mitigates pathological angiogenesis. Angiogenesis 24, 363–377 (2020).
Craige, S. M. et al. NADPH oxidase 4 promotes endothelial angiogenesis through endothelial nitric oxide synthase activation. Circulation 124, 731–740 (2011).
Evangelista, A. M., Thompson, M. D., Bolotina, V. M., Tong, X. & Cohen, R. A. Nox4- and Nox2-dependent oxidant production is required for VEGF-induced SERCA cysteine-674 S-glutathiolation and endothelial cell migration. Free Radic. Biol. Med. 53, 2327–2334 (2012).
Kim, Y. M. et al. ROS-induced ROS release orchestrated by Nox4, Nox2, and mitochondria in VEGF signaling and angiogenesis. Am. J. Physiol. Cell Physiol. 312, C749–C764 (2017).
Schroder, K. et al. Nox4 is a protective reactive oxygen species generating vascular NADPH oxidase. Circ. Res. 110, 1217–1225 (2012).
Tojo, T. et al. Role of gp91phox (Nox2)-containing NAD(P)H oxidase in angiogenesis in response to hindlimb ischemia. Circulation 111, 2347–2355 (2005).
Urao, N. et al. Critical role of endothelial hydrogen peroxide in post-ischemic neovascularization. PLoS ONE 8, e57618 (2013).
Ushio-Fukai, M. Redox signaling in angiogenesis: role of NADPH oxidase. Cardiovasc. Res. 71, 226–235 (2006).
Ushio-Fukai, M. VEGF signaling through NADPH oxidase-derived ROS. Antioxid. Redox Signal. 9, 731–739 (2007).
Ushio-Fukai, M. & Urao, N. Novel role of NADPH oxidase in angiogenesis and stem/progenitor cell function. Antioxid. Redox Signal. 11, 2517–2533 (2009).
Charles, R. L. et al. Protein sulfenation as a redox sensor: proteomics studies using a novel biotinylated dimedone analogue. Mol. Cell. Proteom. 6, 1473–1484 (2007).
Poole, L. B., Karplus, P. A. & Claiborne, A. Protein sulfenic acids in redox signaling. Annu. Rev. Pharmacol. Toxicol. 44, 325–347 (2004).
Poole, L. B. & Nelson, K. J. Discovering mechanisms of signaling-mediated cysteine oxidation. Curr. Opin. Chem. Biol. 12, 18–24 (2008).
Fukai, T., Ushio-Fukai, M. & Kaplan, J. H. Copper transporters and copper chaperones: roles in cardiovascular physiology and disease. Am. J. Physiol. Cell Physiol. 315, C186–C201 (2018).
Narayanan, G., R,, B. S., Vuyyuru, H., Muthuvel, B. & Konerirajapuram Natrajan, S. CTR1 silencing inhibits angiogenesis by limiting copper entry into endothelial cells. PLoS ONE 8, e71982 (2013).
Harris, E. D. A requirement for copper in angiogenesis. Nutr. Rev. 62, 60–64 (2004).
Brewer, G. J. Tetrathiomolybdate anticopper therapy for Wilson’s disease inhibits angiogenesis, fibrosis and inflammation. J. Cell. Mol. Med. 7, 11–20 (2003).
Kaplan, J. H. & Maryon, E. B. How mammalian cells acquire copper: an essential but potentially toxic metal. Biophys. J. 110, 7–13 (2016).
Maryon, E. B., Molloy, S. A., Ivy, K., Yu, H. & Kaplan, J. H. Rate and regulation of copper transport by human copper transporter 1 (hCTR1). J. Biol. Chem. 288, 18035–18046 (2013).
Eisses, J. F. & Kaplan, J. H. The mechanism of copper uptake mediated by human CTR1: a mutational analysis. J. Biol. Chem. 280, 37159–37168 (2005).
Puig, S., Lee, J., Lau, M. & Thiele, D. J. Biochemical and genetic analyses of yeast and human high affinity copper transporters suggest a conserved mechanism for copper uptake. J. Biol. Chem. 277, 26021–26030 (2002).
Clifford, R. J., Maryon, E. B. & Kaplan, J. H. Dynamic internalization and recycling of a metal ion transporter: Cu homeostasis and CTR1, the human Cu+ uptake system. J. Cell Sci. 129, 1711–1721 (2016).
