Abstract
Massive single-cell profiling efforts have accelerated our discovery of the cellular composition of the human body while at the same time raising the need to formalize this new knowledge. Here, we discuss current efforts to harmonize and integrate different sources of annotations of cell types and states into a reference cell ontology. We illustrate with examples how a unified ontology can consolidate and advance our understanding of cell types across scientific communities and biological domains.
Similar content being viewed by others
Main
With collaborations of over 2,000 scientists across more than 1,000 institutes from 76 countries to date, the Human Cell Atlas (HCA) has generated comprehensive molecular profiles of tens of millions of single cells across 18 different organs and systems, which in turn are advancing our understanding of the definition of cell types and states1,2. Technological advances in single-cell and spatial genomics are rapidly expanding the compendium of known cell types3 and accelerating discoveries of a large variety of novel cell populations.
For instance, these efforts have been applied to system-level disciplines such as immunology and neuroscience, both of which require an understanding of vast networks of cells and tissues. In immunology, cell types have been historically recognized and well characterized. Yet the number of discrete cell types and specific cell states identified from single-cell genomics has exceeded expectations, in particular with respect to the diversity of cell states derived from developmental dynamics4, tissue-resident phenotypes5 and activation states6. For example, transcriptomics profiling identified three decidual natural killer cell populations at the maternal–fetal interface that show varying levels of immunoregulatory properties and modulate trophoblast invasion7. Transcriptomics and genomics profiling studies have also captured an increasing variety of cell types and gene programmes in the central and peripheral nervous systems. Cell atlasing (that is, the creation of a cell atlas) of mammalian brains has led to the discovery of previously uncharacterized cell types, including more than 100 cell types in a single region of the neocortex8, as well as of cellular diversity due to species-specific adaptations in the cortex8. A similar dramatic increase in diversity has been reported in the peripheral nervous system such as in the enteric nervous system9,10.
This incredible progress takes us closer to answering a general question that motivates stem and developmental cell biologists, as well as the HCA project: what is the complete cellular makeup of the human body? Annotating cells and gene programmes is crucial not only to address this question but also to fully exploit these data for biological discovery, including in pathological states. This can only be achieved by naming the entities we study in a consolidated manner so that findings can be related between studies and one study can build on findings from multiple previous ones as knowledge is accrued and expanded. However, most annotations of single-cell genomics datasets to date have used uncontrolled free text (that is, arbitrary naming schemes) for cell type names, which makes the cross-searching of annotations across separate datasets challenging and unreliable. In some cases, with a naming scheme absent, cells are described merely by a subset of their molecular characteristics and therefore hard to match between studies.
To fully answer the question of what constitutes the cellular composition of the human body, there is an urgent need to put new discoveries from the HCA into the context of classical cell biology and anatomy, as well as developmental biology, neurobiology and pathology. Cell ontologies, a structured controlled vocabulary for cell types in animals, are a tremendously powerful way of formalizing such knowledge, which in turn opens up opportunities for quantitative scientific interrogation of the HCA data in new and exciting ways.
In this Perspective, we discuss the utility and parts of cell ontologies, review the state of current cell ontologies and conclude with ongoing efforts and how they can be applied to discovery over the coming years.
Using cell ontology for knowledge integration and mining
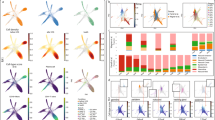
Biomedical ontologies originated in simple controlled vocabularies developed to supplement or replace the free-text metadata in databases, clinical records and medical billing systems11. Standardizing the text used to record, for example, diseases, gene functions, anatomical structures and cell types within and between databases makes it possible to reliably search and group records referring to the same entities (for example, by diseases or cell types). However, controlled vocabularies are not sufficient for searching and grouping records with closely related contents. For example, a user searching a database for records relating to macrophages or liver sinusoid would not find records for Kupffer cells unless the data structures driving the search had some meaningful ways to relate the terms ‘macrophage’, ‘Kupffer cell’ and ‘liver sinusoid’. Cell ontologies provide mechanisms for this integration, which allows us to record a Kupffer cell as a type of macrophage located in the liver sinusoid and then to enrich search results to take advantage of the classification and location relationships (Fig. 1).
A graph showing the relationships between terms for anatomical structures (for example, hepatic sinusoid), cell types (for example, macrophage) and functional roles (for example, erythrocyte clearance). The following relationships are shown: ‘is a’, which records the classification; ‘part of’, which relates cells to their tissues and organs; ‘located in’, which relates cells to spaces such as the hepatic sinusoid; ‘develops in’, which records the developmental origin; and ‘capable of’, which records the function.
