Abstract
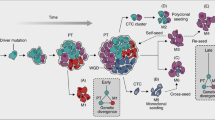
Metastasis is the leading cause of cancer-related deaths and enables cancer cells to compromise organ function by expanding in secondary sites. Since primary tumours and metastases often share the same constellation of driver mutations, the mechanisms that drive their distinct phenotypes are unclear. Here we show that inactivation of the frequently mutated tumour suppressor gene LKB1 (encoding liver kinase B1) has evolving effects throughout the progression of lung cancer, which leads to the differential epigenetic re-programming of early-stage primary tumours compared with late-stage metastases. By integrating genome-scale CRISPR–Cas9 screening with bulk and single-cell multi-omic analyses, we unexpectedly identify LKB1 as a master regulator of chromatin accessibility in lung adenocarcinoma primary tumours. Using an in vivo model of metastatic progression, we further show that loss of LKB1 activates the early endoderm transcription factor SOX17 in metastases and a metastatic-like sub-population of cancer cells within primary tumours. The expression of SOX17 is necessary and sufficient to drive a second wave of epigenetic changes in LKB1-deficient cells that enhances metastatic ability. Overall, our study demonstrates how the downstream effects of an individual driver mutation can change throughout cancer development, with implications for stage-specific therapeutic resistance mechanisms and the gene regulatory underpinnings of metastatic evolution.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
RNA-seq, scATAC–seq and ATAC–seq data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE167381. The human lung adenocarcinoma data were derived from the TCGA Research Network (http://cancergenome.nih.gov/). The dataset derived from this resource that supports the findings of this study is publicly available at https://gdc.cancer.gov/about-data/publications/ATACseq-AWG. All other data supporting the findings of this study are available from the corresponding authors on request. Transcription factor binding motifs were derived from CIS-BP (http://cisbp.ccbr.utoronto.ca/index.php). Source data are provided with this paper.
Code availability
All custom code used in this work is available from the corresponding authors upon request. We also host a Github website that includes the main analysis code used in this study (https://github.com/GreenleafLab/LKB1_2021)44.
References
Cancer Genome Atlas Research Network. Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014).
Waddell, N. et al. Whole genomes redefine the mutational landscape of pancreatic cancer. Nature 518, 495–501 (2015).
Sanchez-Cespedes, M. A role for LKB1 gene in human cancer beyond the Peutz-Jeghers syndrome. Oncogene 26, 7825–7832 (2007).
Ji, H. et al. LKB1 modulates lung cancer differentiation and metastasis. Nature 448, 807–810 (2007).
Carretero, J. et al. Integrative genomic and proteomic analyses identify targets for Lkb1-deficient metastatic lung tumors. Cancer Cell 17, 547–559 (2010).
Shackelford, D. B. & Shaw, R. J. The LKB1-AMPK pathway: metabolism and growth control in tumour suppression. Nat. Rev. Cancer 9, 563–575 (2009).
Jin, L. et al. The PLAG1-GDH1 axis promotes anoikis resistance and tumor metastasis through CamKK2-AMPK signaling in LKB1-deficient lung cancer. Mol. Cell 69, 87–99 (2018).
Calles, A. et al. Immunohistochemical loss of LKB1 is a biomarker for more aggressive biology in KRAS-mutant lung adenocarcinoma. Clin. Cancer Res. 21, 2851–2860 (2015).
Lizcano, J. M. et al. LKB1 is a master kinase that activates 13 kinases of the AMPK subfamily, including MARK/PAR-1. EMBO J. 23, 833–843 (2004).
Kottakis, F. et al. LKB1 loss links serine metabolism to DNA methylation and tumorigenesis. Nature 539, 390–395 (2016).
Hollstein, P. E. et al. The AMPK-related kinases SIK1 and SIK3 mediate key tumor-suppressive effects of LKB1 in NSCLC. Cancer Discov. 9, 1606–1627 (2019).
Murray, C. W. et al. An LKB1-SIK axis suppresses lung tumor growth and controls differentiation. Cancer Discov. 9, 1590–1605 (2019).
Pierce, S. E., Granja, J. M. & Greenleaf, W. J. High-throughput single-cell chromatin accessibility CRISPR screens enable unbiased identification of regulatory networks in cancer. Nat. Commun. 12, 2969 (2021).
Filbin, M. G. et al. Developmental and oncogenic programs in H3K27M gliomas dissected by single-cell RNA-seq. Science 360, 331–335 (2018).
Flavahan, W. A. et al. Altered chromosomal topology drives oncogenic programs in SDH-deficient GISTs. Nature 575, 229–233 (2019).
LaFave, L. M. et al. Epigenomic state transitions characterize tumor progression in mouse lung adenocarcinoma. Cancer Cell 38, 212–228 (2020).
Reiter, J. G. et al. Minimal functional driver gene heterogeneity among untreated metastases. Science 361, 1033–1037 (2018).
Hu, Z., Li, Z., Ma, Z. & Curtis, C. Multi-cancer analysis of clonality and the timing of systemic spread in paired primary tumors and metastases. Nat. Genet. 52, 701–708 (2020).
Turajlic, S. & Swanton, C. Metastasis as an evolutionary process. Science 352, 169–175 (2016).
Robles-Oteiza, C. et al. Recombinase-based conditional and reversible gene regulation via XTR alleles. Nat. Commun. 6, 8783 (2015).
Morgens, D. W. et al. Genome-scale measurement of off-target activity using Cas9 toxicity in high-throughput screens. Nat. Commun. 8, 15178 (2017).
