Abstract
Loss of alveolar type 2 cells (AEC2s) and the ectopic appearance of basal cells in the alveoli characterize severe lung injuries such as idiopathic pulmonary fibrosis (IPF). Here we demonstrate that human alveolar type 2 cells (hAEC2s), unlike murine AEC2s, transdifferentiate into basal cells in response to fibrotic signalling in the lung mesenchyme, in vitro and in vivo. Single-cell analysis of normal hAEC2s and mesenchymal cells in organoid co-cultures revealed the emergence of pathologic fibroblasts and basaloid cells previously described in IPF. Transforming growth factor-β1 and anti-bone morphogenic protein signalling in the organoids promoted transdifferentiation. Trajectory and histologic analyses of both hAEC2-derived organoids and IPF epithelium indicated that hAEC2s transdifferentiate into basal cells through alveolar-basal intermediates that accumulate in proximity to pathologic CTHRC1hi/TGFB1hi fibroblasts. Our study indicates that hAEC2 loss and expansion of alveolar metaplastic basal cells in severe human lung injuries are causally connected through an hAEC2-basal cell lineage trajectory driven by aberrant mesenchyme.
Similar content being viewed by others
Main
Previous studies in mice have shown that mouse alveolar type 2 cells (mAEC2s) are the resident stem cell population in the alveoli that constitute the entire gas exchange surface of the lung1,2. In idiopathic pulmonary fibrosis (IPF), the most deadly and prevalent form of diffuse parenchymal lung disease, human alveolar type 2 cells (hAEC2s) are lost from the alveoli, concurrent with the appearance of metaplastic alveolar KRT5+ basal cells, which normally appear in the conducting airways3,4,5,6,7,8,9. Rigorous genetic lineage tracing has shown that metaplastic KRT5+ cells in the murine alveoli are not derived from mAEC2s, but rather from KRT5−/SOX2+ progenitors in the mouse airway after severe alveolar injury from fibrosis or viral infections5,6,10,11,12. However, it is not clear whether a similar population in the human airway exists that contributes to metaplastic basal cells, as the airways contain key anatomic differences across the two species13. This is a clinically relevant question, because the extent of alveolar KRT5+ basal cells directly correlates with mortality in IPF14. In this study, we made a surprising finding that hAEC2s, but not mAEC2s, can readily transdifferentiate into KRT5+ basal cells in organoid culture and xenotransplant. Moreover, we define pro-fibrotic mesenchymal niche-derived factors that promote hAEC2-to-basal cell transdifferentiation. Finally, quantitative spatial analysis of IPF lung tissue reveals that basal cells and advanced alveolar-basal intermediates are surrounded by aberrant, CTHRC1hi pro-fibrotic mesenchyme. These results identify hAEC2s as a source of metaplastic KRT5+ basal cells in severe alveolar injuries and provide a potential explanation for the reported appearance of aberrant hAEC2s with basaloid features in the transcriptomes of IPF and other severe lung injures such as COVID pneumonia8,9.
Results
hAEC2s transdifferentiate into basal cells in vitro
Previously reported hAEC2 organoids have utilized MRC5, a fetal human lung fibroblast cell line, as feeders to maintain primary adult hAEC2s in vitro, at least in part through local bone morphogenic protein (BMP) and Wnt signalling1,15. We confirmed similar organoid development using low-passage MRC5 cells in our three-dimensional (3D) organoid system and hAEC2s isolated with the HTII-280 antibody (EpCAM+/HTII-280+; Fig. 1b,c and Extended Data Fig. 1a), with high purity of the hAEC2s confirmed by single-cell RNA-sequencing (scRNA-seq) and cytospin analysis (Fig. 1a and Extended Data Fig. 1b,c). We hypothesized that primary mesenchyme obtained from normal adult lungs would provide a more supportive niche. We used a flow cytometry strategy to isolate adult human lung mesenchyme (AHLM; CD45−/CD11b−/CD31−/EpCAM−) that was previously described by scRNA-seq to yield a predominantly PDGFRα+ fibroblast population16 (Extended Data Fig. 1d), and co-cultured AHLM with hAEC2s in a 3D organoid. Although there was no difference in colony-forming capacity between MRC5 cells and AHLM, we saw a dramatic loss of Surfactant Protein-C (SFTPC) accompanied by the gradual appearance of KRT5, an airway basal cell marker, in hAEC2 organoids co-cultured with AHLM (Fig. 1d,e and Extended Data Fig. 1e). By day 14 of co-culture, the majority of the organoids derived from hAEC2–AHLM co-culture contained KRT5+ cells, whereas the majority of organoids in the hAEC2–MRC5 co-culture contained only SFTPC+ cells (Fig. 1c,e). Further immunophenotyping of KRT5+ organoids from hAEC2s co-cultured with either AHLM or MRC5 cells showed that most KRT5+ cells arising from MRC5 co-culture do not stain for other mature basal cell markers such as TP63 or KRT17 (Extended Data Fig. 1f,h), whereas the majority of KRT5+ cells derived from AEC2/AHLM organoids are also TP63+ and KRT17 (Extended Data Fig. 1g,h). Some of the organoids assume apical-basal polarity with KRT5+ basal cells in the outer layer and KRT5−/KRT8+ cells in the inner layer (Extended Data Fig. 1i), which have been identified as a transitional cell type derived from AEC2s17,18,19,20. To further ensure that the basal cells did not result from an outgrowth of contaminants in the HTII-280 sort, we cultured hAEC2s in previously defined mesenchymal-free, alveolar growth medium21 over a month to enrich for hAEC2s (cytospin shows >99% purity) before co-culture with AHLM, which resulted in the same basal cell differentiation as seen with freshly isolated hAEC2s (Extended Data Fig. 1k). Comparison of hAEC2 organoids utilizing passage 0 (freshly sorted) versus passage 2 from the same AHLM donor did not show any significant difference in KRT5 transdifferentiation (Extended Data Fig. 1j). A parallel experiment using mAEC2s co-cultured with primary adult murine lung mesenchyme (AMLM) under identical in vitro conditions produced no KRT5+ organoids (Extended Data Fig. 2a,b). Conversely, culturing hAEC2s with AMLM failed to generate organoids (Extended Data Fig. 2c).
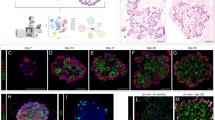
a, Live/EpCAM+/HTII-280+ cells (hAEC2s) were sorted and sequenced, confirming a purified SFTPC+ population. b,c, Immunofluorescence of hAEC2-derived organoids co-cultured with MRC5 cells (b) with quantification of epithelial lineages (c) found in the organoids (SFTPC+ organoids, organoids with both SFTPC+ cells and KRT5+ cells, or KRT5+ organoids). Representative of n = 3 biologically independent replicates. Data are presented as the mean of two replicates. d,e, Immunofluorescence of hAEC2-derived organoids co-cultured with AHLM (d) with quantification of the lineages of the organoids (e). Data are presented as the mean of n = 2 biological samples. Representative of n = 2 independent experiments. f, iPSC-derived AEC2s express tdTomato from one allele of SFTPC and can be flow purified based on their tdTomato expression. g, iAEC2s were co-cultured with either MRC5 cells or AHLM (both passage <5) with CK + DCI medium and organoids were analysed at day 14 for tdTomato expression. Experiments were performed in biological duplicates with AHLM collected from two independent donor lungs. h, Organoids from iAEC2s/MRC5 or iAEC2s/AHLM were collected and stained for alveolar (SFTPC), basal (KRT5 or KRT17) or intermediate (KRT8) markers. i,j, Ninety percent of iAEC2s/MRC5 organoids were SFTPC+/tdTomato+ (i), while ~65% of iAEC2s/AHLM organoids were SFTPClow, with the rest SFTPCneg (j). These organoids were also KRT5−/KRT17+/KRT8+, suggesting only partial differentiation. Representative of n = 2 independent experiments. k,l, iAEC2/AHLM organoids, when co-cultured with a 1:1 mixture of CK/DCI and modified MTEC media show complete transdifferentiation of tdTomato+ iAEC2s into tdTomato−/SFTPC−/KRT5+/NKX2-1+ (86.7% of analysed colonies) lung basal cells. The remaining 13.3% of organoids were SFTPC−/K5−/NKX2-1+. Representative of n = 2 independent experiments.
To further study the capacity of hAEC2s to differentiate into KRT5+ basal cells using a trackable differentiation system, we utilized a previously validated human induced pluripotent stem cell-derived AEC2 (iAEC2) model system22. We generated stably self-renewing iAEC2s in epithelial-only (feeder-free) cultures from the published SPC2 induced pluripotent stem cell (iPSC) line that has been previously engineered with a tdTomato reporter targeted to one allele of the endogenous SFTPC locus to isolate iAEC2s lacking basal cell transcripts (SPC2-ST-B2 clone, referred to as SPC2 cell line)23. As previously shown, we confirmed the purity of the iAEC2s, judged by the absence of basal cell markers and the maintenance of robust tdTomato expression (Fig. 1f). Next, we co-cultured iAEC2s with either MRC5 cells or AHLM in our previously published medium, optimized to maintain iAEC2 fate in culture22,23 (CHIR99021 (CHIR), Keratinocyte growth factor (KGF), dexamethasone, cyclic adenosine monophosphate (cAMP), Isobutylmethylxanthin (IBMX), hereafter called CK/DCI). In stark contrast to iAEC2s co-cultured with MRC5 cells, iAEC2s co-cultured with AHLM dramatically lost SFTPC expression, as evidenced by the loss of tdTomato fluorescence (Fig. 1g). Immunophenotyping confirmed the loss of SFTPC in iAEC2s co-cultured with AHLM in virtually all organoids, along with the emergence of intermediate and early basal cell markers KRT8 and KRT17, respectively, in ~65% of organoids (Fig. 1i,j). Interestingly, iAEC2s co-cultured with AHLM do not completely transdifferentiate into KRT5+ basal cells in CK/DCI. Further experiments revealed CHIR as the main medium component that prevents basal cell differentiation (Extended Data Fig. 2d,e) and, when CHIR was reduced by dilution, there was complete differentiation into basal cells that are also positive for KRT5, KRT14 and p63, while retaining NKX2-1, confirming lung fate (Fig. 1k,l and Extended Data Fig. 2f).
