Abstract
Cells respond to stress by blocking translation, rewiring metabolism and forming transient messenger ribonucleoprotein assemblies called stress granules (SGs). After stress release, re-establishing homeostasis and disassembling SGs requires ATP-consuming processes. However, the molecular mechanisms whereby cells restore ATP production and disassemble SGs after stress remain poorly understood. Here we show that upon stress, the ATP-producing enzyme Cdc19 forms inactive amyloids, and that their rapid re-solubilization is essential to restore ATP production and disassemble SGs in glucose-containing media. Cdc19 re-solubilization is initiated by the glycolytic metabolite fructose-1,6-bisphosphate, which directly binds Cdc19 amyloids, allowing Hsp104 and Ssa2 chaperone recruitment and aggregate re-solubilization. Fructose-1,6-bisphosphate then promotes Cdc19 tetramerization, which boosts its activity to further enhance ATP production and SG disassembly. Together, these results describe a molecular mechanism that is critical for stress recovery and directly couples cellular metabolism with SG dynamics via the regulation of reversible Cdc19 amyloids.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Mass spectrometry data have been deposited to the ProteomeXchange Consortium via the PRIDE51 partner repository (dataset identifier PXD026060). Numerical source data giving rise to graphical representations (with all independent repeats) and unprocessed images of gels and blots are reported in the Source Data files. Detailed experimental procedures and additional data supporting the findings of this study are available from the corresponding author upon reasonable request. Source data are provided with this paper.
Change history
26 October 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41556-021-00799-3
References
Fulda, S. et al. Cellular stress responses: cell survival and cell death. Int. J. Cell Biol. 2010, 214074 (2010).
Aulas, A. et al. Stress-specific differences in assembly and composition of stress granules and related foci. J. Cell Sci. 130, 927–937 (2017).
Protter, D. S. W. & Parker, R. Principles and properties of stress granules. Trends Cell Biol. 26, 668–679 (2016).
Saad, S. et al. Reversible protein aggregation is a protective mechanism to ensure cell cycle restart after stress. Nat. Cell Biol. 19, 1202–1213 (2017).
Begovich, K. et al. Conserved metabolite regulation of stress granule assembly via AdoMet. J. Cell Biol. 219, e201904141 (2020).
Cabrera, M. et al. Chaperone-facilitated aggregation of thermo-sensitive proteins shields them from degradation during heat stress. Cell Rep. 30, 2430–2443 (2020).
Wolozin, B. & Ivanov, P. Stress granules and neurodegeneration. Nat. Rev. Neurosci. 20, 649–666 (2019).
Cao, X., Jin, X. & Liu, B. The involvement of stress granules in aging and aging-associated diseases. Aging Cell 19, e13136 (2020).
Jain, S. et al. ATPase-modulated stress granules contain a diverse proteome and substructure. Cell 164, 487–498 (2016).
Molliex, A. et al. Phase separation by low complexity domains promotes stress granule assembly and drives pathological fibrillization. Cell 163, 123–133 (2015).
Banani, S. F. et al. Biomolecular condensates: organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 18, 285–298 (2017).
Van Treeck, B. et al. RNA self-assembly contributes to stress granule formation and defining the stress granule transcriptome. Proc. Natl Acad. Sci. USA 115, 2734–2739 (2018).
Kroschwald, S. et al. Different material states of Pub1 condensates define distinct modes of stress adaptation and recovery. Cell Rep. 23, 3327–3339 (2018).
Sathyanarayanan, U. et al. ATP hydrolysis by yeast Hsp104 determines protein aggregate dissolution and size in vivo. Nat. Commun. 11, 5226 (2020).
Shattuck, J. E. et al. The prion-like protein kinase Sky1 is required for efficient stress granule disassembly. Nat. Commun. 10, 3614 (2019).
Wang, B. et al. ULK1 and ULK2 regulate stress granule disassembly through phosphorylation and activation of VCP/p97. Mol. Cell 74, 742–757 (2019).
Walters, R. W. et al. Differential effects of Ydj1 and Sis1 on Hsp70-mediated clearance of stress granules in Saccharomyces cerevisiae. RNA 21, 1660–1671 (2015).
Patel, A. et al. ATP as a biological hydrotrope. Science 356, 753–756 (2017).
Hofmann, S. et al. Translation suppression promotes stress granule formation and cell survival in response to cold shock. Mol. Biol. Cell 23, 3786–3800 (2012).