Molloy, S. A. & Kaplan, J. H. Copper-dependent recycling of hCTR1, the human high affinity copper transporter. J. Biol. Chem. 284, 29704–29713 (2009).
Brady, D. C. et al. Copper is required for oncogenic BRAF signalling and tumorigenesis. Nature 509, 492–496 (2014).
Tsai, C. Y., Finley, J. C., Ali, S. S., Patel, H. H. & Howell, S. B. Copper influx transporter 1 is required for FGF, PDGF and EGF-induced MAPK signaling. Biochem. Pharmacol. 84, 1007–1013 (2012).
Turski, M. L. et al. A novel role for copper in Ras/mitogen-activated protein kinase signaling. Mol. Cell. Biol. 32, 1284–1295 (2012).
Chen, G. F. et al. Copper transport protein antioxidant-1 promotes inflammatory neovascularization via chaperone and transcription factor function. Sci. Rep. 5, 14780 (2015).
Gariano, R. F. & Gardner, T. W. Retinal angiogenesis in development and disease. Nature 438, 960–966 (2005).
Liu, Z. et al. Glycolysis links reciprocal activation of myeloid cells and endothelial cells in the retinal angiogenic niche. Sci. Transl. Med. 12, eaay1371 (2020).
Wang, Y. et al. Ephrin-B2 controls VEGF-induced angiogenesis and lymphangiogenesis. Nature 465, 483–486 (2010).
Das, A. et al. Endothelial antioxidant-1: a key mediator of copper-dependent wound healing in vivo. Sci. Rep. 6, 33783 (2016).
Kuo, Y. M., Zhou, B., Cosco, D. & Gitschier, J. The copper transporter CTR1 provides an essential function in mammalian embryonic development. Proc. Natl Acad. Sci. USA 98, 6836–6841 (2001).
Lee, J., Prohaska, J. R. & Thiele, D. J. Essential role for mammalian copper transporter Ctr1 in copper homeostasis and embryonic development. Proc. Natl Acad. Sci. USA 98, 6842–6847 (2001).
Kim, Y. M. et al. Redox regulation of mitochondrial fission protein Drp1 by protein disulfide isomerase limits endothelial senescence. Cell Rep. 23, 3565–3578 (2018).
Shanbhag, V. et al. ATP7A delivers copper to the lysyl oxidase family of enzymes and promotes tumorigenesis and metastasis. Proc. Natl Acad. Sci. USA 116, 6836–6841 (2019).
Baker, A. M. et al. Lysyl oxidase plays a critical role in endothelial cell stimulation to drive tumor angiogenesis. Cancer Res. 73, 583–594 (2013).
Ash, D. et al. The P-type ATPase transporter ATP7A promotes angiogenesis by limiting autophagic degradation of VEGFR2. Nat. Commun. 12, 3091 (2021).
Nelson, K. J. et al. Use of dimedone-based chemical probes for sulfenic acid detection methods to visualize and identify labeled proteins. Methods Enzymol. 473, 95–115 (2010).
Okur, M. N., Russo, A. & O’Bryan, J. P. Receptor tyrosine kinase ubiquitylation involves the dynamic regulation of Cbl-Spry2 by intersectin 1 and the Shp2 tyrosine phosphatase. Mol. Cell. Biol. 34, 271–279 (2014).
Kang, D. H. et al. Peroxiredoxin II is an essential antioxidant enzyme that prevents the oxidative inactivation of VEGF receptor-2 in vascular endothelial cells. Mol. Cell 44, 545–558 (2011).
Lee, J., Petris, M. J. & Thiele, D. J. Characterization of mouse embryonic cells deficient in the ctr1 high affinity copper transporter. Identification of a Ctr1-independent copper transport system. J. Biol. Chem. 277, 40253–40259 (2002).
Rezende, F. et al. Knock out of the NADPH oxidase Nox4 has no impact on life span in mice. Redox Biol. 11, 312–314 (2017).
Ashino, T. et al. Unexpected role of the copper transporter ATP7A in PDGF-induced vascular smooth muscle cell migration. Circ. Res. 107, 787–799 (2010).
Ikeda, S. et al. Novel role of ARF6 in vascular endothelial growth factor-induced signaling and angiogenesis. Circ. Res. 96, 467–475 (2005).
van Lessen, M. et al. Regulation of vascular endothelial growth factor receptor function in angiogenesis by numb and numb-like. Arterioscler. Thromb. Vasc. Biol. 35, 1815–1825 (2015).