hAEC2s transdifferentiate to basal cells in a fibrotic host in vivo
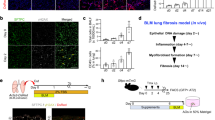
A previous transplant study of freshly sorted murine AEC2s showed no evidence of basal cell transdifferentiation when engrafted in injured lungs24. To determine whether the in vitro transdifferentiation capacity of hAEC2s is maintained in vivo, we transplanted freshly sorted hAEC2s into bleomycin-induced fibrotic lungs of NOD scid gamma (NSG) mice on day 10 (Fig. 2a). hAEC2s were transplanted either alone or with MRC5 or AHLM cells, analogous to ex vivo experiments with 3D organoids. Using the human-specific nuclear antigen (HNA), we were able to identify numerous patches of human cells predominantly in damaged alveolar regions 10 days after transplant (day 20 post bleomycin injury). Co-staining with KRT5 identified HNA+/KRT5+ cells that were SFTPC−, where the metaplastic basal cells appeared to form ‘bronchiolized’ cysts in the murine host (Fig. 2c). Utilizing an antibody specific to human-pro-SP-C, we also noted the presence of HNA+/pro-SP-C+ hAEC2 patches in the lung that were spatially distinct from dysplastic basal patches (Fig. 2b,c). Immunohistology analysis showed that, when transplanted alone or with MRC5 cells, the majority of the hAEC2s retained alveolar fate (~75%), whereas co-transplants of a similar AHLM:AEC2 ratio resulted in a majority of the patches comprising KRT5+ basal cells (~80%) (Fig. 2b,c). To determine whether transplanted AHLM and hAEC2s are associated in fibrotic lung, we labelled AHLM with a lentivirus expressing red fluorescent protein (RFP) followed by co-transplantation of hAEC2s and RFP-labelled AHLM. Interestingly, we found pods of hAEC2s with basaloid feature (SFTPC+/KRT5+/HNA+, Fig. 2d, green arrows) along with basal cells (SFTPC−/KRT5+/HNA+, green arrows) in regions adjacent to engrafted RFP+ AHLM (Fig. 2d, orange arrows). We also detected clusters of cells co-expressing AEC2s (SFTPC) and basal (KRT5) and transitioning (KRT8)17,19 cell markers, adjacent to AHLM (Fig. 2e). The xenotransplantation experiments demonstrate that hAEC2s are capable of reconstituting a fibrotic milieu in vivo and transdifferentiate into basal cells with high efficiency in the presence of AHLM, confirming the in vivo plasticity of hAEC2s.
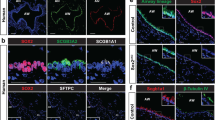
a, Experimental set-up for xenotransplant experiment testing engraftment of hAEC2s alone or co-transplanted with MRC5 or AHLM. b, hAEC2s engrafted in the injured lungs and differentiated towards either KRT5+ (green) basal cells or remaining as pro-SFTPC+ (white) hAEC2s, as judged by co-staining with HNA (red) in the lungs of mice transplanted with hAEC2s alone or hAEC2s + MRC5 or hAEC2s + AHLM. n = 3 mice per group, representative of n = 2 independent replicates. c, Co-transplantation of hAEC2s with MRC5 cells does not change the relative rates of hAEC2s engraftment events when compared with hAEC2s alone. However, co-transplantation of AHLM reduced the number of hAEC2 patches observed in the lungs of mice while increasing basal cell differentiation. Each dot represents the number of engrafted regions (>5 cells) in one section. At least two sections per mouse were analysed and n = 3 mice per group. Data are expressed as the mean ± s.d. The Kolmogorov–Smirnov test was used to determine normality and one-way analysis of variance (ANOVA) followed by the Kruskal–Wallis test was used for multiple comparisons. d, Immunofluorescence of engrafted hAEC2s actively transdifferentiating into KRT5+ basal cells in proximity to engrafted AHLM in the injured lung. Orange arrows, RFP-labelled AHLM; green arrows, KRT5+/SFTPC− basal cells; white arrows, SFTPC+/KRT5− hAEC2s; red arrows, KRT5+/SFTPC+ hybrid cells. n = 3 mice per group, representative of one independent experiment. e, Immunostaining of lungs transplanted with hAEC2s and tdTomato-labelled AHLM showed KRT8low/SFTPC+ hAEC2s (white arrows, inset), KRT8+/SFTPC+ intermediate-hAEC2s (white arrowhead, inset), KRT5+ basal cells (green arrows, inset) and rare SFTPC+/KRT5+ cells (red arrows, inset) adjacent to tdTomato-labelled AHLM (orange arrow) in areas of engraftment. Data are representative of n = 3 mice.
IPF-like mesenchymal subsets emerge during AHLM culture
hAEC2s co-cultured with IPF mesenchyme accelerated basal cell transdifferentiation compared to passage-matched AHLM from normal donors (Extended Data Fig. 3a,b). To determine whether AHLM acquires features of IPF mesenchymal subsets with culture, we analysed the scRNA-seq results for fresh (sorted from donor lung, uncultured) AHLM (normal donors, N = 3), fresh IPF mesenchyme (N = 3) and cultured AHLM at passage 0–1 isolated from organoids (N = 2). IPF lungs were collected from patients undergoing lung transplantation, and AHLM was collected from cadaveric donors without prior history of lung disease. Library preparation was performed separately and mesenchymal cells were segregated based on PDGFRA expression and merged for uniform manifold approximation and projection (UMAP) and clustering into distinct mesenchymal subsets (Fig. 3a). UMAP analysis showed fairly consistent clustering of subsets across AHLM and IPF donors, respectively (Extended Data Fig. 3c). Fresh AHLM consists mainly of proximal (advential), and distal (alveolar) fibroblasts16, along with a small population of smooth muscle (SM; Fig. 3c). However, cultured AHLM underwent a dramatic shift in cellular identity, with the loss of distal alveolar fibroblast identity similar to IPF mesenchyme (Fig. 3c, blue box and Supplementary Table 1). Instead, a CTHRC1-high (CTHRC1hi) fibroblast cluster and a separate HAS-high (HAShi) cluster emerged in cultured AHLM that were rare in fresh AHLM but enriched in IPF (Fig. 3b, red boxes in Fig. 3c and Supplementary Table 1). The merged UMAP results demonstrate a clear overlap in the identities of the CTHRC1hi and HAShi fibroblast subsets derived from fresh IPF and cultured AHLM (Fig. 3c). The emergence of these new subsets in culture was very interesting, because both HAShi (ref. 8) and CTHRC1hi (ref. 25) fibroblast subsets have been previously identified as fibroblast subsets in single-cell analysis of IPF lungs. The analysis of differentially expressed genes (DEGs) demonstrated highly concordant marker expressions in the pathologic mesenchymal niche subsets in cultured AHLM and IPF mesenchyme (Fig. 3d, Extended Data Fig. 3d and Supplementary Table 1). Both CTHRC1hi and HAShi subsets arising in cultured AHLM and IPF display features of fibrotic transformation. The CTHRC1hi fibroblast subset is characterized by high expression of collagen along with TGFB1 (Fig. 3b,d,e), and the HAShi fibroblast subset is characterized by the elevated expression of secreted BMP antagonists that have previously been shown to be elevated in IPF lungs26 (Fig. 3b,d,f). Analysis of immune subsets in IPF lungs shows that TGFB1 is also highly expressed in the macrophage population found in fibrotic lungs (Extended Data Fig. 3e–g). This analysis shows that pathologic fibroblast subsets emerge from cultured AHLM that are present in IPF, and they express niche factors that could potentially promote metaplastic transdifferentiation of hAEC2s into basal cells.
a, scRNA-seq analysis of fresh (sorted from donor lung, uncultured) AHLM (normal donors, N = 3, marked as AHLM fresh), fresh IPF mesenchyme (N = 3, marked as IPF fresh) and cultured AHLM at passage 0–1 isolated from organoids (N = 2). b, Signature genes in CTHRC1+ and HAShi/BMP antagonisthi clusters. c, scRNA-seq analysis showing a decrease of the distal (alveolar) population (blue box) and increase of CTHRC1+ and HAShi/BMP antagonisthi populations (red boxes) in both fresh IPF mesenchyme and cultured AHLM. d, Heatmaps showing the signature genes of each cluster in fresh IPF mesenchyme and cultured AHLM. e, Violin plots showing the signature genes in the CTHRC1+ fibroblasts among AHLM cultured, IPF fresh and AHLM fresh. f, Violin plots showing the BMP antagonists in the HAShi/BMP antagonisthi fibroblasts among AHLM cultured, IPF fresh and AHLM fresh.
Pathological niche signalling modulates transdifferentiation
To further define niche factors promoting hAEC2 transdifferentiation, we examined the transcriptome of cultured AHLM versus MRC5 cells by bulk RNA-seq. DEG analysis shows significant upregulation of multiple secreted BMP antagonists in the AHLM along with transforming growth factor-β (TGF-β) ligands, whereas BMP ligands and hedgehog antagonist, HHIP, were preferentially upregulated in MRC5 cells (Fig. 4a and Supplementary Table 2). HHIP is a secreted antagonist of Sonic Hedgehog (SHH) that has previously been identified in both hAEC2s and mesenchymal niche cells27, and its expression is reduced in IPF8. Hedgehog (Hh) activation can manipulate BMP signalling to modify basal cell metaplasia in murine models26, and addition of recombinant HHIP to hAEC2 organoids co-cultured with AHLM significantly attenuated KRT5 expression while increasing SFTPC expression in the organoids (Fig. 4b), as confirmed by immunophenotyping of organoid sections (Fig. 4c). HHIP treatment attenuated expression of the Hh transcriptional activator, GLI1, in AHLM co-cultured with hAEC2s, concurrent with an increase in the expression of BMP ligands (Fig. 4d). Similar to HHIP, addition of recombinant BMP4 significantly attenuated hAEC2 transdifferentiation into basal cells (Fig. 4e,f) and other airway lineages such as club cells (Extended Data Fig. 4a). Conversely, addition of DMH1 (a small-molecule inhibitor of BMP activation) and recombinant TGF-β1 both augmented the number of basal cells derived from hAEC2s co-cultured with MRC5 cells (Fig. 4g,h). This demonstrates that active TGF-β signalling promotes hAEC2 transdifferentiation into basal cells, while BMP activation in the hAEC2 niche maintains AEC2 fate, which could account for the differential effects of AHLM versus MRC5 cells in their ability to promote basal cell differentiation.