Ontologies of cell types such as those in Cell Ontology (CL)12 and Drosophila Anatomy Ontology13 are increasingly being used to annotate single-cell transcriptomics data. The use of ontology terms in dataset annotation relates annotated data back to hard-earned legacy knowledge, classical terminologies and the accompanying understanding of cell types, anatomies and development. Such annotation makes data cross-searchable, discoverable, integrable and more accessible to general cell biologists. It facilitates cross-dataset analyses, which then allows more quantitative analyses of similarities across thousands of individual cells and leads to more nuanced views of cell types, their classification and their properties.
CL was first developed as a platform in 2004 to collect major cell types from humans and model organisms, and has since been applied to various fields. For example, the Encyclopedia of DNA Elements (ENCODE) Consortium used CL to annotate its compendium of cell types, which yielded a prioritized set of genetic and epigenetic elements14. Because the precise terms used for cell types, anatomical structures and diseases often greatly vary across sources, biomedical ontologies, including CL, typically use a bipartite system of universally resolvable identities (IDs) in the form of URLs for ontology terms, with each linked to an official label. For example, the term with the primary label ‘Kupffer cell’ in CL is identified by the permanent URL http://purl.obolibrary.org/obo/CL_0000091, which is further abbreviated to the compact form CL:0000091 (ref. 15). Critically, using resolvable IDs rather than labels to refer to cell types in database records allows associated metadata (for example, labels, descriptions and references) and their relationships (for example, anatomy, development, functional and pathological relevance) to evolve over time with no cost for the databases and records that use IDs to refer to them (Fig. 1).
Ontologies can serve to link and integrate heterogeneous data types related to the same cell type across multiple modalities. For example, Virtual Fly Brain16,17 and the Fly Cell Atlas18 use the same ontology terms to annotate three-dimensional images of neurons (>70,000 images), connectomics data (>3.5 million pairwise connections) and single-cell transcriptomics data (~600,000 cells). Similarly, CL terms, classifications and relationships are also increasingly being used to define and classify terms in the Gene Ontology database19 (>750 terms) and in widely used ontologies of phenotypes (730 terms in Human Phenotype Ontology20) and diseases (>3,000 terms in Mondo Disease Ontology21). These links make it possible to combine single-cell, phenotype and disease data relating to the same cell types. With the advent of large-scale single-cell transcriptomics atlasing, community-driven nomenclature- and ontology-building projects have emerged and are coordinating with existing ontology-building efforts (for example, HCA Biological Networks2, the Human BioMolecular Atlas Program (HuBMAP)22, BRAIN Initiative Cell Census Network (BICCN)23 and Cell Annotation Platform (http://celltype.info)).
This is already affecting our ability to organize our knowledge of cell types for comparisons of datasets across individual laboratories and, notably, for effectively interpreting health and disease using the knowledge from both classical histopathology and single-cell genomics. For instance, ontological distinctions between fetal and mature cells in the kidney are mirrored by differences in their molecular signatures, which are critical to understanding the divergent origins of paediatric and adult kidney cancers, respectively24. Similarly, datasets that were annotated in a consistent manner facilitated cross-tissue meta-analyses for COVID-19 that identified specialized nasal epithelial cells that were enriched in the expression of entry factors for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)25. Such datasets also enabled the identification of covariates such as age, sex and smoking status associated with SARS-CoV-2 entry factor expression in lung and airway cells26, and a comparison between cells from autopsy tissue samples from patients with COVID-19 and from healthy and other disease conditions27. Together, these studies highlight the necessity and utility of establishing previously agreed ontological classifications.
Considerations in the classification of human cell types
Biologists have long recognized that the natural world lends itself to hierarchical systems of classification, which capture the underlying hierarchical processes that drive biology, such as the phylogenetic classification of species by morphological and molecular observations. Similarly, cell types can be hierarchically classified and categorized in ever-increasing levels of resolution, from a general cell type such as an endothelial cell to more specialized types such as a liver sinusoidal endothelial cell (LSEC) and then down to highly specialized types found in specific locations such as a periportal LSEC. As with taxonomy for a species, various kinds of observations inform the ultimate classification, and these different types of information are often used in concert to arrive at a particular cell-type definition.
