Abstract
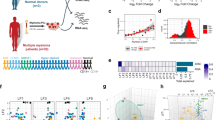
While there is extensive evidence for genetic variation as a basis for treatment resistance, other sources of variation result from cellular plasticity. Using multiple myeloma as an example of an incurable lymphoid malignancy, we show how cancer cells modulate lineage restriction, adapt their enhancer usage and employ cell-intrinsic diversity for survival and treatment escape. By using single-cell transcriptome and chromatin accessibility profiling, we show that distinct transcriptional states co-exist in individual cancer cells and that differential transcriptional regulon usage and enhancer rewiring underlie these alternative transcriptional states. We demonstrate that exposure to standard treatment further promotes transcriptional reprogramming and differential enhancer recruitment while simultaneously reducing developmental potential. Importantly, treatment generates a distinct complement of actionable immunotherapy targets, such as CXCR4, which can be exploited to overcome treatment resistance. Our studies therefore delineate how to transform the cellular plasticity that underlies drug resistance into immuno-oncologic therapeutic opportunities.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The data collected in this paper include next-generation sequencing (NGS) data. The following NGS datasets were produced: (1) scRNA-seq datasets from primary myeloma and CD45+ immune cells and normal donor bone marrow plasma cells and CD45+ immune cells; (2) scRNA-seq datasets from the MMCL MOLP2 treated with PVD or DMSO; (3) scATAC-seq datasets from primary myeloma and normal donor plasma cells; (4) bulk ATAC-seq datasets from the MMCL MOLP2 treated with PVD or DMSO; (5) CUT&Tag data for H3K27 acetylation from the MMCL MOLP2 treated with PVD or DMSO. FASTQ files were generated following Illumina sequencing and further analysed as described below and in the Methods. The scRNA-seq and ATAC-seq datasets and CUT&Tag data generated in this study have been deposited in the Gene Expression Omnibus under the accession number GSE162337. The cytoscape files for the final GRNs are available for download at the Broad Institute Single Cell Portal (https://singlecell.broadinstitute.org/single_cell/study/SCP1511/dynamic-transcriptional-reprogramming-leads-to-novel-immunotherapeutic-vulnerabilities-in-myeloma). Previously published IRF4 Chip-seq data that were re-analysed here are available under accession code PRJEB25605 from the European Nucleotide Archive. Previously published H3K27 acetylation ChIP-seq data that were re-analysed here are available from the European Nucleotide Archive under accession codes PRJEB25605 (for patient samples) and PRJNA608682 (for normal plasma cells). Additionally, we used the following publicly available datasets: the dependency scores available through the depmap download portal (https://depmap.org/portal/download/); the BLUEPRINT dataset12; and the JASPAR motif database. Source data are provided with this paper. All other data supporting the findings of this study are available from the corresponding authors on reasonable request.
Code availability
The following tools are available on github: trimmomatic, STAR aligner, HTSeq, RSEM and the nfcore/atacseq pipeline v.1.0.0; for downstream analysis, R (v.3.6.1 or 4.0) and python (v.2.7.2 or 3.6.0); and the packages mvoutlier (v.2.1.1), PAGODA2 (v.0.1.1), SingleR (v.1.0.1), scran (1.12.1), InferCNV (1.4.0), CONICSmat (v.0.0.0.1), MiXCR, randomFOREST (v.4.6-14), SCANPY (v.1.4.6), CytoTRACE (v.0.1.0), PROGENy (1.6.0), LandSCENT (0.99.3), SCENIC (1.1.2-2), Seurat (v.3.2.1), velocyto, scvelo (v.0.2.0), TissueEnrich (v.1.4.1), Cicero (v.1.2.0), chromVAR (v.1.6.0), GVIZ (v.1.28.3), deeptools (v.3.0.2), ArchR (v.0.9.5), edgeR (v.3.32.0), NMF (v.0.23.0), BRIE2 (v.2) and MISO (v.0.5.4); also FlowJo (v.10), cytoscape (v.3.8.0), GREAT (http://great.stanford.edu/public/html/) and GSEA (https://www.gsea-msigdb.org/gsea/index.jsp).