a, Bulk RNA-seq analysis of AHLM and MRC5 cells with DEG analysis. Significantly differentially expressed genes (Padj < 0.01, log2FC < −0.5 or >0.5; FC, fold change) are marked as red dots and are otherwise in grey. b, Treatment of hAEC2s + AHLM organoids with 2.5 μg ml−1 HHIP increases SFTPC and decreases KRT5 expression in the epithelial fractions at day 14 of co-culture. The expression levels of SFTPC and KRT5 in the epithelial fractions of hAEC2s + MRC5 organoids were used as controls. Data are expressed as the mean ± s.d. SFTPC, *P = 0.0399 and **P = 0.0028; KRT5, *P = 0.0294 and **P = 0.0015. c, Immunofluorescence analysis of HHIP-treated hAEC2s + AHLM organoids at day 14, quantified by the percentage of SFTPC+, KRT5+ and SFTPC+/KRT5+ cells in total cells. Data are expressed as the mean ± s.d. **P = 0.0023. d, HHIP-treated AHLM isolated from day-14 organoid co-culture demonstrates decreased GLI1 and increased BMP3 and BMP4 mRNA levels. Data are expressed as the mean ± s.d. *P = 0.0354, **P = 0.0043 and ***P = 0.0002. e, Immunofluorescence analysis and quantification of the hAEC2s + AHLM organoids at day 14 treated with 50 ng ml−1 BMP4. Data are expressed as the mean ± s.d. *P = 0.0142 and ***P = 0.0005. f, Treatment of hAEC2s + AHLM organoids with BMP4 increases SFTPC and decreases KRT5 mRNA expression in the epithelial fractions at day 14 of co-culture. The expression levels of SFTPC and KRT5 in the epithelial fractions of hAEC2s + MRC5 organoids were used as controls. Data are expressed as the mean ± s.d. SFTPC, *P = 0.0358 and **P = 0.0066; KRT5, **P = 0.0062 (left) and **P = 0.0028 (right). g,h, Immunofluorescence analysis (g) and quantification (h) of day-14 hAEC2s + AHLM organoids treated with DMH-I (1 µM) or TGF-β1 (3 ng ml−1). DMH-I treatment gives rise to either SFTPC+ alveolospheres (g, middle panel inset, 24% SFTPC+ AEC2s) or KRT5+/SFTPC+ basal/AEC2 mixed colonies (35% KRT5+ basal cells). TGF-β1 treatment drives more efficient basal cell transdifferentiation by suppressing alveolar fate (10% SFTPC+ AEC2s) and induces basal fate (57% KRT5+ basal cells). The experiment was performed in a technical triplicate and data from three technical replicates are counted as one biological replicate. *P < 0.05 and **P < 0.01, as determined by unpaired two-tailed Student’s t-test. Each data point represents one technical replicate.
hAEC2-derived basal cells resemble IPF metaplastic basal cells
There is increased recognition of basal cell heterogeneity in normal and diseased lungs, with specific basal cell subsets increased in IPF28. To compare hAEC2-derived basal cells in organoids with basal cells isolated from intact lungs, we analysed the scRNA-seq results of freshly isolated epithelial cells isolated from a distal fragment (not from the trachea) of normal donor lungs (N = 2), IPF lungs (N = 2) and hAEC2-derived basal cells isolated from organoids (N = 2). Library preparation was performed separately, and basal cells were segregated based on KRT5 expression and merged for UMAP and clustering into distinct basal subsets (Fig. 5a and Extended Data Fig. 5a). Clustering revealed four distinct subsets of basal cells with a high degree of gene overlap with previously identified basal cell subsets in vivo28, including secretory primed (SPB), multipotent (MPB), activated (AB) and proliferating (PB) (Fig. 5a–c and Extended Data Fig. 5b). Interestingly, the plurality of hAEC2-derived basal cells are secretory primed basal cells (Fig. 5a and Extended Data Fig. 5c, red box), a population that has been identified as the predominant basal population in the alveoli of IPF lungs28. Basal cell subset-specific marker analysis shows similarities in the expression of genes in hAEC2-derived and IPF basal cells (Fig. 5c).
a, scRNA-seq analysis of basal cells from fresh normal donors (N = 2, marked as NL basal), fresh patients with IPF (N = 2, marked as IPF basal) and hAEC2 + AHLM organoids (N = 2). b, Signature genes in the SPB, MPB, AB and PB clusters. c, Heatmaps showing the signature genes of each cluster in hAEC2-derived basal and IPF basal cells. d, Differential gene expression analysis of hAEC2-derived basal versus IPF basal cells. e, Immunofluorescence analysis of KRT14, MMP7 and KRT5 in normal lung, IPF lung and hAEC2-derived organoids at day 14. Representative of n = 2 independent experiments.
To explore further whether IPF biomarkers are upregulated in hAEC2-derived basal cells, we performed DEG analysis comparing basal cells from normal lung versus hAEC2-derived basal cells. This analysis showed that although hAEC2-derived basal cells express similar canonical basal markers such as SOX2, NGFR and TP63, they over-express markers previously reported to be upregulated in IPF epithelium, such as KRT14, VIM and MMP73,4,8,9,29 (Fig. 5d). Histologic comparison of normal lungs, IPF lungs and hAEC2-derived organoids shows that these IPF biomarkers are preferentially present in basal cells from IPF and hAEC2-derived organoids, but not normal lungs (Fig. 5e). Finally, to identify a common gene signature of IPF and hAEC2-derived basal cells, we performed an overlap analysis of DEGs of IPF basal cells and hAEC2-derived basal cells compared with normal lung basal cells. Hypergeometric probability testing shows significant enrichment of overlap genes in DEGs between hAEC2-derived and IPF basal cells (Extended Data Fig. 5d), generating a large list of DEGs that are shared between IPF and hAEC2-derived basal cells (Extended Data Fig. 5e and Supplementary Table 3). These results demonstrate that, although hAEC2-derived basal cells express canonical basal cell gene programs, they share more similarities with IPF basal cells ectopically localized in the diseased alveoli.
hAEC2 to basal cell transdifferentiation occurs through intermediate states
Recently, AEC2-derived intermediate cell types have been described in the alveolar space of bleomycin-challenged mice and IPF lungs8,9,17,19. To determine whether we observe similar discrete intermediates in the progressive transdifferentiation of hAEC2s toward basal cells in organoids, we analysed the time course of all epithelial cells (based on EPCAM expression) undergoing transdifferentiation in vitro by scRNA-seq. Epithelial cells from day-0 (live sort of HTII-280+ before culture; Fig. 1a and Extended Data Fig. 1a,b), day-7 (seven days after co-culture), day-14 and day-21 organoids were sequenced separately, then merged and clustered by UMAP, whereby six main clusters emerged (Fig. 6a, Extended Data Fig. 6a and Supplementary Table 4). Four clusters were easily identified by lineage markers of hAEC2s, basal cells, club cells and ciliated cells, respectively (Extended Data Fig. 6a,c,d). The time course shows a time-dependent loss of the hAEC2 population and gain of the KRT5+ basal cell population, along with an early emergence of two distinct alveolar-basal intermediates (ABIs) bearing both hAEC2 and basal markers at day 7 that recede with time in culture (Fig. 6a, highlighted by a dashed line). ABI1 cells are identified by persistence of SFTPC, ABCA3 and NAPSA, along with the emergence of KRT17 (Fig. 6i and Extended Data Fig. 6e). ABI2 cells are identified by low levels of AEC2 markers, higher levels of KRT17 and absence of KRT5 (Fig. 6i and Extended Data Fig. 6e). Both these intermediates express high levels of KRT8 (Fig. 6i and Extended Data Fig. 6i), a marker of intermediate cell types, described previously17,19. Of note, we observed a SFTPClow/KRT8+/KRT5− population comparable to ABIs in the epithelial cells derived from hAEC2s in vivo after engraftments in mice (Fig. 2d). Concurrent with the disappearance of the ABIs over time, there is also a progressive emergence of SCGB1A1+ club cells and a FOXJ1+ ciliated population in culture (Fig. 6a and Extended Data Fig. 6c,d), suggesting that transdifferentiated KRT5+ cells are mature basal cells capable of further differentiation into club and ciliated lineages.
a, scRNA-seq of freshly sorted HTII-280+ hAEC2s (day 0) and hAEC2s co-cultured with AHLM at days 7, 14 and 21. b–d, Pseudotime ordering of epithelial cells from day-0, 7, 14 and 21 organoids (b,c) identifies an hAEC2s → ABIs → basal → club → ciliated cell trajectory based on gene expression levels over pseudotime trajectory that was confirmed via RNA velocity (d). e, Immunofluorescence staining of day-7 hAEC2 + AHLM organoids identifies the presence of KRT17−/KRT8high/SFTPC+ ABI1 (14%) and KRT17+/KRT8+/SFTPClow ABI2s (23%) along with SFTPC+ hAEC2s (43%) and KRT17+/KRT5+ basal cells (9%). Representative of n = 2 biologically independent experiments, each performed in technical triplicate. Data are presented as the mean of biological replicates. f, IPA of the DEGs between AEC2s, ABI1 and ABI2s identifies the upregulation of alveolar signalling (NKX2-1, ETV5) in AEC2s, WNT and BMP signalling in AEC2s and ABI1 and fibrotic/airway signalling pathways (TGF-β1, HES1, extracellular matrix genes, SOX2 and TP63) in ABI2s. g,h, Re-analysis of IPF epithelium from Habermann et al.8 (g) to include basal cells along with PATS-like cells (encompassing transitional AEC2s and KRT17+/KRT5− cells on UMAP) with pseudotime ordering (h). i, Heatmaps comparing lineage-specific gene expression of KRT17+/KRT5− basaloid cells in IPF (labelled as PATS-like cells by Kobayashi et al.20) and ABIs in a 3D organoid.
The emergence of ABI1 and ABI2 intermediate populations in the direct line of transdifferentiation to basal cells was confirmed by both Monocle and RNA trajectory analysis (Fig. 6b−d). Visualization of gene expression along the hAEC2 transdifferentiation trajectory shows that the well-established AEC2 marker, LAMP3, and the new hAEC2 marker, HHIP27, are quickly lost first as mature hAEC2s transition into ABIs (Fig. 6c and Extended Data Fig. 6a,b). The emergence of the early basal marker KRT17 and the transitional-state marker KRT817,18,19,20, concurrent with the persistence of AEC2 markers SFTPC, ABCA3 and NAPSA, marks the ABI1 populations seen early in culture (Extended Data Fig. 6b–d). The presence of the ABI1 and ABI2 is confirmed on immunophenotyping of day-7 organoids early in the transdifferentiation, where we see KRT17−/KRT8+/SFTPC+ and KRT17+/KRT8+/SFTPClow cells, in addition to a minority of organoids (9%) that contain only completely transdifferentiated basal cells (KRT17+/KRT5+; Fig. 6e). As transdifferentiation progresses, the ABIs give way to mature basal cells, as all hAEC2 markers are lost with the emergence of mature basal cell markers KRT5 and KRT14 (Fig. 6c,i and Extended Data Fig. 6a,e,i). An upstream IPA analysis of pathways driving hAEC2s, ABI1 and ABI2s confirmed the distinct expression profiles of these intermediates. ABI1 cells exhibited active upstream drivers typical of hAEC2s (FOXA2, GATA4, NKX2-1) relative to ABI2s (Fig. 6f). Conversely, the ABI2s are preferentially activated by basal cell pathways (TP63, HES1, SOX2) along with TGF-β1 signalling and expression of mesenchymal markers (Fig. 6f and Supplementary Table 5).