Taking anatomical locations as an example, CL12 imports information about anatomical structures and features from the Uber-anatomy Ontology (Uberon)28 and relates them to CL terms using, for example, ‘part of’ to relate cell types to the tissues and organs, and ‘located in’ to relate cell types to cavities within structures. For example, the definition of an LSEC in CL includes a ‘part of’ relationship to ‘hepatic sinusoid’, which indicates that the LSEC forms part of the structure of the hepatic sinusoid as defined in Uberon. By contrast, the definition of Kupffer cells records that they are ‘located in’ (the lumen of) the hepatic sinusoid. In an anatomically higher hierarchy, the definition of hepatic sinusoid involves relations to the liver lobule and the liver overall, which is in turn defined by its structure, location and physiological role in the body. The LSEC is therefore hierarchically defined relative to the whole organism down to its individual position in the specific tissue where it is found (Fig. 2a). Furthermore, since CL classifies cell types hierarchically from generic cell types down to more specialized types, an LSEC is also defined as a descendent of the general endothelial cell class in CL. The main LSEC class (officially ‘endothelial cell of hepatic sinusoid’) has its own descendent classes, which represents the following further specializations of LSECs: ‘endothelial cell of periportal hepatic sinusoid’ and ‘endothelial cell of pericentral hepatic sinusoid’.
a, CL has terms for a variety of cell types associated with the hepatic sinusoid (Uberon:0001281). The classification of these cell types allows them to be grouped with other cells from the same location. For example, Kupffer cells (CL:0000091) can be grouped with other tissue-resident macrophages or with cells of the hepatic sinusoid. b, Ontologies can be used to encode transitions through diverse cell states. Examples include T-cell activation following antigen recognition, cell cycling, neuron development and maturation, smooth muscle cell (SMC) contraction and relaxation, and cell destruction after oxidative stress.
Sources of information that contribute to a cell-type categorization include morphological features, developmental origins and functional profiles. Ontologies attempt to capture all terms that are used by different scientific communities to refer to the same cell type, as well as alternative names that may not be commonly used. Historically, different fields in biology have focused on different aspects of cells to drive their naming. For example, many immune cells have been classified according to which cell surface protein(s) they express29,30,31,32,33,34,35,36, whereas cells of the nervous system have been named according to a combination of features, including morphologies, physiologies, connectivities and the roles they play in the neuronal circuitry37. In some systems, such as the retina38, there is strong evidence to indicate that cell types can be consistently classified regardless of the features used to classify them. In these cases, classically defined cell types typically align well with those identified by analyses of single-cell transcriptomics data, which makes cell annotation straightforward. In other cases, different features could in principle lead to different cell-type classifications, which makes consistent annotation more challenging. Formal ontologies are able to support multiple overlapping classification schemes and can therefore potentially help reconcile different classification schemes, at least at the level of more generally grouped classes.
Cell ontologies also represent developmental lineages and, to a more limited extent, cell states such as activation, cycling, morphological changes and stresses (Fig. 2b) either directly or through extensions of existing annotations. Cell-cycle states, for example, can be represented in the annotation system by combining a term from CL with a term from the Gene Ontology Cell Cycle Phase. Developmental or actively regenerating tissues present particular challenges to cell ontology development, as a plethora of intermediate states and continuous branching lineages can be partitioned. In such a setting, cell annotation needs to emphasize the relative ordering of states or their positions on a continuous differentiation path. There are also striking examples of developmental convergence (developmental homoplasy). Somatosensory neurons, for example, can be of mixed origin from the neural crest or sensory placodes39. Similarly, dermal fibroblasts in different parts of the trunk or face are derived from distinct embryonic lineages despite molecular and phenotypic similarities40. Nevertheless, cell ontologies record gross lineage relationships, with limited temporal resolution between developing/progenitor and mature cell types using specific relations where these relationships are stereotyped and consistent. To date, CL records lineage and differentiation relationships for more than 1,900 cell types and connects developing cell types to developing tissues and stages via links to Uberon.
Many processes that drive cell diversifications, including ontogeny (cell differentiation), morphogenesis (often driven by continuous gradients) and the dual impact of the differentiation history and tissue context of a cell, are imprinted in the molecular properties of the cell and can be captured by hierarchical representations. Therefore, molecular features can serve as the basis for robust cell-type classification that reflects these underlying processes (even when the process is not explicitly known). Currently, cell types and states can be elucidated from single-cell transcriptomics, epigenomics and proteomics expression profiles using different software such as SCCAF41. Further complemented by morphological, physiological, developmental and functional properties, this data-driven framework makes cell annotations comparable across independent ontology efforts and makes the inferred cell types understandable across different communities. Of note, while these inferences are unbiased, it is important to reconcile them with conventional biological and clinical understanding and terminologies.