References
Sharma, S. V. et al. A chromatin-mediated reversible drug-tolerant state in cancer cell subpopulations. Cell 141, 69–80 (2010).
Hangauer, M. J. et al. Drug-tolerant persister cancer cells are vulnerable to GPX4 inhibition. Nature 551, 247–250 (2017).
Shaffer, S. M. et al. Rare cell variability and drug-induced reprogramming as a mode of cancer drug resistance. Nature 546, 431–435 (2017).
Risom, T. et al. Differentiation-state plasticity is a targetable resistance mechanism in basal-like breast cancer. Nat. Commun. 9, 3815 (2018).
Verfaillie, A. et al. Decoding the regulatory landscape of melanoma reveals TEADS as regulators of the invasive cell state. Nat. Commun. 6, 6683 (2015).
Wouters, J. et al. Robust gene expression programs underlie recurrent cell states and phenotype switching in melanoma. Nat. Cell Biol. 22, 986–998 (2020).
LaFave, L. M. et al. Epigenomic state transitions characterize tumor progression in mouse lung adenocarcinoma. Cancer Cell 38, 212–228.e13 (2020).
Paiva, B. et al. Differentiation stage of myeloma plasma cells: biological and clinical significance on behalf of GEM/PETHEMA cooperative study groups Europe PMC Funders Group. Leukemia 31, 382–392 (2017).
Joseph, D., Gonsky, J. P. & Blain, S. W. Macrophage inhibitory factor-1 (MIF-1) controls the plasticity of multiple myeloma tumor cells. PLoS ONE 13, e0206368 (2018).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Fan, J. et al. Characterizing transcriptional heterogeneity through pathway and gene set overdispersion analysis. Nat. Methods 13, 241–244 (2016).
Fernández, J. M. et al. The BLUEPRINT data analysis portal. Cell Syst. 3, 491–495.e5 (2016).
Ledergor, G. et al. Single cell dissection of plasma cell heterogeneity in symptomatic and asymptomatic myeloma. Nat. Med. 24, 1867–1876 (2018).
Broyl, A. et al. Gene expression profiling for molecular classification of multiple myeloma in newly diagnosed patients. Blood 116, 2543–2553 (2010).
Kinker, G. S. et al. Pan-cancer single-cell RNA-seq identifies recurring programs of cellular heterogeneity. Nat. Genet. 52, 1208–1218 (2020).
Hay, S. B., Ferchen, K., Chetal, K., Grimes, H. L. & Salomonis, N. The human cell atlas bone marrow single-cell interactive web portal. Exp. Hematol. 68, 51–61 (2018).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Gulati, G. S. et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science 367, 405–411 (2020).
Teschendorff, A. E. & Enver, T. Single-cell entropy for accurate estimation of differentiation potency from a cell’s transcriptome. Nat. Commun. 8, 15599 (2017).
Chen, W. et al. Single-cell landscape in mammary epithelium reveals bipotent-like cells associated with breast cancer risk and outcome. Commun. Biol. 2, 306 (2019).
Setz, C. S. et al. PI3K-mediated Blimp-1 activation controls B cell selection and homeostasis. Cell Rep. 24, 391–405 (2018).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Kas, K. et al. ESE-3, a novel member of an epithelium-specific ets transcription factor subfamily, demonstrates different target gene specificity from ESE-1. J. Biol. Chem. 275, 2986–2998 (2000).
Joshi, S. et al. TEAD transcription factors are required for normal primary myoblast differentiation in vitro and muscle regeneration in vivo. PLoS Genet. 13, e1006600 (2017).
Goenawan, I. H., Bryan, K. & Lynn, D. J. DyNet: visualization and analysis of dynamic molecular interaction networks. Bioinformatics 32, 2713–2715 (2016).
Cuitiño, M. C. et al. Two distinct E2F transcriptional modules drive cell cycles and differentiation. Cell Rep. 27, 3547–3560.e5 (2019).
Fulciniti, M. et al. Non-overlapping control of transcriptome by promoter- and super-enhancer-associated dependencies in multiple myeloma. Cell Rep. 25, 3693–3705.e6 (2018).