Weitzel, G., Pilatus, U. & Rensing, L. The cytoplasmic pH, ATP content and total protein synthesis rate during heat-shock protein inducing treatments in yeast. Exp. Cell. Res. 170, 64–79 (1987).
Soini, J. et al. Transient increase of ATP as a response to temperature up-shift in Escherichia coli. Micro. Cell Fact. 4, 9 (2005).
Xu, Y. F. et al. Regulation of yeast pyruvate kinase by ultrasensitive allostery independent of phosphorylation. Mol. Cell 48, 52–62 (2012).
Fiechter, A., Fuhrmann, G. F. & Käppeli, O. Regulation of glucose metabolism in growing yeast cells. Adv. Micro. Physiol. 22, 123–183 (1981).
Flores, C. L. et al. Carbohydrate and energy-yielding metabolism in non-conventional yeasts. FEMS Microbiol. Rev. 24, 507–529 (2000).
Grignaschi, E. et al. A hydrophobic low-complexity region regulates aggregation of the yeast pyruvate kinase Cdc19 into amyloid-like aggregates in vitro. J. Biol. Chem. 293, 11424–11432 (2018).
Anastasiou, D. et al. Pyruvate kinase M2 activators promote tetramer formation and suppress tumorigenesis. Nat. Chem. Biol. 8, 839–847 (2012).
Blázquez, M. A. et al. Trehalose-6-phosphate, a new regulator of yeast glycolysis that inhibits hexokinases. FEBS Lett. 329, 51–54 (1993).
van Vaeck, C. et al. Analysis and modification of trehalose 6-phosphate levels in the yeast Saccharomyces cerevisiae with the use of Bacillus subtilis phosphotrehalase. Biochem. J. 353, 157–162 (2001).
Peeters, K. et al. Fructose-1,6-bisphosphate couples glycolytic flux to activation of Ras. Nat. Commun. 8, 922 (2017).
Cereghetti, G. et al. Reversible, functional amyloids: towards an understanding of their regulation in yeast and humans. Cell Cycle 17, 1545–1558 (2018).
Villar-Piqué, A. et al. Screening for amyloid aggregation: in-silico, in-vitro and in-vivo detection. Curr. Protein Pept. Sci. 15, 477–489 (2014).
Linder, T. Evaluation of the chitin-binding dye Congo red as a selection agent for the isolation, classification, and enumeration of ascomycete yeasts. Arch. Microbiol. 200, 671–675 (2018).
Feng, Y. et al. Global analysis of protein structural changes in complex proteomes. Nat. Biotechnol. 32, 1036–1044 (2014).
Cappelletti, V. et al. Dynamic 3D proteomes reveal protein functional alterations at high resolution in situ. Cell 184, 545–559 (2021).
Boles, E. et al. Characterization of a glucose-repressed pyruvate kinase (Pyk2p) in Saccharomyces cerevisiae that is catalytically insensitive to fructose-1,6-bisphosphate. J. Bacteriol. 179, 2987–2993 (1997).
Orlandi, I. et al. Ethanol and acetate acting as carbon/energy sources negatively affect yeast chronological aging. Oxid. Med. Cell. Longev. 2013, 802870 (2013).
Bell, W. et al. Composition and functional analysis of the Saccharomyces cerevisiae trehalose synthase complex. J. Biol. Chem. 273, 33311–33319 (1998).
Monteiro, F. et al. Measuring glycolytic flux in single yeast cells with an orthogonal synthetic biosensor. Mol. Syst. Biol. 15, e9071 (2019).
Ji, F. et al. Determination of intracellular metabolites concentrations in Escherichia coli under nutrition stress using liquid chromatography-tandem mass spectrometry. Talanta 189, 1–7 (2018).
Persson, L.B., Ambati, V.S. & Brandman, O. Cellular control of viscosity counters changes in temperature and energy availability. Cell 183, 1572–1585 (2020).
Fenton, A. W. & Blair, J. B. Kinetic and allosteric consequences of mutations in the subunit and domain interfaces and the allosteric site of yeast pyruvate kinase. Arch. Biochem. Biophys. 397, 28–39 (2002).
Cherkasov, V. et al. Coordination of translational control and protein homeostasis during severe heat stress. Curr. Biol. 23, 2452–2462 (2013).
Mogk, A., Bukau, B. & Kampinga, H. H. Cellular handling of protein aggregates by disaggregation machines. Mol. Cell 69, 214–226 (2018).