Pan, Q. et al. Blocking neuropilin-1 function has an additive effect with anti-VEGF to inhibit tumor growth. Cancer Cell 11, 53–67 (2007).
Tsai, C. Y., Larson, C. A., Safaei, R. & Howell, S. B. Molecular modulation of the copper and cisplatin transport function of CTR1 and its interaction with IRS-4. Biochem. Pharmacol. 90, 379–387 (2014).
Lee, S., Howell, S. B. & Opella, S. J. NMR and mutagenesis of human copper transporter 1 (hCtr1) show that Cys-189 is required for correct folding and dimerization. Biochim. Biophys. Acta 1768, 3127–3134 (2007).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Clement, K. et al. CRISPResso2 provides accurate and rapid genome editing sequence analysis. Nat. Biotechnol. 37, 224–226 (2019).
Zhou, H. J. et al. SUMOylation of VEGFR2 regulates its intracellular trafficking and pathological angiogenesis. Nat. Commun. 9, 3303 (2018).
Pitulescu, M. E., Schmidt, I., Benedito, R. & Adams, R. H. Inducible gene targeting in the neonatal vasculature and analysis of retinal angiogenesis in mice. Nat. Protoc. 5, 1518–1534 (2010).
Lee, M., Choy, W. C. & Abid, M. R. Direct sensing of endothelial oxidants by vascular endothelial growth factor receptor-2 and c-Src. PLoS ONE 6, e28454 (2011).
Oshikawa, J. et al. Novel role of p66Shc in ROS-dependent VEGF signaling and angiogenesis in endothelial cells. Am. J. Physiol. Heart Circ. Physiol. 302, H724–H732 (2012).
Oshikawa, J. et al. Extracellular SOD-derived H2O2 promotes VEGF signaling in caveolae/lipid rafts and post-ischemic angiogenesis in mice. PLoS ONE 5, e10189 (2010).
Pasula, S. et al. Endothelial epsin deficiency decreases tumor growth by enhancing VEGF signaling. J. Clin. Invest. 122, 4424–4438 (2012).
Acknowledgements
We acknowledge X. Fang for assisting mouse genotyping in our initial study; and staff at the core facilities of the Genome Research Division within the Research Resource Center at the University of Illinois at Chicago for DNA sequencing and the Transgenic Mouse and Embryonic Stem Cell Facility at the University of Chicago for embryo injections. This work was supported by National Institute of Health grants R01HL135584 (to M.U.-F.), R01HL147550 (to M.U.-F. and T.F.), R01HL133613 (to T.F. and M.U.-F.), R01HL116976 (to T.F. and M.U.-F.), R01HL070187 (to T.F.), R35GM135179 (to L.B.P.), R01EY011766, R01EY030500 and R21EY032265 (to R.B.C.), 1K99EY029373-01A1 (to A.Y.F.); American Heart Association (AHA) grant 17POST33660754 (to D.A.); Veterans Administration Merit Review Award 2I01BX001232 (to T.F.), 101BX001233 (to R.B.C.); The VA Career Scientist Award IK6BX005228 (to R.B.C.). R.B.C. is the recipient of a Research Career Scientist Award from the Department of Veterans Affairs. The contents do not represent the views of the Department of Veterans Affairs or the United States Government. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
M.U.-F., T.F. and A.D. designed the study. A.D. (major), D.A., S.V., Y.H., Y.-M.K., B.K.S., A.Y.F., R.B.C., F.Z.H., R.L. and H.S. performed/assisted research. M.U.-F., T.F., A.D. and Y.H. analysed data. J.H.K. and L.B.P. discussed data and provided input. M.M. performed mouse genotyping. M.R.R. and B.J.M. generated the CRISPR–Cas9 KI mutant mice. J.H.K. and L.B.P. provided materials. M.U.-F., T.F. and A.D. wrote the manuscript. J.H.K. and L.B.P. edited the manuscript. All of the authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Donita Brady, Wang Min and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 CTR1 expression in endothelial cells and characterization of inducible Endothelial CTR1-KO mice.