Jin, Y. et al. Active enhancer and chromatin accessibility landscapes chart the regulatory network of primary multiple myeloma. Blood 131, 2138–2150 (2018).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Fulco, C. P. et al. Activity-by-contact model of enhancer-promoter regulation from thousands of CRISPR perturbations. Nat. Genet. 51, 1664–1669 (2019).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Salomonis, N. et al. Alternative splicing regulates mouse embryonic stem cell pluripotency and differentiation. Proc. Natl Acad. Sci. USA 107, 10514–10519 (2010).
Lee, J. H., Kwon, E. J. & Kim, D. H. Calumenin has a role in the alleviation of ER stress in neonatal rat cardiomyocytes. Biochem. Biophys. Res. Commun. 439, 327–332 (2013).
Campbell, K. S., Cohen, A. D. & Pazina, T. Mechanisms of NK cell activation and clinical activity of the therapeutic SLAMF7 antibody, elotuzumab in multiple myeloma. Front. Immunol. 9, 2551 (2018).
Roccaro, A. M. et al. CXCR4 regulates extra-medullary myeloma through epithelial–mesenchymal-transition-like transcriptional activation. Cell Rep. 12, 622–635 (2015).
Ghobrial, I. M. et al. A Phase Ib/II trial of the first-in-class anti-CXCR4 antibody ulocuplumab in combination with lenalidomide or bortezomib plus dexamethasone in relapsed multiple myeloma. Clin. Cancer Res. 26, 344–353 (2020).
DiPersio, J. F. et al. Plerixafor and G-CSF versus placebo and G-CSF to mobilize hematopoietic stem cells for autologous stem cell transplantation in patients with multiple myeloma. Blood 113, 5720–5726 (2009).
Mandal, M. et al. CXCR4 signaling directs Igk recombination and the molecular mechanisms of late B lymphopoiesis. Nat. Immunol. 20, 1393–1403 (2019).
Barwick, B. G., Gupta, V. A., Vertino, P. M. & Boise, L. H. Cell of origin and genetic alterations in the pathogenesis of multiple myeloma. Front. Immunol. 10, 1121 (2019).
Ordonez, R. et al. Chromatin activation as a unifying principle underlying pathogenic mechanisms in multiple myeloma. Genome Res. 30, 1217–1227 (2020).
Flavahan, W. A., Gaskell, E. & Bernstein, B. E. Epigenetic plasticity and the hallmarks of cancer. Science 357, eaal2380 (2017).
Rauscher, B., Henkel, L., Heigwer, F. & Boutros, M. Lineage specific core-regulatory circuits determine gene essentiality in cancer cells. Preprint bioRxiv https://doi.org/10.1101/609552 (2019).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Lun, A. T., McCarthy, D. J. & Marioni, J. C. A step-by-step workflow for low-level analysis of single-cell RNA-seq data with Bioconductor. F1000Research 5, 2122 (2016).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Muller, S., Cho, A., Liu, S. J., Lim, D. A. & Diaz, A. CONICS integrates scRNA-seq with DNA sequencing to map gene expression to tumor sub-clones. Bioinformatics 34, 3217–3219 (2018).
Bolotin, D. A. et al. MiXCR: software for comprehensive adaptive immunity profiling. Nat. Methods 12, 380–381 (2015).
Gaujoux, R. & Seoighe, C. A flexible R package for nonnegative matrix factorization. BMC Bioinformatics 11, 367 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297 (2012).
Puram, S. V. et al. Single-cell transcriptomic analysis of primary and metastatic tumor ecosystems in head and neck cancer. Cell 171, 1611–1624.e24 (2017).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Liberzon, A. et al. The molecular signatures database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Aran, D., Hu, Z. & Butte, A. J. xCell: digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 18, 220 (2017).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Schubert, M. et al. Perturbation-response genes reveal signaling footprints in cancer gene expression. Nat. Commun. 9, 20 (2018).
Anand, P. et al. Single-cell RNA-seq reveals developmental plasticity with coexisting oncogenic states and immune evasion programs in ETP-ALL. Blood 137, 2463–2480 (2021).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Huang, Y. & Sanguinetti, G. BRIE2: computational identification of splicing phenotypes from single-cell transcriptomic experiments. Genome Biol. 22, 251 (2021).