Gutierres, M. B. B., Bonorino, C. B. C. & Rigo, M. M. ChaperISM: improved chaperone binding prediction using position-independent scoring matrices. Bioinformatics 36, 735–741 (2020).
Narayanaswamy, R. et al. Widespread reorganization of metabolic enzymes into reversible assemblies upon nutrient starvation. Proc. Natl Acad. Sci. USA 106, 10147–10152 (2009).
Plata, M. R. et al. Determination of carbohydrates present in Saccharomyces cerevisiae using mid-infrared spectroscopy and partial least squares regression. Anal. Bioanal. Chem. 405, 8241–8250 (2013).
Yuan, M. et al. A positive/negative ion-switching, targeted mass spectrometry-based metabolomics platform for bodily fluids, cells, and fresh and fixed tissue. Nat. Protoc. 7, 872–881 (2012).
Sonderegger, M. & Sauer, U. Evolutionary engineering of Saccharomyces cerevisiae for anaerobic growth on xylose. Appl. Environ. Microbiol. 69, 1990–1998 (2003).
Lamprecht, I., Schaarschmidt, B. & Welge, G. Microcalorimetric investigation of the metabolism of yeasts. V. Influence of ploidy on growth and metabolism. Radiat. Environ. Biophys. 13, 57–61 (1976).
Piazza, I. et al. A map of protein-metabolite interactions reveals principles of chemical communication. Cell 172, 358–372 (2018).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Acknowledgements
We thank C. Boone (University of Toronto) for providing strains for chaperone overexpression; K. Weis (ETH Zurich) for antibodies and yeast strains; C. Kraft (University of Freiburg) for help with yeast cell semi-permeabilization; S.-S. Lee, L. Garbani Marcantini, J. Schleicher, D. M. Szymala and A. Timofiiva for their help with microscopy and data analysis; ScopeM and FGCZ for their technical support; A. Smith for critical editing; and P. Arosio, D. Jarosz, A. Sengör and the members of the Peter laboratory for their helpful discussions and comments on the manuscript. This work was funded by the Synapsis Foundation, ETH Zurich, and the Swiss National Science Foundation (grant no. SNF 200426). In addition, P.P. received funding from the European Research Council (ERC-CoG) and the EPIC-XS Consortium.
Author information
Authors and Affiliations
Contributions
Conceptualization: G.C., R.D. and M.P. Formal analysis: G.C. and I.P. Investigation: G.C., C.W.-Z., V.M.K., M.D., A.A., I.P., H.Y. and S.S. Writing–original draft preparation: G.C. and M.P. Writing–review and editing: G.C. and M.P., with input from all authors. Visualization: G.C. and M.P. Supervision: M.P., P.P., U.S. and D.A.D. Funding acquisition: M.P., P.P., U.S. and D.A.D.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Cell Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cdc19 aggregates are amyloids both in vitro and in vivo.
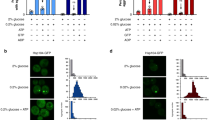
a, Schematic representation of the domains of Cdc19. The four mutated residues located within the LCR in the Cdc19irrev mutant are indicated (red circles). b,c, Both Cdc19WT and Cdc19irrev form ThT- and Congo Red (CR)-positive aggregates upon heat shock in vitro. Purified Cdc19WT, Cdc19irrev and a non-aggregating Cdc19 mutant as negative control (Cdc19ΔPEP4) were incubated with ThT (b) or CR (c) at 4 °C or after heat shock (42 °C, 10 min). Fluorescence emission was measured at 490 nm or 614 nm, respectively. The mean ± s.e.m. is shown (n = 3 independent experiments, two-tailed t-tests, ThT: PWT = 0.0000149, Pirrev = 0.0091, CR: PWT = 0.0003, Pirrev = 0.0128). d, In vivo-formed Cdc19WT and Cdc19irrev aggregates are CR-positive. Cells expressing GFP-tagged Cdc19WT or Cdc19irrev were harvested when exponentially growing or after heat shock (42 °C, 30 min) and lysed. Cdc19–GFP was immobilized in a GFP-trap microfluidic chamber and stained with CR. GFP and CR signals were detected by fluorescence microscopy, and merged to visualize co-localization. Arrowheads indicate CR-positive Cdc19–GFP aggregates (n = 3 independent experiments). Scale bars; 10 μm. e, Limited-Proteolysis Mass Spectrometry (LiP-MS) results indicate that Cdc19WT and Cdc19irrev undergo comparable structural transitions upon aggregation in vitro and in vivo. Purified soluble or aggregated (42 °C, 10 min) Cdc19WT or Cdc19irrev, as well as cell extracts obtained from cells expressing Cdc19WT-GFP or Cdc19irrev-GFP harvested during exponential growth or stationary phase (2 d) to induce aggregation were analysed by LiP-MS as described in the Methods (n = 3 independent experiments). Peptides detected in soluble and aggregated Cdc19WT and Cdc19irrev are displayed in volcano plots, and upregulated (red) or downregulated (blue) conformation-sensitive peptides are highlighted. Conformation-specific LiP-MS-peptides detected in vitro and in vivo were mapped to the Cdc19 schematic drawing (green). f, Intracellular ATP levels (mM) were determined in the indicated strains after heat shock (42 °C, 30 min) and recovery (30 °C, 60 min). The mean ± s.e.m. of n = 5 independent experiments is shown (two-tailed Mann-Whitney test, **P = 0.0079); a.u., arbitrary units. Source data for all graphical representations are found in Source Data Extended Data Fig. 1.