Current state of ontologies
First developed as platforms to integrate cross-species ontology information, CL and Uberon are now species-neutral ontologies with a strong focus on mammalian cell types and anatomies, with standard mechanisms for recording the species applicability of terms. To date, CL has 2,401 terms covering all major cell types. The granularity of this coverage is variable, with the greatest coverage currently for the immune system (>500 cell types). Uberon defines over 14,000 types of anatomical structures and records many types of relationships between them. In practical terms, CL and Uberon are tightly integrated with each other. Almost 2,000 cell types in CL are linked by ‘part of’ relationships to the anatomical structures defined in Uberon. Further combining CL with newly discovered cell populations from HCA data, we are beginning to extensively cover major organs and cell types in the human body (Table 1).
The human-applicable components of CL and Uberon are under active development as part of multiple collaborative efforts. For human data, terms are being added in a coordinated fashion to both ontology platforms in response to the requests of individual laboratories, as well as to the annotation needs of atlasing projects including the Data Coordination Platform of the HCA2 (https://data.humancellatlas.org) and the Cambridge Cell Atlas portal (www.cambridgecellatlas.org). Editing of CL and Uberon is coordinated by a team of researchers drawn from a growing number of collaborating projects, including the HCA (Chan Zuckerberg Initiative), HuBMAP (National Institutes of Health (NIH)), the Monarch Initiative (NIH) and the Cell Annotation Platform (a collaborative effort funded by Schmidt Futures). This team of editing researchers runs regular open training sessions, and anyone trained to edit the ontology terms can join the editing team. Edits are coordinated and reviewed on GitHub (https://github.com/obophenotype/cell-ontology), with all changes and releases subject to automated quality-control tests before approval. Issues not resolved after discussion on open tickets are coordinated via monthly editor video conferences, which also coordinate the general focus of CL and Uberon efforts. These calls frequently feature guest speakers with a particular interest in extending CL or Uberon in specific areas. CL and Uberon are both members of the Open Biological and Biomedical Ontology Foundry group of ontologies15, which is a loose alliance of ontologies committed to adopting common standards and aligning semantics and ontology infrastructure. All these endow CL and Uberon with the ability to continuously evolve with inputs from various projects and perspectives and to supply formalized ontology information back to the projects (Table 2). Examples of the co-evolution of CL and human cell ontology-building efforts are listed below.
The Brain Data Standards Initiative, which is part of the NIH BRAIN Initiative Cell Census Network, is extending CL with terms for cortical cell types defined by single-cell transcriptomics, with a current focus on the primary motor cortex of human, marmoset and mouse42. This work leverages existing efforts on nomenclature standards43, but importantly aims to use the quantitative hierarchical cell-type classification from single-cell genomics as a data-driven foundation for ontological definitions. Different data types about these cell types are integrated at different levels of the hierarchy, including their spatial tissue distributions, morphological and physiological properties, and axonal projection targets. Ultimately, such a data-driven approach may be used across the entire human body to provide a common metric in gene usage to measure similarities and potential common developmental origins across organs.
The ASCT+B effort44, presented as an accompanying Perspective in this issue, is a HuBMAP, Human Tumor Atlas (HTAN) and HCA community-wide project to build tables representing the human anatomy and cell-type terminology needed for annotating single cell RNA sequencing (scRNA-seq) datasets, and to record expert-approved lists of markers for cell types. Entries in these tables are mapped to existing CL or Uberon terms where possible or turned into term requests for these data resources when new terms are needed. The relationships between cell types and anatomical structures encoded in these tables are validated against CL and Uberon. The results of this validation are relayed to improve the tables, Uberon and CL through discussions and agreement with experts. For example, the ASCT+B project is building an expert-validated ontological model of the human vasculature that is feeding hundreds of new terms and relationships back into Uberon. An important outcome of this work will be a curated subset of CL and Uberon terms to annotate human scRNA-seq data in a reliable manner, both for the healthy HCA data as well as disease samples.