Katz, Y., Wang, E. T., Airoldi, E. M. & Burge, C. B. Analysis and design of RNA sequencing experiments for identifying isoform regulation. Nat. Methods 7, 1009–1015 (2010).
Wang, E. T. et al. Alternative isoform regulation in human tissue transcriptomes. Nature 456, 470–476 (2008).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 53, 403–411 (2021).
Ewels, P. A. et al. The nf-core framework for community-curated bioinformatics pipelines. Nat. Biotechnol. 38, 276–278 (2020).
Jia, Y. et al. Myeloma-specific superenhancers affect genes of biological and clinical relevance in myeloma. Blood Cancer J. 11, 32 (2021).
Kaya-Okur, H. S. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat. Commun. 10, 1930 (2019).
Pliner, H. A. et al. Cicero predicts cis-regulatory DNA interactions from single-cell chromatin accessibility data. Mol. Cell 71, 858–871.e8 (2018).
Acknowledgements
The authors would like to thank A. Letai for discussion and critical comments on the manuscript. This project was supported by grants from the National Cancer Institute (K08CA191026 to J.G.L.), (K08CA191091 to B.K.), the Anna Fuller Fund (to J.G.L.) and a Scholar Award by the V Foundation for Cancer Research (to J.G.L.). K.C.A. is an American Cancer Society Clinical Research Professor. J.F. acknowledges support from the German Research Foundation (DFG) and the American Association for Cancer Research (AACR).
Author information
Authors and Affiliations
Contributions
J.F. designed and performed experiments, analysed data and wrote the manuscript. P.A., R.A.P., N.S. and H.S. performed data analyses. M.S.N. performed experiments and data analyses. T.V., J.M.W., S.P., J.A.K., A.K. and V.D. performed sample processing, sequencing or provided scientific input. N.S.R. and A.J.Y. provided project leadership, coordinated sample acquisition and clinical data integration. M.M., K.C.A., P.G.R. and J.P.L. oversaw clinical data analyses and integration. B.K. and J.G.L. designed experiments, provided project leadership, supervised the analysis and wrote the manuscript. All authors discussed the results and reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.J.Y. has a consulting role for Adaptive Biotechnologies, Amgen, BMS, Celgene, GSK, Janssen, Karyopharm, Sanofi and Takeda. The remaining authors declare no competing interests.
Additional information
Peer review statement Nature Cell Biology thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Sorting strategy.
a) Sorting strategy for myeloma cells and normal donor plasma cells with representative flow cytometry plots. CD38+ CD138+ cells were sorted after EasySep enrichment of CD138+ cells from bone marrow or peripheral blood. b) Sorting strategy for immune cell subsets. CD3+, CD19+, CD14+ and CD45+ Lin- cells were sorted following exclusion of dead cells and doublets. c) Sorting strategy for NK cells. CD56+ CD16+ cells (Q1-3) were sorted after exclusion of dead cells, doublets, CD3+ cells and CD138+ cells.
Extended Data Fig. 2 Quality assessment and filtering of single cell RNA data.
a) Distribution of library size (i), number of detected genes (ii), percentage of counts mapping to mitochondrial genes (iii), and percentage of counts mapping to house-keeping genes (HKGs) (iv) per cell. b) Scatter plot depicting principal component analysis using the top two dimensions. The PCA was performed on the four features depicted in a) for all cells in the unfiltered dataset. The outliers, highlighted in orange, were identified using the mvoutlier package. c) The distribution of features shown in a) after filtering out cells identified as outliers. d) Scatter plot depicting expression frequency vs mean read counts per gene in the filtered dataset. e) Density plot showing the contribution of various technical factors (timepoint, run date, individual, total features, total counts, percentage of counts mapping to mitochondrial genes, and percentage of counts mapping to house-keeping genes (HKGs)) to the total variation observed in the dataset.
Extended Data Fig. 3 Characterization of scRNA-Seq dataset of primary myeloma cells.