Extended Data Fig. 2 Genetic screening identifies trehalose metabolism as a regulator of reversible Cdc19 aggregation.
a, Screening protocol that identified a single point mutation (G386D) in TPS3 as a suppressor of stress-induced growth arrest of cdc19irrev cells. b,d, Serial dilutions of the indicated exponentially growing strains were spotted on agar plates before or after heat shock (42 °C, 30 min), and imaged after 3 d at 30 °C (n = 3 independent experiments). c, Exponentially growing tps3-G385D cells expressing Cdc19irrev-GFP and Pab1–CFP were cultivated at 30 °C, then heat shocked (42 °C, 30 min) and allowed to recover at 30 °C. The mean ± s.e.m. percentage of cells with Cdc19 aggregates is indicated (n = 3 independent experiments, >30 cells per sample per experiment). Scale bar; 5 μm. e, Intracellular disaccharides were measured in the indicated strains before, during and after heat shock (42 °C, 30 min). The mean ± s.e.m. is shown (n = 5 independent experiments for wild-type, n = 3 independent experiments for tps1Δ and tps2Δ, two-tailed Mann-Whitney test, **P = 0.0079). f,g, The indicated exponentially growing strains were heat shocked (42 °C, 30 min) and allowed to recover at 30 °C ± antimycin A (1 or 2 μM, respectively). f, Serial dilutions were spotted on agar plates ± antimycin A before or after heat shock, and imaged after 3 d at 30 °C (n = 3 independent experiments). g, Plots indicate mean percentage of cells that re-solubilized Cdc19 (n = 2 independent experiments). h, Mean Cdc19–GFP levels relative to Pgk1 in the indicated strains are shown (n = 2 independent experiments). i, Exponentially growing tps2Δ cells expressing Cdc19irrev-–GFP and Pab1–CFP were heat shocked (42 °C, 30 min) and allowed to recover at 30 °C. Plot indicates the mean ± s.e.m. percentage of cells with Cdc19 aggregates (n = 3 independent experiments, >30 cells per sample per experiment). Scale bar; 5 µm. j, Intracellular disaccharides (mainly trehalose46) and FBP were measured in the indicated strains before, during and after heat shock (42 °C, 30 min), and plotted as mean ± s.e.m. (n = 4 independent experiments for cdc19irrev tps2Δ, n = 5 for cdc19irrev, two-tailed Mann-Whitney test, *PDisaccharides = 0.0159, *PFBP = 0.0317); a.u., arbitrary units. Source data for all graphical representations and unprocessed western blots available in Source Data Extended Data Fig. 2.