As part of the human cell-focused Sanger–European Bioinformatics Institute (EBI) Cambridge Cell Atlas portal (https://www.cambridgecellatlas.org), an effort to make results from human single-cell gene expression experiments easily accessible to a broad community of users, including clinicians, CL is being enriched and extended based on contributions from pathologists and clinicians. This will introduce human cell types annotated with details of specific immunohistochemical markers that are in routine clinical use in diagnostic pathology. This ontology can then be integrated into the search functionality of the Cambridge Cell Atlas platform to enable searching based on a specific immunohistochemical marker or panel of markers. This will enable the identification of the normal cell type(s) (and potentially pathogenic cell types) that express the marker(s). This functionality could be useful to pathologists in interpreting and contextualizing the range of cell types stained by different immunohistochemical markers on histological sections, cytological preparations or by flow cytometry, and in understanding perturbations in staining patterns in pathological states.
Applications of a cell ontology
Cell ontologies provide the community a single place to look up cell types. Through this portal, knowledge can be aggregated and standardized in an encyclopaedic sense. First, cross-modal data integration can reinforce or refine the identity of a cell type. For example, the survey on the mammalian neocortex revealed the correspondence of various cellular properties when overlapping imaging, electrophysiology and connectivity data with transcriptomics profiles37. Second, mining of an ontological classification system can reveal major trends with respect to shared cell types across organ-specific atlases (for example, immune, stromal and endothelial cells) versus specialized types (for example, goblet cells in the gut and lung). This emphasizes the concept of a tissue being the collective of its cells operating in concert in a specific three-dimensional organization.
Importantly, with more single-cell resources employing the cell and anatomy ontologies, including but not limited to the Fly Cell Atlas, EBI’s Single Cell Expression Atlas and the Sanger–EBI Cambridge Cell Atlas, cell ontologies can link scientific and medical communities through common nomenclatures and markers for human cell biology, pathology and disease. This link, in a broader sense, represents cross-community research whereby a common cell-type reference can be referred. For example, a well-defined cell-type classification of human head and neck tumours, which covered major immune and non-immune cell populations, was utilized as the reference to interrogate the cellular signals contributing to bulk samples of head and neck squamous cell carcinoma from The Cancer Genome Atlas (TCGA)45. This analysis revealed the association of tumour-infiltrating regulatory T cells with improved survival in head and neck cancer45.
At the same time, immunohistochemical markers in routine clinical use (such as those listed by Pathology Outlines, https://www.pathologyoutlines.com/stains.html), which are linked to the non-pathological cell types by the Cambridge Cell Atlas project, could also be curated and further linked to pathological tissues and cell states that express them. This would provide hundreds of antibodies to link cell types and anatomical structures with CL and Uberon, albeit with a focus on pathological states (CL and Uberon currently focus on healthy homeostatic states).
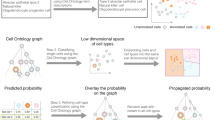
The application of cell ontologies will be most pertinent in the context of interactive and automated systems for the interpretation and annotation of single-cell genomics datasets. A number of efforts to design such systems are under way, including automated cell annotation projection pipelines46,47,48,49,50,51,52. For example, as part of the HCA initiative, the Cell Annotation Platform (CAP) aims to provide a general repository for cell annotations of different datasets in combination with interactive tools for annotating new datasets. For a cell of interest, CAP user interfaces will suggest the appropriate ontology terms based on text search, learned synonyms and eventually molecular signatures themselves. Where no appropriate term is available from CL, free-text annotation will be used as the basis for the addition of a new term to CL. Similarly, the HuBMAP data portal assigns cell annotations to scRNA-seq datasets with an Azimuth-based label transfer procedure49 based on a vocabulary of cell types from CL, which aims to assess cellular diversities at different levels of resolution. With an initial focus on immune cells, CellTypist uses an expandable cross-tissue cell reference before predicting cell identities with a logistic regression-based label transfer pipeline, with all derived cell types directly interpretable by CL48. Conversely, the resulting knowledge base of commonly used annotation terms and associated molecular signatures will provide a useful resource to extend ontologies as well as to train and optimize machine-learning models that automate the annotation task. In parallel to these efforts, data-driven ontology development is advancing community engagement in specific research domains such as Neuroscience Multi-Omic (NeMO) Analytics for the brain (https://nemoanalytics.org) and gene expression analysis resource (gEAR) for the ear53.
Summary and outlook
Resolving the cellular makeup of the human body warrants the categorization of cells in a standardized framework. The CL database offers one such avenue to consolidating this knowledge in an encyclopaedic manner, with applications from cell and tissue biology all the way to the clinic. Despite potential cell classification ambiguities and transient cellular states, each facet of a cell, ranging from morphological to molecular features, can be taken into account until a defining status is reached and recognized by the community.