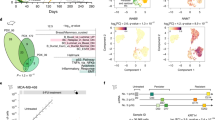
a) Patient characteristics: Genetic aberrations detected by FISH. b) Copy number profiles of myeloma patients were inferred from scRNA expression data using InferCNV. Normal donor plasma cells served as a control. See the enlarged version in Supplementary Information. c-e) Random forest model identifying genes that best discriminate myeloma from normal plasma cells. c) Plot depicting the error vs number of trees used by random forest model on malignant (green), non-malignant (red) and combined (black) cells. d) Relative importance of each gene in the model (mean decrease in Gini coefficient). e) Confusion matrix showing classification and error rates during training of the model, for prediction on the training set (predict_train) and the test set (predict_test). f) Detailed heatmap showing classification of malignant and normal plasma cells based on CNVs, CDR3 sequence, expression of translocation targets, and genes identified by random forest (RF) model that best discriminate between normal and malignant cells, and composite malignancy score for patient MM5. g) tSNE plots showing PAGODA2 clustering highlighting cells from patient MM5 with malignancy scores ≥ 1 or < 1. h) Heatmap with relative expression of marker genes for individual patients in single myeloma and normal plasma cells.
Extended Data Fig. 4 Transcriptional diversity is increased in MM.
a) tSNE plots showing expression of selected genes not normally expressed in plasma cells (log2-transformed counts). Depicted are CD20 (MS4A1), the B Lymphoid Tyrosine Kinase BLK, usually expressed in earlier stages of B cell development and the myeloid restricted serine protease C (CTSC). b) Heatmap with lineage scores from single cell RNA-Seq derived datasets from the Human Cell Atlas in single myeloma cells from patients, single plasma cells from normal donors or single B cells. HSC, hematopoietic stem cell; CLP, common lymphoid progenitor; CMP, common myeloid progenitor; MKP, MK progenitor; ERP, ER progenitor; CD34 Gran, CD34 + Granulocyte progenitor, MixLin, mixed lineage progenitor; PreB, Pre-B cell; ProB, Pro-B cell; PC, plasma cell. c) Heatmap showing relative expression of individual genes from different lineages from the Human Cell Atlas dataset in single myeloma cells, single normal donor plasma cells or single B cells. The enlarged heatmaps in b-c are provided in Supplementary Information. d) UMAP colored by patient. e) RNA velocity estimates of single myeloma cells and normal donor plasma cells projected onto two-dimensional UMAP. Normal donor plasma cells are indicated in red. f) Cells colored based on cell cycle phase. g) Number of genes detected per cell in single myeloma vs normal donor plasma cells. Boxplots show the median and interquartile range, whiskers extend to 1.5x the interquartile range. n = 1,162 cells. p < 2.2e-16 by two-sided Wilcoxon-test. h,i) Entropy is increased in myeloma vs normal donor plasma cells. Boxplots show the median and interquartile range, whiskers extend to 1.5x the interquartile range. n = 1,162 cells. p < 2.2e-16 by two-sided Wilcoxon-test. j) Validation of entropy using an alternative protein-protein interaction network. Boxplots show the median and interquartile range, whiskers extend to 1.5x the interquartile range. n = 1,162 cells. p < 2.2e-16 by two-sided Wilcoxon-test. k) Predicted ordering by CytoTRACE, which orders MM cells based on their developmental potential from most mature (lowest values) to most immature (highest values). Boxplots show the median and interquartile range, whiskers extend from min to max. l) tSNE plots showing expression of plasma cell lineage transcription factors XBP1, IRF4, PRDM1, FOS, POU2AF1 and ZBTB20 (log2-transformed counts) in single myeloma cells, normal donor plasma cells and B cells.
Extended Data Fig. 5 Gene regulatory network activity in different cell types.