Identification of AEC2 to basal cell ABIs in fibrotic human lungs
Several recent studies in mice have reported that mAEC2s undergo a transitional state when differentiating into mAEC1s in vitro and in vivo18,19,20. The pre-alveolar type 1 transitional cells (PATS) described in mouse20 were equated with both the transitional AEC2s and KRT17+/KRT5− basaloid cells described earlier8,9 and labelled as ‘PATS-like cells’ in human. However, basal cells were omitted from the original analysis20, even though the KRT17+/KRT5− basaloid cells express low levels of canonical basal cell transcription factor TP638,20. Therefore, we hypothesized that PATS-like cells, inclusive of transitional AEC2s and KRT17+/KRT5− basaloid cells, also represent intermediate cells between AEC2s and basal cells. To this end, we re-analysed IPF epithelial transcriptomes to include basal cells from the original dataset (from Haberman et al.8, analysed by Kobayashi et al.20), and then performed Monocle trajectory analysis (Fig. 6g,h). This analysis confirmed an hAEC2–hAEC1 trajectory through transitional AEC2s. However, inclusion of basal cells in the analysis also revealed an equally prominent hAEC2-to-basal cell trajectory through the KRT17+/KRT5− basaloid cells (Fig. 6g,h). We then directly compared the genes found to be upregulated in PATS-like cells in IPF20 with the organoid ABIs (ABI1 and ABI2) and observed an almost identical expression pattern (Fig. 6i and Extended Data Fig. 6f−i). The organoid ABI1 population appears similar to the transitional AEC2s, while the organoid ABI2 population appears similar to the KRT17+/KRT5− basaloid cells (Fig. 6i). Together, these findings suggest that ABIs derived from hAEC2s in our 3D organoid culture recapitulate previously identified intermediate cell populations in IPF lungs and suggest a direct trajectory from hAEC2s to basal cells through these intermediate cell types.
Consistent with our in silico prediction of a continuous trajectory from hAEC2s to ABIs to basal cells, we found evidence of all four cell types in the same cystic region (Fig. 7a and Extended Data Fig. 7a), as judged by protein staining for hAEC2s (SFTPC+/KRT8low/KRT17−/KRT5−), ABI1 (SFTPC+/KRT8hi/KRT17low/KRT5−), ABI2 (SFTPClow/KRT8hi/KRT17+/KRT5−) and basal cells (SFTPC−/KRT8−/KRT17+/KRT5+). Furthermore, ABI2s can be subdivided based on the presence of TP63, as we found several examples of both TP63+/KRT17+/KRT5− and TP63−/KRT17+/KRT5− ABI2s in the lungs of patients with IPF (Extended Data Fig. 7a), comparable to previously described basaloid (KRT17+/KRT5− cells that are either TP63+ or TP63−) populations in IPF8,9,20. Next, we systematically inspected IPF lungs for the presence of ABIs by RNA in situ hybridization as a function of the extent of alveolar remodelling using a combination of markers for ABI1, ABI2 and basal cells, all of which express varying levels of KRT17 messenger RNA (mRNA). As expected, ABI1 or ABI2 were extremely rare in the alveolar regions of normal lung (Extended Data Fig. 7b). Considering the highly heterogeneous histology of an IPF lung, we focused our analysis of ABIs in histologically defined regions (Fig. 7b). The relative frequencies of ABI1s and ABI2s correlated with progression of disease. In the normal-appearing and thickened alveolar regions of IPF lungs reflecting histological regions of minor injury, >80% of the KRT17+ cells were SFTPC+ ABI1s, and the remainder were KRT17+/SFTPC-/KRT5− ABI2s (Fig. 7c). Conversely, in areas of moderate to severe injury reflected by the presence of microcystic regions, the majority of KRT17+ cells were ABI2s (63%) and with emergence of a small number of KRT17+/KRT5+ basal cells (10% of KRT17+ cells) (Fig. 7c). Finally, in areas with complete bronchiolization and heavy fibrosis, we found few ABI1s (2%) and ABI2s (30%), with 68% of KRT17+ cells now acquiring complete basal identity, as judged by KRT5 expression (Fig. 7c). Furthermore, by immunostaining we found evidence of ABIs in lungs with scleroderma and COVID-19, showing linearly connected AEC2s (SFTPC+), ABI1s (SFTPC+/KRT17+), ABI2s (SFTPC−/KRT17+/KRT5−) and basal cells (KRT17+/KRT5+) cells in the same area of the disease lung (Fig. 7d and Extended Data Fig. 7c). The results strongly suggest that ABI populations in vivo, similar to the population identified in our 3D organoids, appear as a function of alveolar injuries that result in transdifferentiation of hAEC2s to basal cells.
a, Immunostaining of IPF lungs finds hAEC2s (SFTPC+), ABI1 (SFTPC+/KRT17low), ABI2 (SFTPClow/KRT17+/KRT5−) and basal cells (KRT5+/KRT17+) in the same cystic regions. The heatmap shows the average expression of select candidate genes in each cell type in organoids. Representative of n = 3 IPF specimens. b,c, In situ analysis of IPF lungs identifying decreasing frequency of ABI1s (SFTPC+/KRT17+; orange arrows) and increasing frequency of ABI2s (SFTPC−/KRT17+/KRT5−; red arrows) and basal cells (SFTPC−/KRT17+/KRT5+; white arrows) as the severity of fibrosis increases. Green arrows indicate AEC2s (SFTPC+/KRT17−) (b). Representative of n = 6 independent IPF specimens used to define normal, thickened, microcystic and bronchiolized regions. Each region included at least four images taken at ×20 from at least three IPF specimens. Data in c are presented as the average of all the images and error bars show s.d. d, Immunostaining of scleroderma lung identifying linear progression from hAEC2s → ABI1 → ABI2 → basal cells in the same region. Images are representative of n = 1 scleroderma lung.
Analysis of AHLM co-cultured with hAEC2s in our organoid platform identified a TGFB1hi mesenchymal population that is marked by CTHRC1 gene expression, which has been shown previously to be associated with fibroblastic foci25. Because ABI2s have elevated TGF-β signalling (Fig. 6f), which can antagonize BMP signalling30,31, we tested the hypothesis that aberrant mesenchyme providing a pro-fibrotic TGFB1hi niche is spatially associated with ABIs. We probed IPF lung specimens in situ with ABI markers (SFTPC and KRT17) and the pro-fibrotic mesenchymal marker CTHRC1, followed by protein staining for basal marker KRT5 (Fig. 8a). Consistent with our hypothesis, ~10% of ABI1s were adjacent to CTHRC1+ cells, but 83% of ABI2s were found adjacent to CTHRC1+ mesenchyme, suggesting a high degree of association between metaplastic intermediate cells and TGFβ1hi pro-fibrotic mesenchymal cells (Fig. 8b). We also found CTHRC1+ mesenchyme to be also associated with 77% of all basal cells found in actively remodelling regions (Fig. 8b). These data indicate that hAEC2s transdifferentiate into metaplastic basal cells through a series of intermediates, at least in part in a fibrotic niche made of TGFβnhi, pro-fibrotic mesenchymal populations (Fig. 8c).
a,b, RNA in situ hybridization was performed for SFTPC, KRT17 and CTHRC1 followed by KRT5 protein staining (a). 83% of ABI2s (SFTPC−/KRT17+/KRT5−) and 77% of basal cells (KRT17+/KRT5+) found in actively remodelling regions were in close proximity to clusters of CTHRC1high fibroblasts (b). Representative of n = 3 independent IPF lungs analysed. Each dot in b represents an average of at least a 3 × 3 tiled region captured at ×20. In total, 160 images were captured at ×20 and counted across three IPF specimens. Data are presented as mean ± s.d. Significance is calculated by one-way ANOVA followed by the Kruskal–Wallis test for multiple comparisons: **P = 0.0014 and ****P < 0.0001. c, Schematic illustrating a transdifferentiation pathway of hAEC2s to metaplastic basal cells through discrete ABIs and as a likely consequence the accumulation of the dysplastic alveolar structures prominent in IPF. Transdifferentiation develops as a function of the transition of surrounding mesenchyme from a BMP/Wnt high supportive niche (green) to an aberrant, pro-fibrotic niche composed of fibroblasts high in anti-BMPs, TGF-β1 and extracellular matrix (ECM) proteins (red). Although the schematic highlights active TGF-β1 deriving from fibroblasts, ABIs are also a prominent source of TGF-β1 activation and signalling (Fig. 6 and refs. 20 and 19), probably contributing to the development of aberrant mesenchyme and a pro-fibrotic milieu.
Discussion
There is currently an assumption that the regenerative capacity of AEC2s, the resident stem cell of the alveolus, is highly conserved between mouse and human32. However, the unexpected finding that, unlike mAEC2s, hAEC2s robustly transdifferentiate into functional basal cells with cues from pathological mesenchymal cells arising in vitro invited the possibility that a parallel process occurs in human diseases characterized by severe alveolar injuries in vivo (for example, IPF and COVID-19). Single-cell analysis of the cultured mesenchymal niche cells responsible for the transdifferentiation (AHLM) demonstrated the emergence of two fibroblast subpopulations recently identified in freshly isolated IPF lungs that are enriched in either secreted TGF-β1 (CTHRC1hi) or BMP antagonists (HAShi). Similarly, single-cell analysis of the hAEC2-to-basal cell trajectory in vitro revealed the presence of intermediate cell types and basal cell subsets previously identified in IPF lungs. These findings validate our AHLM/hAEC2 organoid assay as an in vitro model for hAEC2-mesenchymal crosstalk that results in stem cell metaplasia seen in severe alveolar injury, and provides experimental confirmation of a stem cell trajectory that is seen in diseased human, but not murine lung.