Extended Data Fig. 3 FBP specifically reduces aggregation of purified Cdc19WT but not Cdc19ΔFBP and Cdc19irrev mutant proteins.
a–c, Purified wild-type Cdc19 (a), FBP binding-deficient Cdc19ΔFBP mutant (b) or Cdc19irrev mutant (c) proteins were mixed as indicated with 5 mM FBP or for control 5 mM F6P, 5 mM ATP, 5 mM PEP, 5 mM trehalose or buffer and incubated at 30 °C for 14 h. Cdc19 aggregates were separated from soluble protein by centrifugation and the supernatant (Sup) and pellet fractions were analysed by SDS-PAGE and Coomassie blue staining. The amount of Cdc19 was quantified in the pellet and supernatant by measuring Cdc19 band intensities using ImageJ and normalizing to buffer controls, and is displayed as mean ± s.e.m. (n = 3 independent experiments, two-tailed t-test, *P = 0.0342). d, Addition of FBP alone is not sufficient to re-solubilize pre-formed Cdc19WT amyloids in vitro. Purified Cdc19WT (Input) was incubated for 10 min at 42 °C to trigger its aggregation, and the resulting amyloids were incubated with (20 mM) or without (0 mM) FBP for several hours. Cdc19 re-solubilization was then tested by centrifugation and analysis of the resulting supernatant (Sup) and pellet fractions by SDS-PAGE and Coomassie blue staining (n = 3 independent experiments); a.u., arbitrary units. Unprocessed original scans of gels are shown in Source Data Extended Data Fig. 3.
Extended Data Fig. 4 The chaperones Hsp104 and Ssa2 cooperate with FBP to efficiently disassemble Cdc19 amyloids in vivo.
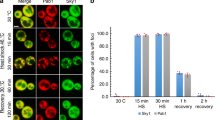
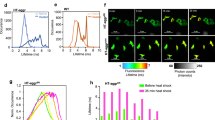
a, Co-overexpression of Hsp104 and Ssa2 partially restores growth of cdc19irrev cells after heat stress. Wild-type and cdc19irrev cells overexpressing as indicated Hsp104 or Ssa2, or both together by addition of 10 mM estradiol for 3 h, were heat-shocked for 30 min at 42 °C. Serial dilutions were spotted on agar plates and grown at 30 °C for 3 d (n = 3 independent experiments). b, Hsp104 protein levels were quantified in the indicated strains by western blotting either in the absence (-) or presence (+) of 10 mM estradiol for 3 h (n = 3 independent experiments). c, Increased FBP levels and Hsp104 cooperate to efficiently restart growth after stress in cdc19irrev cells. Exponentially growing wild-type or tps1Δ cells expressing either Cdc19WT or Cdc19irrev were subjected to a 30 min heat shock at 42 °C. Where indicated, overexpression of Hsp104 was induced by treating cells with 10 mM estradiol for 3 h. Growth restart after stress release of the indicated strains was quantified by measuring the cell density (OD600) over time after inoculation of equal cell numbers at 30 °C. Mean ± s.e.m. cell density 22 h after stress release is shown (n = 3 independent experiments). Note that Hsp104 overexpression and increased FBP levels cooperate to rescue cdc19irrev cells after heat shock. d, Wild-type or tps1Δ cells co-expressing mCherry-tagged Hsp104 and either GFP-tagged Cdc19WT or Cdc19irrev mutant were heat shocked (42 °C, 30 min), and imaged by fluorescence microscopy at the times indicated. Representative GFP- (left row) and mCherry images (middle row) are shown, together with the merged image (bottom row) to visualize co-localization of Cdc19 aggregates and Hsp104 (n = 3 independent experiments). Scale bars; 5 μm; a.u., arbitrary units. Source data for the graphical representation and unprocessed western blots can be found in Source Data Extended Data Fig. 4.
Supplementary information
Supplementary Table
Supplementary Table 1. Yeast strains used in this study. Supplementary Table 2. Plasmids used in this study
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed gels and western blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed western blot.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed gels.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed western blot.
Rights and permissions
About this article
Cite this article
Cereghetti, G., Wilson-Zbinden, C., Kissling, V.M. et al. Reversible amyloids of pyruvate kinase couple cell metabolism and stress granule disassembly. Nat Cell Biol 23, 1085–1094 (2021). https://doi.org/10.1038/s41556-021-00760-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-021-00760-4
This article is cited by
-
Activation of L-lactate oxidase by the formation of enzyme assemblies through liquid–liquid phase separation
Scientific Reports (2023)
-
Amyloid formation as a protein phase transition
Nature Reviews Physics (2023)
-
Mechanisms and pathology of protein misfolding and aggregation
Nature Reviews Molecular Cell Biology (2023)
-
Adaptive preservation of orphan ribosomal proteins in chaperone-dispersed condensates
Nature Cell Biology (2023)
-
A comparative meta-analysis of membraneless organelle-associated proteins with age related proteome of C.elegans
Cell Stress and Chaperones (2022)