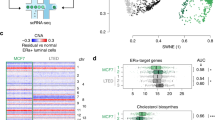
a-d) Gene regulatory network activity for different cell types was determined from the BLUEPRINT dataset. a) Gene expression in normal plasma cells. b) Gene expression in hematopoietic stem cells (HSCs). c) Gene expression in CLP (common lymphoid progenitor) population. d) Gene expression in macrophages. e) Network layouts for normal hematopoiesis based on the BLUEPRINT dataset (top) and based on our single-cell RNA data (bottom) illustrating extensive rewiring, gain of new connections and changes in relative activity. Edges are colored based on regulon activity where high activity is indicated in red, low activity in blue. Target genes are depicted in white, transcription factors which are not among regulons are shown in green. f) Changes in regulon activity in myeloma compared to normal donor plasma cells are projected onto the network. Transcription factors with the largest difference in regulon activity in myeloma compared to normal donor plasma cells are highlighted in insets. g) tSNE plots showing regulon activity (area under the curve, AUC) of plasma cell lineage transcription factors XBP1, IRF4, PRDM1, and FOS in single myeloma and normal plasma cells.
Extended Data Fig. 6 Quality assessment and filtering of single cell ATAC data.
a) Filtering of single cell ATAC profiles based on TSS enrichment and number of unique nuclear fragments. b) Fragment length distribution of filtered scATAC profiles showing characteristic distribution with nucleosome-free region and mononucleosome peaks. c) TSS enrichment after filtering per sample. d) Number of unique fragments per sample. e,f) tSNE plot colored by clusters (e) or individual (f). g,h) tSNE plots following batch effect removal by Harmony colored by clusters (g) or individual (h). i-k) Defining cluster identities following integration with scRNA-seq data. i) tSNE plot colored by predicted cell type identities. j) tSNE plot showing cell type identities by cluster. k) Heatmap showing confusion matrix for predicted cell type identities by cluster. l-o) Tracks with aggregated ATAC profiles for each cluster for marker genes SDC1, LYZ, MS4A1 and FCGR3A, respectively. p-s) tSNE plots colored by gene scores for marker genes SDC1, LYZ, MS4A1 and FCGR3A, respectively.
Extended Data Fig. 7 Annotation of peaks from aggregated scATAC data.
a) Processing of scATAC profiles. b) Intersection of peaks in aggregate scATAC samples showing the overlap of peaks between MM and normal donor (ND) samples. c) Number of peaks significantly different by DESeq2 enriched in ND or MM (FDR ≤ 0.1, absolute log2FC ≥ 1). d) Heatmap showing differentially accessible peaks from c) in ND and MM, sorted by the ND sample. e) ChromHMM state annotation in aggregate scATAC samples. Depicted is the fraction of peaks in each of the indicated states. f) Number of peaks in ChromHMM state 13 corresponding to heterochromatin. **p = 0.0029 by two-sided t-test. g) Fraction of peaks in indicated genomic regions. h) Boxplot comparing distance to TSS in ND vs MM. ****p < 2.2e-16 by two-sided Wilcoxon test. Boxplots show the median and interquartile range, whiskers extend to 1.5x the interquartile range. n = 46,935 peaks and 68,752 peaks, respectively. i) Heatmap showing accessibility of enhancers (n = 15,748) associated with the top multi-enhancer genes, sorted by the ND sample. j) Barplot showing genes with ≥ 2 enhancer interactions by ABC model. k) Heatmap showing multi-enhancer genes by ABC model (n = 16,635), sorted by the ND sample. l) Barplot showing gene set enrichment analysis for multi-enhancer genes in MM by ABC model. m) Barplot showing differential motif enrichment analysis of the single cell ATAC-Seq dataset comparing myeloma cells with normal donor plasma cells. Shown are the top 40 differentially enriched transcription factor motifs ordered by FDR. n) Venn diagram showing overlap of differentially enriched motifs and rewired transcription factors determined by DyNet algorithm.
Extended Data Fig. 8 Alternative splicing following treatment in MM.
a) Quantification of exon inclusion/exclusion ratio (percent of spliced in, psi). Volcano plot showing differential splicing at C5D1 vs screening timepoints with FDR < 0.05. b) Barplot showing gene set enrichment analysis of differentially spliced transcripts. c) UMAP colored by Louvain clusters based on calculated psi (percent of spliced in) values. d) psi-based UMAP colored by individual. e) psi-based UMAP colored by timepoint. f) Violin plots showing the single cell distribution of logit (percent spliced in) values at screening and C5D1 for selected differentially spliced transcripts. Boxplots show the median and interquartile range, whiskers extend to 1.5x the interquartile range. n = 1,374 cells. **** p ≤ 0.0001 and * p ≤ 0.05 by two-sided Wilcoxon test. g-j) Miso plots visualizing splice junctions and potential exon skipping events in differentially spliced transcripts for TSC1 (g), CD200 (h) and CALU (i) as well as SLAMF7 (j). Splice probabilities are shown on the right.