Many HCA-related resources, such as cellxgene54, have been using CL for de novo cell annotation. Cell ontologies serve other sources of data by retrieving or delivering ontology-level information. We anticipate that the synergy between the HCA project and CL will continue to grow over the coming years and beyond the completion of HCA, with dimensions of human genetic variation, ageing and disease on the horizon. HCA single-cell omics data provide a foundation for the development of cell ontologies, which are powerful resources to define cell types that are universal across the entire body or specific to subsets of tissues and will facilitate future research. This will become more pressing and clearer as the number of HCA studies of individual tissues and organs increases. The HCA Biological Networks will provide nucleation points for expert community efforts to achieve gold standard, consensus cell annotations with cell ontology terms. With such a quantitative approach, common phenotypes and developmental origins of cell types will become understandable through shared gene usage, and functional similarities will be revealed in gene patterns. Whole-body consequences of disease will be understandable through differential gene usage in differently located cells. This will create opportunities for a new and different kind of quantitative data-driven framework that extends and potentially transforms existing ontology efforts.
References
Regev, A. et al. The Human Cell Atlas. eLife 6, e27041 (2017).
Rozenblatt-Rosen, O., Stubbington, M. J. T., Regev, A. & Teichmann, S. A. The Human Cell Atlas: from vision to reality. Nature 550, 451–453 (2017).
Aldridge, S. & Teichmann, S. A. Single cell transcriptomics comes of age. Nat. Commun. 11, 4307 (2020).
Popescu, D. M. et al. Decoding human fetal liver haematopoiesis. Nature 574, 365–371 (2019).
Madissoon, E. et al. scRNA-seq assessment of the human lung, spleen, and esophagus tissue stability after cold preservation. Genome Biol. 21, 1 (2019).
Hagai, T. et al. Gene expression variability across cells and species shapes innate immunity. Nature 563, 197–202 (2018).
Vento-Tormo, R. et al. Single-cell reconstruction of the early maternal–fetal interface in humans. Nature 563, 347–353 (2018).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
Drokhlyansky, E. et al. The human and mouse enteric nervous system at single-cell resolution. Cell 182, 1606–1622.e23 (2020).
Elmentaite, R. et al. Cells of the human intestinal tract mapped across space and time. Nature 597, 250–255 (2021).
Bodenreider, O. & Stevens, R. Bio-ontologies: current trends and future directions. Brief. Bioinform 7, 256–274 (2006).
Diehl, A. D. et al. The Cell Ontology 2016: enhanced content, modularization, and ontology interoperability. J. Biomed. Semant. 7, 44 (2016).
Costa, M., Reeve, S., Grumbling, G. & Osumi-Sutherland, D. The Drosophila anatomy ontology. J. Biomed. Semant. 4, 32 (2013).
Consortium, E. P. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Smith, B. et al. The OBO Foundry: coordinated evolution of ontologies to support biomedical data integration. Nat. Biotechnol. 25, 1251–1255 (2007).
Milyaev, N. et al. The Virtual Fly Brain browser and query interface. Bioinformatics 28, 411–415 (2012).
Osumi-Sutherland, D., Costa, M., Court, R. & O’Kane, C. J. Virtual Fly Brain—using OWL to support the mapping and genetic dissection of the Drosophila brain. CEUR Workshop Proc. 1265, 85–96 (2014).
Li, H. et al. Fly Cell Atlas: a single-cell transcriptomic atlas of the adult fruit fly. Preprint at bioRxiv https://doi.org/10.1101/2021.07.04.451050 (2021).
Gene Ontology, C. The Gene Ontology resource: enriching a GOld mine. Nucleic Acids Res. 49, D325–D334 (2021).
Mungall, C. J. et al. The Monarch Initiative: an integrative data and analytic platform connecting phenotypes to genotypes across species. Nucleic Acids Res. 45, D712–D722 (2017).
Jacqz, E., Branch, R. A., Heidemann, H. & Aujard, Y. [Prevention of nephrotoxicity of amphotericin B during the treatment of deep candidiasis]. Ann. Biol. Clin. (Paris) 45, 689–693 (1987).
Hu, B. C. The human body at cellular resolution: the NIH Human Biomolecular Atlas Program. Nature 574, 187–192 (2019).
Ecker, J. R. et al. The BRAIN Initiative Cell Census Consortium: lessons learned toward generating a comprehensive brain cell atlas. Neuron 96, 542–557 (2017).
Young, M. D. et al. Single-cell transcriptomes from human kidneys reveal the cellular identity of renal tumors. Science 361, 594–599 (2018).