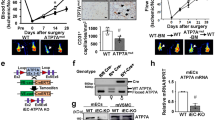
A. Co-staining for CTR1 and isolectin B4 (IB4) in a postnatal day (P)5 mouse retina in CTR1 WT and CTR1iECKO mice in developmental retina angiogenesis models. Arrows indicate the tip sprouts of vessels. B. Co-staining for CTR1 and CD31 (EC marker) or Mac-3 (macrophage marker) and their colocalization (merged, white arrows) on day 3 (upper) and day 14 (lower), respectively, in ischemic gastrocnemius muscles in hindlimb ischemia models. Representative images from n = 3 independent experiments are shown. C. Strategy to generate tamoxifen-inducible EC-specific CTR1 knockout (CTR1iECKO) mice by crossing CTR1flox/flox mice with VE-Cadherin (Cdh5)-ERT2 Cre delete mice, which specifically express Cre in ECs upon tamoxifen administration. D and E. mRNA and proteins from aortic ECs, liver and lung isolated from WT and CTR1iECKO mice and analyzed by qPCR and Western blotting using CTR1 antibody or Actin antibody (loading control) (n = 6 biologically independent cells/samples), two-tailed unpaired t-test, **p = 0.0085. F. Real time qPCR analysis of CTR1 mRNA in HUVECs transfected with control or CTR siRNAs. (n = 3 biologically independent experiments), two-tailed unpaired t-test, **p = 0.0023. NS = not significant. Data are mean ± SEM). Source numerical data and unprocessed blots are available in source data.
Extended Data Fig. 2 CTR1+/− mice show impaired postnatal developmental and reparative angiogenesis in vivo.
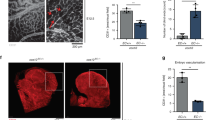
A. Retinal whole-mount staining with Isolectin B4 (IB4) of P5 WT and CTR1+/− mice. Right panels show quantification of vascular progression length, numbers of branch point and tip cells. (n = 6 samples each for WT and CTR1+/−, compared with Two-tailed unpaired t-test. *p = 0.0378). B and C. WT and CTR1+/− mice were used for skin wound healing model. Representative images show time-course for wounded skin and graph represents the wound closure rates expressed as % of wound area from control at day 0 after wounding (B). Wounded tissues at day 7 were used to measure CD31+ capillary density (C). Boxes showed magnified images. Right panel showed the quantification. (n = 6 mice per group, representative of two independent experiments, compared with two-way ANOVA followed by Bonferroni’s multiple comparison analysis (B) or two-tailed unpaired t-test (C)). D and E. Irradiated WT or CTR1+/− mice were transplanted with bone marrow (BM) from WT or CTR1+/− mice. After 6 weeks of BM transplantation (BMT), mice were subjected to hindlimb ischemia and limb blood flow was measured at indicated days after surgery using a laser Doppler flow analyzer (D). In E, CD31+ capillary density (angiogenesis) in ischemic and non-ischemic gastrocnemius muscles was measured at day 21 after surgery. (n = 6 mice per group, representative of two independent experiments, compared with two-way ANOVA followed by Bonferroni’s multiple comparison analysis (D), or Two-tailed unpaired t-test (E), ***p < 0.001. NS = not significant. Data are mean ± SEM). Source numerical data are available in source data.
Extended Data Fig. 3 Cu-dependent LOX activity is not required for VEGF-induced EC proliferation.
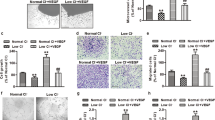
A and D. HUVECs treated with Cu chelator, TTM (20 nM) for 24 hr were used to measure LOX activity in conditioned media (A) or cell proliferation measured by BrdU incorporation (D) after VEGF stimulation for 24 hr. B. HUVECs treated with Cu chelator TTM for 24 hr were stimulated with CuCl2 (25 µM) for 5 min, and lysates were used to immunoblotted (IB) with anti-p-MEK1/2 or p-ERK1/2 and their total proteins antibodies. C. HUVECs treated with specific LOX inhibitor, BAPN (100 µM) for 24 h were used to measure VEGF-induced cell proliferation as described. (n = 3 biologically independent experiments). A and B, two-tailed unpaired t-test. C and D, one-way ANOVA followed by Tukey’s multiple comparisons analysis, *p < 0.05, **p < 0.01, ***p < 0.001. NS = not significant. Data are mean ± SEM. Source numerical data and unprocessed blots are available in source data.
Extended Data Fig. 4 CuCl2 does not induce CTR1 Sulfenylation.