Another group of recent studies have also identified an intermediate cell state during AEC2 to AEC1 differentiation, first described in mice, that is marked by elevated expression of KRT8 and increased TGF-β1 signalling17,19,20. These murine intermediate cell types bear a striking resemblance to the human ABIs identified in our study, suggesting that a KRT8 high intermediate state may not be unique during AEC2-to-AEC1 transdifferentiation in mice, but also evolved to include AEC2-to-basal transdifferentiation in human. Furthermore, ABI1 and ABI2 found in vitro are remarkably similar to the discrete intermediate cell types—that is, transitional AEC2s and KRT5−/KRT17+ aberrant basaloid cells—described in IPF lungs. A comparative IPA analysis of hAEC2, ABI1 and ABI2 transcriptomes (Fig. 6f) reveals the extensive transcriptional reprogramming involved in the transdifferentiation through these intermediate states, including progressive loss of AEC2-promoting signalling, such as NKX2-1, and the emergence of numerous activities promoting mesenchymal features, especially the strong activation of TGF-β signalling in ABI2s. We focused on TGF-β1, because TGF-β1 promotes hAEC2 transdifferentiation to basal cells in organoids, and CTHRC1hi fibroblasts known to be TGFB1hi emerge both in our organoid assay and in IPF tissues. Indeed, over 80% of the ABI2 clusters and 67% of alveolar basal cells were found adjacent to CTHRC1hi fibroblasts (Fig. 8). These spatial relationships are consistent with the high TGF-β1 signalling observed as an upstream driver of ABI2 signalling (Fig. 6f).
That hAEC2s undergo progressive transdifferentiation to metaplastic basal cells is not unique to IPF. Alveolar metaplastic basal cells are also common in sections of scleroderma and COVID-19 lungs, and these are intermingled with ABI1 and ABI2 intermediates in areas of active remodelling (Fig. 7d). The common finding of ABIs in hAEC2-derived organoids as well as hAEC2 xenografts, as well as in histologic analyses of fibrotic lungs, suggests that hAEC2s are a major source of metaplastic basal cells in diseases with severe alveolar injury. Future studies are needed to clarify whether and under what circumstances hAEC2 reprogramming towards metaplastic basal cells in the alveoli is reversible, and whether other components of the fibrotic niche such as endothelial cells and immune cells are able to drive the metaplastic phenotype.
Methods
This study complies with all relevant ethical regulations approved by the UCSF Institutional Review Board (IRB) and Institutional Animal Care and Use Committee (IACUC).
Human lung tissue
Studies involving human tissue were approved by the UCSF IRB. All subjects provided written informed consent. Peripheral regions of the normal lungs were obtained to select for the distal-most regions of the lung from brain-dead donors that were rejected for lung transplantation. IPF, scleroderma and acute respiratory distress syndrome (COVID-19) lung specimens were taken from the periphery of the lung at the time of lung transplant. The age and sex of tissue donors are listed in Supplementary Table 6.
Animal studies and treatment
Mice were housed in accordance with the UCSF IACUC protocol in humidity- and temperature-controlled rooms on a 12-h light–dark cycle with free access to food and water. Eight- to twelve-week-old animals were used for the experiments. For labelling mAEC2s, SftpccreERT2/+:R26RmTmG/+ mice33,34 were intraperitoneally administered with tamoxifen (cat no. T5648, Sigma) at 200 mg per kg body weight per day for three consecutive days. NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ (NSG) mice have been described previously35,36. Animal studies utilized a minimum of four mice per group. Mice were injured with oral aspiration of bleomycin (2.1 U per kg body weight). Mice were weighed twice a week. Mice were euthanized between days 17 and 20 post injury for histopathological analysis.
Histology and immunofluorescence
Paraffin embedding
Mouse lungs were inflated with and fixed in 4% paraformaldehyde (PFA) overnight at 4 °C. Human lung pieces were fixed in 4% PFA overnight at 4 °C. The lungs were then washed with PBS four times for 30 min each at 4 °C, then dehydrated in a series of ethanol (30%, 50%, 70%, 95% and 100%). The dehydrated lungs were incubated with xylene for 1 h at room temperature (r.t.), then embedded in paraffin. The lungs were sectioned at 8 µm on a microtome.
Optimal Cutting Temperature (OCT) embedding
Lungs inflated with 94%OCT/2%PFA/4%PBS were fixed with 4% PFA for 1 h at r.t., washed with PBS for 4 h at r.t. and embedded in OCT after 30% and 15% sucrose gradient washing. Organoids in 3D Matrigel were fixed with 4% PFA for 30 min at r.t. or overnight at 4 °C, then washed in PBS overnight three time, followed by embedding in OCT. Sections (8-µm) were cut on a cryostat.
Immunofluorescent staining
Paraffin sections were twice incubated in xylene for 10 min, then rehydrated in ethanol washes (100%, 95%, 70%, 50% ethanol) for 5 min each. OCT-embedded slides were fixed in 4% PFA at r.t. for 10 min, then washed with PBS. For both paraffin- and OCT-embedded slides, antigen retrieval (cat. no. DV2004MX, Biocare) was performed for 30 min at 95 °C or at 155 °C followed by incubation with sodium borohydride (Sigma) in PBS. Slides were washed with 0.1% Tween-20 in PBS (PBST), blocked (3% donkey serum in PBST) for 1 h, and then incubated with primary antibodies overnight at 4 °C. The following primary antibodies were used: anti-SFTPC (cat. no. AB3786, Millipore, 1:2,000), anti-SFTPC (cat. no. sc-518029, Santa Cruz, 1:100), anti-KRT5 (cat. no. 905901, BioLegend, 1:500), anti-KRT14 (cat. no. CBL197, Millipore, 1:200), anti-KRT17 (cat. no. sc-393002, Santa Cruz, 1:100), anti-p63 (cat. no. 13109, CST, 1:100), anti-KRT8 (cat. no. TROMA-I, DSHB, 0.045 µg ml−1), anti-LAMP3 (cat. no. AF0487, R&D, 1:200), anti-ABCA3 (cat. no. ab99856, Abcam, 1:500), anti-human nuclear antigen (cat. no. ab86129, Abcam, 1:500), anti-MMP7 (cat. no. MAB3315, Millipore, 1:200), anti-Ac-Tuba1a (cat. no. T7451, Sigma, 1:500), anti-p-SMAD1/5/8 (cat. no. AB3848-I, Millipore, 1:100), anti-beta IV tubulin (cat. no. ab11315, Abcam, 1:200) and anti-SCGB1A1 (cat. no. MAB4218, R&D, 1:500). Slides were washed with PBST and then incubated with secondary antibodies for 1 h at r.t. The following secondary antibodies were used at 1:250: anti-chicken IgY (H + L) AF647 (cat. no. 703-605-155, Jackson ImmunoResearch), anti-rabbit immunoglobulin-G (IgG; H + L) AF555 (cat. no. A-31572, Thermo Fisher), anti-rabbit IgG (H + L) AF488 (cat. no. A-21206, Thermo Fisher), anti-mouse IgG (H + L) AF555 (cat. no. A-31570, Thermo Fisher) and anti-rat IgG (H + L) AF488 (cat. no. A-21208, Thermo Fisher). 4′,6-Diamidino-2-phenylindole (DAPI) was added for 5 min, then the slides were mounted. Images were captured using a Zeiss Imager M1 and analysed using AxioVision 4.8.2 or Zeiss ZEN v3.1 software(Zeiss). Where indicated, multiple images at ×20 were captured using the ‘MosaiX’ function and stitched together using the ‘Tile Stitch’ function in AxioVision or ZEN. Final images were prepared in Affinity Designer v1.10.1.1142.
Organoid composition calculation
An organoid comprising SFTPC+ cells and no KRT5+ cells was counted as an alveolar organoid (SFTPC+), an organoid comprising both SFTPC+ and KRT5+ cells was counted as a hybrid organoid (SFTPC+/KRT5+) and an organoid comprising KRT5+ cells and no SFTPC+ cells was counted as a basal organoid (KRT5+). Organoids with no detectable staining of either SFTPC or KRT5 were counted as unstained organoids (SFTPC−/KRT5−). To characterize and compare basal cells derived from AEC2/MRC5 and AEC2/AHLM co-cultures, organoids were collected and stained at day 14 with basal cell markers. Individual KRT5+ cells were counted for protein expression of other basal markers and represented as a percentage of total KRT5+ cells. In lineage analysis of day-7 organoids, and testing the effects of BMP4, HHIP, CHIR, KGF, TGF-β1 and DMH-I on the transdifferentiation of hAEC2s into KRT5+ basal cells, the percentages of SFTPC+, KRT5+ and SFTPC+/KRT5+ cells and ABIs were calculated by counting all the cells across multiple organoids on at least one slide.
Lung digestion and fluorescence-activated cell sorting
Human lung pieces were washed in PBS (2×) and HBSS (1×) for 10 min at r.t., compressed to remove liquid, and dissected into 1-cm3 pieces. Dispase II (15 U ml−1; cat. no. 17105041, Thermo Fisher), 225 U ml−1 collagenase type I (cat. no. 17100017, Thermo Fisher), 100 U ml−1 Dnase I (cat. no. DN25, Sigma-Aldrich) and 1% Pen/Strep in 1× HBSS were used to digest the pieces for 2 h at 37 °C. Fungizone (1:400) was added for the final 30 min of the digestion. The digested tissue was liquified in a blender. The suspension was serially filtered through gauze and 100-µm, 70-µm and 40-µm strainers. Red blood cells were removed using red blood cell lysis buffer (Sigma). After Fc blocking, immune and endothelial cells were depleted using biotinylated CD45 (cat. no. 368534, BioLegend, 1:200), CD31 (cat. no. 13-0319-80, eBioscience, 1:200) and CD11b (cat. no. 301304, BioLegend, 1:200) antibodies and running through streptavidin beads (cat. no. 17663, Stemcell Technologies) at 25 µl ml−1. The following antibodies were used at 1:200: anti-CD45-APC-Cy7 (cat. no. 304014, BioLegend), anti-CD11b-APC-Cy7 (cat. no. 557754, BD), anti-CD31-APC-Cy7 (cat. no. 303120, BioLegend), anti-CD326-PE (cat. no. 324206, BioLegend), anti-HTII-280 (cat. no. 303118, Terrace Biotech) and anti-mouse IgM-AF488 (cat. no. A-21042, Thermo Fisher, 1:1,000). Doublets and dead cells were excluded based on forward and side scatters and DRAQ7 (cat. no. 7406S, Cell Signaling, 1:200) or DAPI fluorescence. hAEC2s were sorted as live/EpCAM+/HTII-280+ cells, and AHLM cells were sorted as live/CD45−/CD11b−/CD31−/EpCAM− cells.