Sungnak, W. et al. SARS-CoV-2 entry factors are highly expressed in nasal epithelial cells together with innate immune genes. Nat. Med. 26, 681–687 (2020).
Muus, C. et al. Single-cell meta-analysis of SARS-CoV-2 entry genes across tissues and demographics. Nat. Med. 27, 546–559 (2021).
Delorey, T. M. et al. COVID-19 tissue atlases reveal SARS-CoV-2 pathology and cellular targets. Nature 595, 107–113 (2021).
Mungall, C. J., Torniai, C., Gkoutos, G. V., Lewis, S. E. & Haendel, M. A. Uberon, an integrative multi-species anatomy ontology. Genome Biol. 13, R5 (2012).
Bernard, A., Boumsell, L., Daussett, J., Milstein, C. & Schlossman, S. F. (eds) Leucocyte Typing: Human Leucocyte Differentiation Antigens Detected by Monoclonal Antibodies: Specification, Classification, Nomenclature = Typage Leucocytaire: Antigènes de Différenciation Leococytaire Humains Révélés par les Anticorps Monoclonaux (Springer, 1984).
Reinherz, E. L., Haynes, B. F., Nadler, L. M. & Bernstein, I. D. (eds) Leukocyte Typing II (Springer, 1986).
McMichael, A. J. (ed.) Leucocyte Typing III: White Cell Differentiation Antigens (Oxford Univ. Press, 1987).
Knapp, W. et al. (eds) Leucocyte Typing IV: White Cell Differentiation Antigens (Oxford Univ. Press, 1989).
Schlossman, S. F. (ed.) Leucocyte typing V: white cell differentiation antigens. In Proc. Fifth International Workshop and Conference held in Boston, USA, 3–7 November, 1993 (Oxford Univ. Press, 1995).
Kishimoto, T. (ed.) Leucocyte typing VI: white cell differentiation antigens. In Proc. Sixth International Workshop and Conference held in Kobe, Japan, 10–14 November 1996 (Garland, 1998).
Mason, D. (ed.) Leucocyte typing VII: white cell differentiation antigens. In Proc. Seventh International Workshop and Conference held in Harrogate, United Kingdom (Oxford University Press: Oxford, 2002).
Zola, H., Swart, B., Nicholson, I. & Voss, E. Leukocyte and Stromal Cell Molecules: The CD Markers (Wiley-Liss, 2007).
Yuste, R. et al. A community-based transcriptomics classification and nomenclature of neocortical cell types. Nat. Neurosci. 23, 1456–1468 (2020).
Shekhar, K. et al. Comprehensive classification of retinal bipolar neurons by single-cell transcriptomics. Cell 166, 1308–1323.e30 (2016).
Vermeiren, S., Bellefroid, E. J. & Desiderio, S. Vertebrate sensory ganglia: common and divergent features of the transcriptional programs generating their functional specialization. Front. Cell Dev. Biol. 8, 587699 (2020).
Driskell, R. R. & Watt, F. M. Understanding fibroblast heterogeneity in the skin. Trends Cell Biol. 25, 92–99 (2015).
Miao, Z. et al. Putative cell type discovery from single-cell gene expression data. Nat. Methods 17, 621–628 (2020).
Bakken, T. E. et al. Comparative cellular analysis of motor cortex in human, marmoset and mouse. Nature 598, 111–119 (2021).
Miller, J. A. et al. Common cell type nomenclature for the mammalian brain. eLife 9, e59928 (2020).
Börner, K. et al. Anatomical structures, cell types and biomarkers of the Human Reference Atlas. Nat. Cell Biol. https://doi.org/10.1038/s41556-021-00788-6 (2021).
Qi, Z. et al. Single-cell deconvolution of head and neck squamous cell carcinoma. Cancers (Basel) 13, 2387 (2021).
Kiselev, V. Y., Yiu, A. & Hemberg, M. scmap: projection of single-cell RNA-seq data across data sets. Nat. Methods 15, 359–362 (2018).
Kimmel, J. C. & Kelley, D. R. Semi-supervised adversarial neural networks for single-cell classification. Genome Res. 31, 1781–1793 (2021).
Domínguez, C.C. et al. Cross-tissue immune cell analysis reveals tissue-specific adaptations and clonal architecture across the human body. Preprint at bioRxiv https://doi.org/10.1101/2021.04.28.441762 (2021).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Bernstein, M. N., Ma, Z., Gleicher, M. & Dewey, C. N. CellO: comprehensive and hierarchical cell type classification of human cells with the Cell Ontology. iScience 24, 101913 (2021).