HUVECs were stimulated with CuCl2 (25 µM) for indicated times or VEGF (20 ng/ml) for 5 min (for positive control), and DCP-Bio1-labelled lysates were pulled down with streptavidin beads and then IB with CTR1 or actin antibody to detect their CysOH formation. Bottom panel represents averaged CTR1-CysOH/total CTR1 level expressed by fold change from VEGF-induced CTR1-CysOH level as 1.0. (n = 3 biologically independent experiments) two-tailed unpaired t-test. ***p < 0.001. NS = not significant. Data are mean ± SEM. Source numerical data and unprocessed blots are available in source data.
Extended Data Fig. 5 Nox4-ROS-CTR1 Cys189OH axis is required for VEGF-induced VEGFR2 downstream signaling in ECs.
A, HUVECs transfected with Flag-hCTR1-WT, or Flag-hCTR1-C189A were infected with Ad.null (control) or Ad.shNox4 and stimulated with VEGF for 5 min to measure VEGF signaling using IB with antibodies indicated. Graphs represent the averaged fold change of phosphorylated proteins/total proteins over the basal control. (n = 3 biologically independent experiments) two-tailed unpaired t-test. ***p = 0.0008, *p = 0.013, ***P = 0.0002, **P = 0.0019, **P = 0.0019, **P = 0.0028. Data are mean ± SEM. B and C. HUVECs transfected with Flag-CTR1-WT or Flag-hCTR1-C189A or Flag-hCTR1-H190A were stimulated with VEGF (20 ng/ml) for 5 min to measure DCF fluorescence with DAPI staining (B). In C, lysates were used for IB with Flag antibody to verify expression of transfected CTR1 proteins. Tubulin is a loading control. B, C. The experiment is representative of 3 independent experiments that yielded similar results. Source numerical data and unprocessed blots are available in source data.
Extended Data Fig. 6 Generation of Cys oxidation defective ‘redox dead’ mouse mCtr1-C187A (corresponding to human CTR1-C189A) knock-in (KI) mutant (mCtr1-KI) mice by using CRISPR-Cas9 genome editing.
A. Alignment of partial amino acid sequences from human and mouse CTR1, indicating homology (boxes) between human Cys189 and mouse Cys187. Schematic of the 20-nucleotide sgRNA target sequence of the mCtr1 (blue) and the PAM (green). The red arrowhead indicates the Cas9 cleavage site. ssODN, which contains 90 base pairs (bp) of homology sequence flanking each side of the target site was used as HDR template. ssODN incorporates point mutations (red) and BamHI restriction enzyme site (underlined by black). B. CTR1 CRISPR mice genotyping for mCtr1C187A mutant and mCtr1WT after cross breeding with mCtr1KI/+ and mCtr1KI/+. Multiplex PCR genotyping of mCtr1 (C187A) progeny. One common reverse primer was used for both genotypes. The PCR products were in between 300-200 bp. HDR indicates homology directed repair; sgRNA, single-guide RNA; ssODN, single-strand oligoDNA. The experiment is representative of 6 independent experiments that yielded similar results. C. Body weight of WT, mCtr1KI/+ and mCtr1KI/KI mice. (n = 12 mice per group, compared with two-tailed unpaired t-test. NS = not significant. Data are mean ± SEM). Source numerical data are available in source data.
Extended Data Fig. 7 Ectopic expression of Flag-hCTR1-WT, Flag-hCTR1-C189A, or Flag-hCTR1-M154A in bovine aortic endothelial cells (BAECs) transfected with bovine siCont or siCTR1.
Lysates from Fig. 4b were used for IB with anti-Flag antibody to verify ectopic hCTR1 expression or Tubulin antibody (loading control)(A). RNA samples were used for real time qPCR analysis to measure bovine CTR1 mRNA (B). These data suggest successful knockdown of bovine CTR1 in BAEC with expression of various human CTR1 constructs. (n = 3 biologically independent experiments) two-tailed unpaired t-test. **p < 0.01. Data are mean ± SEM. Source numerical data and unprocessed blots are available in source data.
Extended Data Fig. 8 VEGF promotes internalization of CTR1 and VEGFR2 from cell surface in a dynamin- and VEGFR2-dependent manner but CuCl2 promotes CTR1 internalization not VEGFR2.