Mouse lungs were incubated in the same digestion cocktail for 45 min at 37 °C. The mixture was passed through a 70-µm cell strainer, resuspended in RBC lysis buffer, and filtered through a 40-µm cell strainer. Cells were stained with antibodies for 30 min at 4 °C. Antibodies were used at 1:200: CD45-AF700 (cat. no. 560510, BD), CD31-APC/Fire750 (cat. no. 102528, BioLegend) and CD326-BV421 (cat. no. 563214, BD). mAEC2s were sorted using endogenous green fluorescent protein (GFP) from tamoxifen-induced SftpccreERT2/+:R26RmTmG/+ lungs. AMLM cells were sorted based on the selection of live EpCAM−/CD45−/CD31−.
Freezing/thawing primary human cells
Cells were resuspended in F12 medium at 107 cells per ml, followed by the addition of an equal volume of 2× freeze solution (2% 1.5 M HEPES, 10% FBS, 78% F12 and 10% DMSO). The desired volume of cell aliquots was then frozen at −80 °C and stored in liquid nitrogen. Frozen cells were thawed and an equal volume of warm recovery medium (DMEM with 10% FBS) was added and incubated for 1 min, followed by the addition of 12 ml of warm DMEM to the centrifuge tube. Cells were spun at 550g for 4 min and stained for fluorescence-activated cell sorting (FACS).
Cell culture
Mesenchymal cells were cultured in DMEM/F-12 (cat. no. 11330032, Thermo Fisher) with 10% FBS and 1% Pen/Strep. Cells were used within the first five passages of either being received from ATCC (cat. no. CCL-171) for MRC5 cells or being isolated from donor lungs for AHLM. The construct pLKO5-tRFP (Addgene plasmid 57823) was used to generate RFP-expressing lentiviruses (Lenti-RFP). For labelling AHLM, AHLM cells were transduced by Lenti-RFP with 10 μg ml−1 polybrene (cat. no. TR-1003-G, Millipore). iPSC-derived AEC2s with SFTPC-tdTomato reporter (iAEC2s)23 were generated and maintained in CK + DCI medium as described previously22. The purity of iAEC2 culture was assessed at every passage by flow cytometry, with a cell population with >96% tdTomato+ cells considered pure, as described previously22,23.
Organoid assay
AEC2s and mesenchymal cells were co-cultured (5,000 AEC2s: 30,000 mesenchymal cells per well) in modified MTEC medium diluted 1:1 in growth factor-reduced Matrigel (cat. no. CB-40230A, Thermo Fisher). Modified MTEC culture medium is composed of small airway basal medium (SABM, cat. no. CC-3118, Lonza) with insulin, transferrin, bovine pituitary extract, retinoic acid and Epidermal Growth Factor (EGF) as per the SAGM Bullet Kit and 0.1 µg ml−1 cholera toxin (cat. no. C8052, Sigma), 5% FBS and 1% Pen/Strep. The cell suspension–Matrigel mixture was placed in a transwell and incubated with 10 µM ROCK inhibitor (cat. no. 72252, Stemcell) for the first 48 h. Each experimental condition was performed in triplicate. Where applicable, BMP4 (cat. no. 314-BP-010, R&D Systems, 50 ng ml−1), TGF-β1 (cat. no. 100-21, Peprotech, 3 ng ml−1), DMH-I (cat. no. 73632, Stemcell, 1 µM), HHIP (cat. no. 9280-HP-050, R&D, 2.5 µg ml−1), CHIR (cat. no. 4423, Tocris, 3 µM) and KGF (cat. no. 251KG01050, R&D, 100 ng ml−1) were added to the medium after 48 h and replenished in every medium change. iAEC2s were co-cultured with either MRC5 or AHLM cells (5,000 AEC2s: 30,000 mesenchymal cells per well; both MRC5 and AHLM with passage <5) and maintained with either CK/DCI medium or a 1:1 mixture of CK/DCI and modified MTEC medium as described above. Colonies were assayed after 7, 14 and 21 days. For the mesenchyme-free hAEC2 culture, hAEC2s were isolated from donor lungs via FACS as described above and cultured as previously described21. After two or three passages, the purity of hAEC2s was assessed by protein staining for SFTPC and KRT5. Pure hAEC2s were then co-cultured with either MRC5 or AHLM cells as described above.
To extract RNA from organoids, the cell–Matrigel mixture in the transwell was washed with PBS and incubated in the lung digestion cocktail for 1 h at 37 °C with intermittent resuspension. The mixture was removed from the transwell and resuspended in TrypLE (cat. no. 12563011, Thermo Fisher) and shaken at 37 °C for 20 min. Cells were blocked with human FcR blocking reagent (cat. no. 564220, BD, 1:50) for 10 min at 4 °C, then stained with biotin anti-CD326 (cat. no. 324216, BioLegend, 1:250) for 30 min at 4 °C. Streptavidin beads (cat. no. 17663, STEMCEL, 1:50) were added to isolate the epithelial cells, and the rest of the cells were mesenchymal cells. For sequencing, organoids were digested as above and FACS-sorted for live EpCAM+ and EpCAM− cells.
Quantitative polymerase chain reaction
RNA was extracted from cells using a PicoPure RNA isolation kit (cat. no. KIT0204, Applied Biosystems). Complementary DNA (cDNA) was synthesized from total RNA using a SuperScript Strand Synthesis System (cat. no. 18080044, Thermo Fisher). Quantitative PCR (qPCR) was performed using SYBR Green (cat. no. F415L, Thermo Fisher). Relative gene expression levels were defined using the ∆∆Ct method. The qPCR primers (IDT) are listed in Supplementary Table 7.
RNA in situ hybridization
PFA-fixed OCT-embedded sections were used for RNA in situ hybridization with an RNAScope multiplex fluorescent v2 assay (ACDBio). Briefly, 7-µm sections of normal or IPF lungs were washed, protease-dependent antigen retrieval was performed, and probes were hybridized for 2 h at 40 °C, followed by step-wise amplification of each probe. RNA probes for SFTPC (cat. no. 452561-C1 and -C2, ACDBio), KRT17 (cat. no. 463661-C3, ACDBio) and CTHRC1 (cat. no. 413331, ACDBio) were used. Following completion of RNA in situ hybridization, immunostaining was performed after blocking, as already described.
Xenotransplantation assay
Freshly sorted hAEC2s, 700,000 alone or mixed with 200,000 cultured AHLM or MRC5 cells (passage <4) in a 40-µl volume (1× PBS), were transplanted at 10 days post bleomycin injury into the lungs of NSG mice via oral aspiration. Transplanted mice were euthanized 8–12 days post-transplant (a total of 18–22 days post bleomycin injury).
Single-cell transcriptomics
Single-cell sequencing was performed on a 10X Chromium instrument (10X Genomics) at the Institute of Human Genetics in UCSF as described before37. Briefly, cells were isolated via FACS and then loaded on a Chromium Controller instrument to generate single-cell Gel Bead-In-EMulsions (GEMs). Libraries were prepared by performing reverse transcription on a Bio-Rad C1000 Touch Thermal Cycler (Bio-Rad), then GEMs were collected to amplify cDNA using a Bio-Rad C1000 Touch Thermal Cycler. SPRIselect (Beckman Coulter) was used to select for amplified cDNA. Indexed sequencing libraries were constructed using the Chromium Single-Cell 3′ library kit (10X Genomics) and sequenced on a NovaSeq 6000 instrument (Illumina) with the following parameters: Read 1 (26 cycles), Read 2 (98 cycles) and i7 index (eight cycles) to obtain a sequencing depth of ~100,000 reads per cell. Reads were aligned to the appropriate mouse or human genome and quantified using the Cell Range Single-Cell Software Suite.
Quantification and statistical analysis
Statistical analysis
All statistical analyses were performed in GraphPad Prism. Unpaired one-tailed t-tests were used to determine the P values, and the data in the graphs are presented as mean ± s.d. For quantification of xenotransplants, the Kolmogorov–Smirnov test was used to determine normality and one-way ANOVA followed by the Kruskal–Wallis test was used for multiple comparisons. For normally distributed data, ordinary one-way ANOVA followed by Tukey’s multiple comparisons test was performed. For all tests, P values < 0.05 were considered significant. The representation factor was calculated to represent the number of overlapping genes divided by the number of expected overlapping genes drawn from two independent groups, as calculated on nemates.org with a base value of 30,000 genes in the human genome. A representation factor of >1 indicates more overlapping genes than expected of two independent groups.
Immunofluorescence image quantification
Sections were imaged for quantification on a Zeiss Lumar V12 or Zeiss AxioImager.M1 microscope. Cell counts for stained cells were performed on Fiji using the ‘Cell Counter’ plug-in. The results were averaged between each specimen and s.d. values were calculated per condition. For quantification of xenografts, at least three mice in each group were analysed with at least two sections taken at 200 µm apart to capture different regions of the lung. Any HNA+ region of more than five cells was counted as one engrafted region. At least three lobes were analysed at each section for three mice per condition, and a minimum of two sections per mouse were analysed. Data are presented as mean ± s.d., **P < 0.01 (one-way ANOVA followed by Kruskal–Wallis for multiple comparison). For quantification of ABIs, at least six distinct areas per IPF specimen (n = 3), covering at least three different zones of disease severity, were counted. Zones of disease severity were defined as follows: (1) normal-looking alveoli, alveoli with a single layer of DAPI+ cells; (2) thickened alveolar walls, alveoli with multiple layers of DAPI+ cells in the alveolar wall; (3) microcysts, cyst-like structures that line and overlay the fibrotic regions as judged by accumulated DAPI+ cells in alveolar space; (4) heavily fibrotic/bronchiolized zone, fibrotic lesions with autofluorescent collagen deposition around bronchiolized epithelium. SFPTC+/KRT17+/KRT5− cells were detected by mRNA in situ for SFTPC and KRT17 and protein staining for KRT5 and were counted as ABI1s, SFTPC−/KRT17+/KRT5− cells were counted as ABI2s and SFTPC−/KRT17+/KRT5+ cells were counted as mature basal cells. IPF lung specimens (n = 3) were probed with probes against SFTPC, KRT17 and CTHRC1 mRNA and KRT5 protein staining to identify ABIs and adjacent CTHRC1+ mesenchyme. At least a 3 × 3 mosaic image was captured at ×20 (nine images covering an area of 3.28 mm2), stitched together and counted as one data point. In total, 160 images were captured at × 20 across three IPF lungs and every instance of ABI1, ABI2 or basal cell in the actively remodelled region was counted. A cell cluster of at least five cells was counted as an ABI or a CTHRC1+ mesenchymal region. A mesenchymal region was marked as adjacent to ABIs if it was within a three-cell distance of an ABI cluster.