Hou, R., Denisenko, E. & Forrest, A. R. R. scMatch: a single-cell gene expression profile annotation tool using reference datasets. Bioinformatics 35, 4688–4695 (2019).
Orvis, J. et al. gEAR: gene expression analysis resource portal for community-driven, multi-omic data exploration. Nat. Methods 18, 843–844 (2021).
Megill, C. et al. cellxgene: a performant, scalable exploration platform for high dimensional sparse matrices. Preprint at bioRxiv https://doi.org/10.1101/2021.04.05.438318 (2021).
Stewart, B. J. et al. Spatiotemporal immune zonation of the human kidney. Science 365, 1461–1466 (2019).
James, K. R. et al. Distinct microbial and immune niches of the human colon. Nat. Immunol. 21, 343–353 (2020).
Vieira Braga, F. A. et al. A cellular census of human lungs identifies novel cell states in health and in asthma. Nat. Med. 25, 1153–1163 (2019).
Travaglini, K. J. et al. A molecular cell atlas of the human lung from single-cell RNA sequencing. Nature 587, 619–625 (2020).
Ramachandran, P. et al. Resolving the fibrotic niche of human liver cirrhosis at single-cell level. Nature 575, 512–518 (2019).
Aizarani, N. et al. A human liver cell atlas reveals heterogeneity and epithelial progenitors. Nature 572, 199–204 (2019).
Litvinukova, M. et al. Cells of the adult human heart. Nature 588, 466–472 (2020).
Park, J. E. et al. A cell atlas of human thymic development defines T cell repertoire formation. Science 367, eaay3224 (2020).
Reynolds, G. et al. Developmental cell programs are co-opted in inflammatory skin disease. Science 371, eaba6500 (2021).
Garcia-Alonso, L. et al. Mapping the temporal and spatial dynamics of the human endometrium in vivo and in vitro. Nat. Genetics https://doi.org/10.1038/s41588-021-00972-2 (2021).
Acknowledgements
We are grateful to J. Eliasova (scientific illustrator) for support with the figure, to R. Vento-Tormo for comments on the figure and texts, and to the following clinicians and researchers for information on standard pathology markers for tissues and cells: L. Campos, A. Dean, L. Moore, N. Sebire, T. Brevini, M. Haniffa, J. E. Kwa, J. McCaffrey and A. Kreins. We also thank all members of the CL and Uberon editorial teams, including C. Mungall, N. Matentzoglu, A. Diehl, N. Washington, S. Tan, P. Roncaglia, T. Lubiana and D. Goutte-Gattat. Research reported in this publication was supported by the Wellcome Trust (grant 108413/A/15/D), the Office of the Director, National Institutes of Health of the National Institutes of Health (under award number OT2OD026682’), grants from the CZI (Chan Zuckerberg Initiative DAF, an advised fund of Silicon Valley Community Foundation), and Schmidt Futures (Grant 74). This publication is part of the HCA (www.humancellatlas.org/publications/).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Since January 2019, S.A.T. has been remunerated for consulting and SAB membership by Foresite Labs, GlaxoSmithKline, Biogen, Roche and Genentech, and is a founder and equity holder of Transition Bio. A.R. is a cofounder and equity holder in Celsius Therapeutics, an equity holder in Immunitas Therapeutics, and was a scientific advisory board member for ThermoFisher Scientific, Asimov, Syros Pharmaceuticals and Neogene Therapeutics until 31 July 2020. From 1 August 2020, A.R. is an employee of Genentech, a member of the Roche group. A.R. is a named inventor on several patents and patent applications filed by the Broad Institute in the area of single-cell and spatial genomics. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Osumi-Sutherland, D., Xu, C., Keays, M. et al. Cell type ontologies of the Human Cell Atlas. Nat Cell Biol 23, 1129–1135 (2021). https://doi.org/10.1038/s41556-021-00787-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-021-00787-7
This article is cited by
-
Challenges and perspectives in computational deconvolution of genomics data
Nature Methods (2024)
-
Cellular development and evolution of the mammalian cerebellum
Nature (2024)
-
Unsupervised and supervised discovery of tissue cellular neighborhoods from cell phenotypes
Nature Methods (2024)
-
Integrating single-cell multi-omics and prior biological knowledge for a functional characterization of the immune system
Nature Immunology (2024)
-
Representing and extracting knowledge from single-cell data
Biophysical Reviews (2024)