A, B, C. HUVECs stimulated with VEGF (20 ng/ml) for indicated times (A) or incubated with dynasore, a dynamin-associated endocytic inhibitor (200 nM) (B) or transfected with control or VEGFR2 siRNAs were stimulated with VEGF (20 ng/ml) for 30 min (C). Cells were labeled with cell surface biotinylation reagent, 1 mM EZ-Link Sulfo-NHS-LC-Biotin, followed by wash and then lysates were pulled down with streptavidin beads, followed by IB with antibodies indicated to detect cell surface CTR1 or VEGFR2 or Na,K-ATPase (cell surface marker). D. HUVECs stimulated with CuCl2 (25 µM) for indicated times were labeled with cell surface biotinylation reagent, 1 mM EZ-Link Sulfo-NHS-LC-Biotin. After wash, lysates were pulled down with streptavidin beads, followed by IB with antibodies indicated to detect cell surface CTR1 or VEGFR2 or Na,K-ATPase. Bottom panels represent the averaged cell surface CTR1 and VEGFR2 levels expressed as fold changes from the basal control. (n = 3 biologically independent experiments). A, B and D two-tailed unpaired t-test. C, one-way ANOVA followed by Tukey’s multiple comparisons analysis, *p < 0.05, **p < 0.01, ***p < 0.001. NS = not significant. Data are mean ± SEM). Source numerical data and unprocessed blots are available in source data.
Extended Data Fig. 9 CTR1 sulfenylation at Cys189 is required for VEGF-induced internalization of CTR1 and VEGFR2.
A. HUVECs were transfected with Flag-hCTR1-WT or Flag-hCTR1-C189A or Flag-hCTR1-H190A, cells stimulated with VEGF (20 ng/ml) for 30 min were used for measurement of cell surface CTR1 or VEGFR2 or Na/K ATPase using cell surface biotinylation assay, as in Extended Data Fig. 9. B. BAECs transfected with bovine sibovine siCTR1 or siControl, together with either Flag-hCTR1-WT, or Flag-hCTR1-C189A were stimulated with VEGF (20 ng/ml) for 30 mins. Cells were used to measure cell surface VEGFR2 or Na,K-ATPase protenin expression using cell surface biotinylation assay, as described.) (n = 3 biologically independent experiments). one-way ANOVA followed by Tukey’s multiple comparisons analysis, *p < 0.05, **p < 0.01, ***p < 0.001. NS = not significant. Data are mean ± SEM. Source numerical data and unprocessed blots are available in source data.
Extended Data Fig. 10 CTR1 sulfenylation in tissue resident cells is required for ischemia-induced angiogenesis in vivo.
A and B. Irradiated WT or mCTR1KI/KI mice were transplanted with BM from WT mice. After 6 weeks of BMT, mice were subjected to hindlimb ischemia and limb blood flow using a laser Doppler flow analyzer (A). CD31+ capillary density in ischemic gastrocnemius muscle at day 14 after ischemic injury were measured (B). (n = 6 mice per group, representative of two independent experiments, compared with two-way ANOVA followed by Bonferroni’s multiple comparison analysis (A) or two-tailed unpaired t-test (B), ***p < 0.001. Data are mean ± SEM). Source numerical data are available in source data.
Supplementary information
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 2
Unprocessed western blots and/or gels.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 3
Unprocessed western blots and/or gels.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed western blots and/or gels.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unprocessed western blots and/or gels.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 6
Unprocessed western blots and/or gels.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 8
Statistical source data.
Source Data Fig. 8
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 7
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 8
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 9
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Das, A., Ash, D., Fouda, A.Y. et al. Cysteine oxidation of copper transporter CTR1 drives VEGFR2 signalling and angiogenesis. Nat Cell Biol 24, 35–50 (2022). https://doi.org/10.1038/s41556-021-00822-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-021-00822-7
This article is cited by
-
Endothelial β-catenin upregulation and Y142 phosphorylation drive diabetic angiogenesis via upregulating KDR/HDAC9
Cell Communication and Signaling (2024)
-
NIR-IIb fluorescence antiangiogenesis copper nano-reaper for enhanced synergistic cancer therapy
Journal of Nanobiotechnology (2024)
-
Identification of a new gene signature for prognostic evaluation in cervical cancer: based on cuproptosis-associated angiogenesis and multi-omics analysis
Cancer Cell International (2024)
-
Understanding the molecular mechanism of regeneration through apoptosis-induced compensatory proliferation studies - updates and future aspects
Apoptosis (2024)
-
Exosomal circRNA: emerging insights into cancer progression and clinical application potential
Journal of Hematology & Oncology (2023)