Analysis of single-cell RNA-sequencing
FASTQ files were run through CellRanger v2.1.1 software with default settings for de-multiplexing, aligning reads with STAR software to Hg19 or GRCh38, and counting unique molecular identifiers (UMIs). Seurat package v4.038 in R v3.5.1 was used for downstream analysis. Low-quality cells were filtered (expressing fewer than 200 genes, >10% mitochondrial reads and >6,000 unique gene counts). Principal component analysis was performed on log-normalized and scaled data using 2,000 variable genes. The top 10 principal component analyses were used for clustering and visualized using the UMAP algorithm in the Seurat package. Monocle trajectory analysis was performed using Monocle 339 by importing the counts from the Seurat object. RNA velocity was calculated using the scVelo v0.1.16 package40 in Python v3.6 and velocity calculations were overlaid on UMAP projections calculated in Seurat. Single-cell transcriptomes of healthy and IPF lungs were obtained from GSE135893 and GSE132771 and processed using Seurat. AEC2s, AEC1s, transitional AEC2s, KRT17+/KRT5− cells and basal cells were extracted and analysed. RNA values were averaged for each cluster using the AverageExpression function followed by the DoHeatmap function in Seurat to generate heatmaps. Genes specific to the PATS-like cells reported in Kobayashi et al.20 were picked for heatmaps. The lists of DEGs were identified with a Model-based Analysis of Single-cell Transcriptomics (MAST) test. Upstream regulator analysis of gene lists containing significantly differentially expressed genes (Padj < 0.05, logFC < −0.15 or >0.15) was done with Ingenuity Pathway Analysis (Qiagen).
Bulk RNA-sequencing analysis
Total RNA was extracted from AHLM and MRC5 cells using the RNeasy Kit (cat. no. 74004, Qiagen). Sequencing was performed using HiSeq 4000. Quality control of reads was conducted using FastQC (Babraham Bioinformatics). Ligation adaptors were removed using Cutadapt and Sickle 1.33. Sequencing reads were aligned using STAR 2.7.9a, and UCSC human GRCh38/hg38 was used as the reference genome. The differential gene expression list was generated using DEseq2.
Statistics and reproducibility
No statistical method was used to predetermine sample size. No data were excluded from analyses. Sample ID was blinded to the investigator for image quantification.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this Article.
Data availability
The RNA-seq data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under the following accession codes: GSE148992 (human bulk RNA-seq), GSE150068 (organoids and IPF) and GSE150247 (human scRNA-seq). Previously published scRNA-seq data that are re-analysed here are available at GSE135893 and GSE132771. All other data supporting the findings of this study are available from the corresponding author on reasonable request. Source data are provided with this paper.
Code availability
No custom codes were developed and used in this manuscript. All codes are available by request to the corresponding author.
References
Barkauskas, C. E. et al. Type 2 alveolar cells are stem cells in adult lung. J. Clin. Invest. 123, 3025–3036 (2013).
Rock, J. R. et al. Multiple stromal populations contribute to pulmonary fibrosis without evidence for epithelial to mesenchymal transition. Proc. Natl Acad. Sci. USA 108, E1475–E1483 (2011).
Smirnova, N. F. et al. Detection and quantification of epithelial progenitor cell populations in human healthy and IPF lungs. Respir. Res. 17, 83 (2016).
Xu, Y. et al. Single-cell RNA sequencing identifies diverse roles of epithelial cells in idiopathic pulmonary fibrosis. JCI Insight 1, e90558 (2016).
Xi, Y. et al. Local lung hypoxia determines epithelial fate decisions during alveolar regeneration. Nat. Cell Biol. 19, 904–914 (2017).
Vaughan, A. E. et al. Lineage-negative progenitors mobilize to regenerate lung epithelium after major injury. Nature 517, 621–625 (2015).
Liang, J. et al. Hyaluronan and TLR4 promote surfactant-protein-C-positive alveolar progenitor cell renewal and prevent severe pulmonary fibrosis in mice. Nat. Med. 22, 1285–1293 (2016).
Habermann, A. C. et al. Single-cell RNA sequencing reveals profibrotic roles of distinct epithelial and mesenchymal lineages in pulmonary fibrosis. Sci. Adv. 6, eaba1972 (2020).
Adams, T. S. et al. Single-cell RNA-seq reveals ectopic and aberrant lung-resident cell populations in idiopathic pulmonary fibrosis. Sci. Adv. 6, eaba1983 (2020).
Kumar, P. A. et al. Distal airway stem cells yield alveoli in vitro and during lung regeneration following H1N1 influenza infection. Cell 147, 525–538 (2011).
Ray, S. et al. Rare SOX2+ airway progenitor cells generate KRT5+ cells that repopulate damaged alveolar parenchyma following influenza virus infection. Stem Cell Rep. 7, 817–825 (2016).
Yang, Y. et al. Spatial-temporal lineage restrictions of embryonic p63+ progenitors establish distinct stem cell pools in adult airways. Dev. Cell 44, 752–761 (2018).
Pan, H., Deutsch, G. H., Wert, S. E. & Ontology, S. NHLBI Molecular Atlas of Lung Development Program Consortium Comprehensive anatomic ontologies for lung development: a comparison of alveolar formation and maturation within mouse and human lung. J. Biomed. Semant. 10, 18 (2019).
Prasse, A. et al. BAL cell gene expression is indicative of outcome and airway basal cell involvement in idiopathic pulmonary fibrosis. Am. J. Respir. Crit. Care Med. 199, 622–630 (2019).
Zacharias, W. J. et al. Regeneration of the lung alveolus by an evolutionarily conserved epithelial progenitor. Nature 555, 251–255 (2018).
Wang, C. et al. Expansion of hedgehog disrupts mesenchymal identity and induces emphysema phenotype. J. Clin. Invest. 128, 4343–4358 (2018).
Jiang, P. et al. Ineffectual AEC2-to-AEC1 differentiation in IPF: persistence of KRT8hi transitional state. Am. J. Respir. Crit. Care Med. 201, 1443–1447 (2020).
Choi, J. et al. Inflammatory signals induce AT2 cell-derived damage-associated transient progenitors that mediate alveolar regeneration. Cell Stem Cell 27, 366–382 (2020).
Strunz, M. et al. Alveolar regeneration through a KRT8+ transitional stem cell state that persists in human lung fibrosis. Nat. Commun. 11, 3559 (2020).
Kobayashi, Y. et al. Persistence of a regeneration-associated, transitional alveolar epithelial cell state in pulmonary fibrosis. Nat. Cell Biol. 22, 934–946 (2020).
Katsura, H. et al. Human lung stem cell-based alveolospheres provide insights into SARS-CoV-2-mediated interferon responses and pneumocyte dysfunction. Cell Stem Cell 27, 890–904 (2020).
Jacob, A. et al. Differentiation of human pluripotent stem cells into functional lung alveolar epithelial cells. Cell Stem Cell 21, 472–488 (2017).
Hurley, K. et al. Reconstructed single-cell fate trajectories define lineage plasticity windows during differentiation of human PSC-derived distal lung progenitors. Cell Stem Cell 26, 593–608 (2020).
Weiner, A. I. et al. Mesenchyme-free expansion and transplantation of adult alveolar progenitor cells: steps toward cell-based regenerative therapies. NPJ Regen. Med 4, 17 (2019).
Tsukui, T. et al. Collagen-producing lung cell atlas identifies multiple subsets with distinct localization and relevance to fibrosis. Nat. Commun. 11, 1920 (2020).
Cassandras, M. et al. Gli1+ mesenchymal stromal cells form a pathological niche to promote airway progenitor metaplasia in the fibrotic lung. Nat. Cell Biol. 22, 1295–1306 (2020).
Travaglini, K. J. et al. A molecular cell atlas of the human lung from single-cell RNA sequencing. Nature 587, 619–625 (2020).
Carraro, G. et al. Single-cell reconstruction of human basal cell diversity in normal and idiopathic pulmonary fibrosis lungs. Am. J. Respir. Crit. Care Med 202, 1540–1550 (2020).
Rosas, I. O. et al. MMP1 and MMP7 as potential peripheral blood biomarkers in idiopathic pulmonary fibrosis. PLoS Med. 5, e93 (2008).
Ning, J., Zhao, Y., Ye, Y. & Yu, J. Opposing roles and potential antagonistic mechanism between TGF-beta and BMP pathways: implications for cancer progression. EBioMedicine 41, 702–710 (2019).
Dituri, F., Cossu, C., Mancarella, S. & Giannelli, G. The interactivity between TGFβ and BMP signaling in organogenesis, fibrosis and cancer. Cells 8, 1130 (2019).
Basil, M. C. et al. The cellular and physiological basis for lung repair and regeneration: past, present and future. Cell Stem Cell 26, 482–502 (2020).
Chapman, H. A. et al. Integrin α6β4 identifies an adult distal lung epithelial population with regenerative potential in mice. J. Clin. Invest. 121, 2855–2862 (2011).
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007).
Coughlan, A. M. et al. Myeloid engraftment in humanized mice: impact of granulocyte-colony stimulating factor treatment and transgenic mouse strain. Stem. Cells Dev. 25, 530–541 (2016).
Shultz, L. D. et al. Human lymphoid and myeloid cell development in NOD/LtSz-scid IL2Rγnull mice engrafted with mobilized human hemopoietic stem cells. J. Immunol. 174, 6477–6489 (2005).
Kathiriya, J. J., Brumwell, A. N., Jackson, J. R., Tang, X. & Chapman, H. A. Distinct airway epithelial stem cells hide among club cells but mobilize to promote alveolar regeneration. Cell Stem Cell 26, 346–358 (2020).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Acknowledgements
We thank A. Baldwin for providing technical assistance, Parnassus Flow Cytometry Core for assistance with cell sorting for bulk and single-cell RNA analysis (P30DK063720), E. Wan and the Institute for Human Genetics Core for processing of single-cell RNA samples and high-throughput sequencing. The GEO accession number for raw RNA-sequencing data is given in the Methods. This work is supported by NIH grants nos. DP2AG056034, R01HL142552 and R01HL155622 to T.P., R01HL128484, R35HL150767 and U01HL134766 to H.A.C. and F32HL143931-01A1 and K99HL155785-01 to J.J.K., a tobacco-related disease research programme postdoctoral award to C.W. and a Nina Ireland Program Award to M.M. for human lung collection.