Extended Data Fig. 9 Transcriptional diversity following treatment.
a) Number of genes expressed in primary MM cells before and after treatment (p < 2.2e-16 by two-sided t-test). Boxplots show the median and interquartile range, whiskers extend to 1.5x the interquartile range. n = 1,374 cells. b) Bar graph showing relative proportion of cells in each cell cycle phase at screening and C5D1. c) Shown are the lineage TF regulons downregulated and upregulated upon treatment. d) Change in regulon activity of lineage TFs upon treatment. e,f) GO-term enrichment of ATAC-Seq peaks gained (e) and lost (f) in MOLP2 cells following 72 h treatment with PVD. g) Gene set enrichment analysis of the top 500 genes gaining enhancers following treatment with PVD. h) Genes with ≥ 5 enhancer interactions by ABC model. i) Gene set enrichment analysis of genes gaining enhancer interactions following treatment with PVD by ABC model. j) Top rewired TFs with differentially accessible motifs upon treatment.
Extended Data Fig. 10 Surface marker expression in MM.
a) Genes upregulated upon treatment. Shown is a barplot with log2 fold change values compared to screening timepoint. b) Motif enrichment in differential ATAC-Seq peaks in MMCL MOLP2 after PVD treatment (right), comparing to untreated (left). p values are calculated using binomial test. c-e) Live cell counts following 72 h of treatment with pomalidomide (Pom), bortezomib (Bor), dexamethasone (Dex) and combination of all three drugs (PVD) in MM cell lines MOLP2 (c), MM1.S (d) and OPM2 (e) as a percentage of total cell numbers. f,g) Quantification of CXCR4 surface levels by flow cytometry following treatment with pomalidomide (Pom), bortezomib (Bor), dexamethasone (Dex) and combination of all three drugs (PVD) in myeloma cell lines MOLP2 (f) and MM1.S (g). c-g) Significance was assessed using a two-sided t-test with Welch’s correction, with * p ≤ 0.05 and **** p ≤ 0.0001. Data are presented as mean + /- SD, n = 3 independent experimental replicates. h,i) Cell viability following treatment with CXCR4 inhibitors BKT140 (h) and Plerixafor (i) following pre-treatment with PVD in MMCL MOLP2. Data are presented as mean + /- SD, n = 3 independent experiments. n.s. p > 0.05, * p ≤ 0.05, *** p ≤ 0.001 and **** p ≤ 0.0001 by two-sided t-test.
Supplementary information
Supplementary Information
Enlarged version of heatmaps with dendrograms.
Supplementary Tables
Supplementary Tables 1–11.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Frede, J., Anand, P., Sotudeh, N. et al. Dynamic transcriptional reprogramming leads to immunotherapeutic vulnerabilities in myeloma. Nat Cell Biol 23, 1199–1211 (2021). https://doi.org/10.1038/s41556-021-00766-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-021-00766-y
This article is cited by
-
Murine skin-derived multipotent papillary dermal fibroblast progenitors show germline potential in vitro
Stem Cell Research & Therapy (2023)
-
Single-cell technologies in multiple myeloma: new insights into disease pathogenesis and translational implications
Biomarker Research (2023)
-
Dynamic single-cell RNA-seq analysis reveals distinct tumor program associated with microenvironmental remodeling and drug sensitivity in multiple myeloma
Cell & Bioscience (2023)
-
Cell-free DNA for the detection of emerging treatment failure in relapsed/ refractory multiple myeloma
Leukemia (2022)
-
Estimation of tumor cell total mRNA expression in 15 cancer types predicts disease progression
Nature Biotechnology (2022)