Author information
Authors and Affiliations
Contributions
J.J.K., C.W., H.A.C. and T.P. conceived the experiments and wrote the manuscript. C.W., J.J.K., M.Z., A.B., C.L.S., M.C. assandras, M. Cohen, B.W. and K.-D.A. performed the experiments, collected samples and analysed data. D.N.K., P.W., B.W. and M.M. provided materials and input to the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Cell Biology thanks Bi-Sen Ding and the other, anonymous, reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Purified hAEC2s transdifferentiate into KRT5+ basal cells when co-cultured with AHLM.
(a) FACS plot shows the strategy for isolating hAEC2s. After selecting live/CD45-/CD31-/CD11b- cells, hAEC2s were further isolated by gating EpCAM + /HTII-280+ cells. (b) Quantification of SFTPC + and KRT5 + cells in freshly sorted HTII-280+ cells by cytospins. Data are mean ± SD, representative of n = 4 biologically independent human lung cytospins analyzed. (c) confirmation of the purity of HTII-280+ cells by scRNA-seq showed no appreciable detection of non-AEC2 markers. (d) scRNA-seq of AHLM isolated as live/CD31-/CD45-/CD11b-/EpCAM- cells that express PDGFRA. (e) Colony forming efficiency of hAEC2/MRC5 and hAEC2/AHLM co-culture at D14. Data are mean ± SD, representative of n = 3 biologically independent for AHLM and hAEC2 and representative of n = 3 independent experiments for MRC5. (f) and (g) IF analysis of p63 and KRT17, and KRT5 in the hAEC2s + MRC5 and hAEC2s+AHLM organoids at D14. Images are representative of n = 2 (in f) and n = 3 (in g) biological replicates. (h) Quantification of KRT17 + and p63+ cells in total KRT5 + basal cells in the hAEC2s + MRC5 and hAEC2s+AHLM organoids at D14. n = 2 (for MRC5) and n = 3 (for AHLM) biologically independent samples were analyzed. Data are mean ± SD. (i) IF analysis of KRT8 in the hAEC2s+AHLM organoids at D14. Representative of n = 2 independent experiments. (j) IF analysis and quantification of KRT5 + and SFTPC + cells in the hAEC2-derived organoids co-cultured with freshly sorted AHLM (Passage 0) and cultured AHLM (Passage 2) at D14. Each dot represents a technical replicate. Representative of n = 2 independent experiments. (k) Enrichment of hAEC2s using a mesenchyme-free system, confirmed by IF analysis and quantification of SFTPC + and KRT5 + cells (left), and co-culture of the enriched hAEC2s with AHLM or MRC5, the organoids are analyzed at D14 (right). Data are expressed as the mean ± SD, representative of n = 3 biologically independent experiments.
Extended Data Fig. 2 Difference in niche provided by mouse and human lung mesenchyme and components of the CK/DCI media on hAEC2s-to-basal transdifferentiation.
(a) IF analysis of mouse AEC2s (mAEC2s)-derived organoids co-cultured with the adult mouse lung mesenchyme (AMLM) at D14. mAEC2s (GFP + ) were sorted from the lung of adult SftpcCreERT2/+:R26RmTmG/+ mouse. Representative of n = 3 independent experiments. (b) Quantification of lineages of organoids (SFTPC + organoids, organoids with both SFTPC + cells and KRT5 + cells, or KRT5 + organoids) in mAEC2 + AMLM organoids. (c) hAEC2s co-cultured with either adult murine lung mesenchyme (AMLM) or AHLM. (d,e) IF analysis and quantification of hAEC2 + AHLM organoids treated with 3 µM CHIR or 100 ng/ml KGF at D14. Representative of n = 2 independent experiments. Data are expressed as the mean ± SD, representative of n = 3 technical triplicates. ** p = 0.0012. (f) IF analysis of human iAEC2 + AHLM organoids cultured in 1:1 dilution of CK-DCI:SABM/MTEC standard media show canonical basal markers (KRT14, KRT17, P63) and lung lineage (NKX2-1). Data are expressed as the mean ± SD, representative of n = 2 independent experiments. ** indicates p < 0.01 as determined by unpaired one-tailed student’s t-test.
Extended Data Fig. 3 Comparison of pathologic mesenchymal niche subsets arising from IPF and cultured AHLM.
(a, b) IF analysis of hAEC2s-derived organoids co-cultured with human adult normal lung (AHLM) and IPF mesenchyme at D7, and quantified by the percentage of SFTPC + , KRT5 + , and SFTPC + /KRT5 + cells in total cells. Data are expressed as the mean ± SD, representative of n = 3 technical replicates. * p = 0.0182 and ** p = 0.0066. Representative of n = 2 independent experiments. (c) UMAP plots of mesenchyme from each individual IPF or normal donor that was used in scRNA-seq analysis. (d) Comparison of top 100 markers in CTHRC1 + (left) and HAShigh/BMP anatgonisthigh (right) clusters of uncultured IPF (IPF fresh) and cultured AHLM with statistical comparison for degree of gene overlap of two independent gene sets denoted by representation factor (see methods). (e, f, and g) UAMP plots show BMP antagonists and TGFB1 expression in IPF lungs (from IPF Cell Atlas, Habermann et al.). Data are expressed as the mean ± SD. * indicates p < 0.05 and ** indicates p < 0.01 as determined by unpaired one-tailed student’s t-test. Each data point on the graph represents a technical replicate.
Extended Data Fig. 4 Effect of HHIP and BMP4 in modulating metaplastic hAEC2 transdifferentiation.
(a) IF analysis of SCGB1A1 in the hAEC2+ AHLM organoids treated with 2.5 μg/ml HHIP and 50 ng/ml BMP4, quantified by percentage of organoid containing SCGB1A1 + cells. Data are expressed as the mean ± SD. * p = 0.0441. (b) IF analysis of pSMAD1/5/8 in the hAEC2+ AHLM organoids treated with 2.5 μg/ml HHIP, quantified by percentage of pSMAD1/5/8+ cells in total cells. Data are expressed as the mean ± SD. ** p = 0.0079. Experiment was performed in a technical triplicate. * indicates p < 0.05 and ** indicates p < 0.01 as determined by unpaired two-tailed student’s t-test.
Extended Data Fig. 5 Comparison of hAEC2-derived and IPF basal cells.
(a) Feature plots show KRT5 and KRT17 expression in the merged basal cell data from fresh normal donors (N = 2, marked as NL basal), fresh IPF patients (N = 2, marked as IPF basal), and hAEC2+AHLM organoids (N = 2). (b) Comparison of top 100 markers in SPB, MPB, AB, and PB clusters of this study and Carraro et al.3 with statistical comparison for degree of gene overlap of two independent gene sets denoted by representation factor was done via http://nemates.org (see methods) (see methods). (c) UMAP plots of basal cells from IPF, normal donors, and hAEC2-derived organoids with contributing cell number in each cluster. (d) Comparison of top 500 differentially expressed genes in hAEC2-derived and IPF basal cells against normal lung basal cells with statistical comparison for degree of gene overlap of two independent gene sets denoted by representation factor was done via http://nemates.org (see methods). (e) Representative upregulated genes in top 50 of IPF and hAEC2-derived vs. normal lung basal cells. SPB = secretory primed basal, MPB = multipotent basal, AB = activated basal, PB = proliferating basal.
Extended Data Fig. 6 Expression of ABI markers in organoid and comparison with PATS-like cells in IPF.
(a) Heatmap of lineage marker expressions in different cellular subsets arising from hAEC2-derived organoids. (b) Expression along pseudotime trajectory of AEC2, ABI, and basal cell markers. (c) Featureplots of organoid scRNA-seq (D0-D21) show SCGB1A1 + club cells and FOXJ1 + ciliated cells. (d) IF staining of hAEC2+AHLM organoids (Day 21) show SCGB1A1 + club and rare TUBB4 + ciliated cells. Representative of n = 2 independent experiments. (e) Top 15 significantly differentially expressed genes between AEC2s, ABI1s, ABI2s, and basal cells show AEC2-like profile of ABI1s and basal-like profile of ABI2s in addition to high ECM related gene signature of ABI2s. (f) UMAP of hAEC2-derived organoids co-cultured with AHLM and IPF epithelium where transitional AEC2s and KRT17 + /KRT5- cells as originally described by Habermann et al4. appear. The combination of transitional AEC2s and KRT17 + /KRT5- cells were labelled as PATS-like cells by Kobayashi et al1. Feature plots comparing (g) hAEC2 lineage markers expression, (h) PATS-like markers expression, and (i) basal lineage marker expression in the UMAPs of organoid and IPF.
Extended Data Fig. 7 Identification of ABIs in normal, IPF, and COVID-19 lungs.
(a) Immunostaining of IPF lung with AEC2, ABIs, and basal markers show (1) linearly connected AEC2, ABI1, ABI2, and basal cell, (2) a cluster of basal cells (KRT17 + /KRT5 + /TP63 + ), and two types of ABI2s (KRT17 + /KRT5-): (3) TP63neg and (4) TP63pos. Representative of n = 3 independent IPF lungs. (b) In situ analysis hybridization of normal lung with SFTPC and KRT17 identifies rare ABI1s (SFTPC + /KRT17 + ) as a fraction of total SFTPC + cells. Data are represented as mean ± SD, representative of n = 2 biologically independent samples. (c) Immunostaining of COVID-19 lung shows presence of AEC2s, ABI1 and ABI2s, and basal cells in the same bronchiolized region. Representative of n = 1 sample.
Supplementary information
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
About this article
Cite this article
Kathiriya, J.J., Wang, C., Zhou, M. et al. Human alveolar type 2 epithelium transdifferentiates into metaplastic KRT5+ basal cells. Nat Cell Biol 24, 10–23 (2022). https://doi.org/10.1038/s41556-021-00809-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-021-00809-4
This article is cited by
-
The use of cultured human alveolar basal cells to mimic honeycomb formation in idiopathic pulmonary fibrosis
Respiratory Research (2024)
-
A WNT mimetic with broad spectrum FZD-specificity decreases fibrosis and improves function in a pulmonary damage model
Respiratory Research (2024)
-
Human alveolar hydrogels promote morphological and transcriptional differentiation in iPSC-derived alveolar type 2 epithelial cells
Scientific Reports (2023)
-
Alveolar epithelial progenitor cells require Nkx2-1 to maintain progenitor-specific epigenomic state during lung homeostasis and regeneration
Nature Communications (2023)
-
Autocrine TGF-β-positive feedback in profibrotic AT2-lineage cells plays a crucial role in non-inflammatory lung fibrogenesis
Nature Communications (2023)