Mi, H., Muruganujan, A., Ebert, D., Huang, X. & Thomas, P. D. PANTHER version 14: more genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Res. 47, D419–D426 (2019).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Corces, M. R. et al. Lineage-specific and single-cell chromatin accessibility charts human hematopoiesis and leukemia evolution. Nat. Genet. 48, 1193–1203 (2016).
Corces, M. R. et al. The chromatin accessibility landscape of primary human cancers. Science 362, eaav1898 (2018).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Kaufman, J. M. et al. A transcriptional signature identifies LKB1 functional status as a novel determinant of MEK sensitivity in lung adenocarcinoma. Cancer Res. 77, 153–163 (2017).
Winslow, M. M. et al. Suppression of lung adenocarcinoma progression by Nkx2-1. Nature 473, 101–104 (2011).
Park, K.-S., Wells, J. M., Zorn, A. M., Wert, S. E. & Whitsett, J. A. Sox17 influences the differentiation of respiratory epithelial cells. Dev. Biol. 294, 192–202 (2006).
Laughney, A. M. et al. Regenerative lineages and immune-mediated pruning in lung cancer metastasis. Nat. Med. 26, 259–269 (2020).
Satpathy, A. T. et al. Massively parallel single-cell chromatin landscapes of human immune cell development and intratumoral T cell exhaustion. Nat. Biotechnol. 37, 925–936 (2019).
Granja, J. M. et al. Single-cell multiomic analysis identifies regulatory programs in mixed-phenotype acute leukemia. Nat. Biotechnol. 37, 1458–1465 (2019).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Walkinshaw, D. R. et al. The tumor suppressor kinase LKB1 activates the downstream kinases SIK2 and SIK3 to stimulate nuclear export of class IIa histone deacetylases. J. Biol. Chem. 288, 9345–9362 (2013).
Parra, M. Class IIa HDACs - new insights into their functions in physiology and pathology. FEBS J. 282, 1736–1744 (2015).
Zhang, H. et al. Lkb1 inactivation drives lung cancer lineage switching governed by Polycomb Repressive Complex 2. Nat. Commun. 8, 14922 (2017).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Morgens, D. W., Deans, R. M., Li, A. & Bassik, M. C. Systematic comparison of CRISPR/Cas9 and RNAi screens for essential genes. Nat. Biotechnol. 34, 634–636 (2016).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Adamson, B. et al. A multiplexed single-cell CRISPR screening platform enables systematic dissection of the unfolded protein response. Cell 167, 1867–1882 (2016).
Chuang, C.-H. et al. Molecular definition of a metastatic lung cancer state reveals a targetable CD109-Janus kinase-Stat axis. Nat. Med. 23, 291–300 (2017).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Gen. 53, 403–411 (2021).
Granja, J. M. GreenleafLab/LKB1_2021: Release_1.0.1 Zenodo https://doi.org/10.5281/zenodo.5035694 (2021).
Acknowledgements
We thank J. Sage, A. Trevino and members of the Greenleaf and Winslow laboratories for comments. We thank the Stanford Shared FACS facility and the Veterinary Service Center for technical support. We thank A. Orantes for administrative support. S.E.P was supported by an NSF Graduate Research Fellowship Award and the Tobacco-Related Diseases Research Program Predoctoral Fellowship Award (grant number T31DT1900). This work was supported by National Institutes of Health (NIH) grant numbers R01-CA204620 and R01-CA230919 (to M.M.W.), RM1-HG007735 and UM1-HG009442 (to H.Y.C. and W.J.G.), R35-CA209919 (to H.Y.C.), UM1-HG009436 and U19-AI057266 (to W.J.G.), and in part by the Stanford Cancer Institute support grant (NIH grant P30-CA124435).
Author information
Authors and Affiliations
Contributions
S.E.P., J.M.G., M.M.W. and W.J.G. conceived the project and designed the experiments. S.E.P. led the experimental data production together with contributions from J.M.G., M.R.C., J.J.B., M.K.T., A.B.P., R.T. and P.C. S.E.P. and J.M.G. led the data analysis. S.E.P. performed the CRISPR screen analysis and RNA-seq analysis. J.M.G. and S.E.P. performed the ATAC–seq and scATAC–seq analysis. J.M.G. was supervised by H.Y.C and W.J.G. S.E.P. was supervised by M.C.B., W.J.G. and M.M.W. S.E.P., J.M.G., W.J.G. and M.M.W. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
W.J.G. and H.Y.C. are consultants for 10x Genomics, which has licensed IP associated with ATAC–seq. W.J.G. has additional affiliations with Guardant Health (consultant) and Protillion Biosciences (co-founder and consultant). M.M.W. is a co-founder of, and holds equity in, D2G Oncology, Inc. H.Y.C. is a co-founder of Accent Therapeutics, Boundless Bio, and a consultant for Arsenal Biosciences and Spring Discovery. The remaining authors declare no competing interests.
Additional information
Peer review Information Nature Cell Biology thanks Kwon-Sik Park, Tomi Makela and Skirmantas Kriaucionis for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Validation and quality control of inducible LKB1 restoration model and genome-scale CRISPR/Cas9 screen.
a. Schematic of restorable Lkb1TR/TR alleles. SA = splice acceptor, SD = splice donor, FRT = flippase recognition target. b. Schematic of the derivation of LKB1-restorable cell lines. c. Expression of LKB1 by immunoblot over a time-course of 4-OHT treatment, represented in hours (h) and days (d). HSP90 is a sample processing control. 25% and 10% of input after six days of 4-OHT treatment is shown for a visual comparison. d. Expression of LKB1 by immunoblot in LR1 and LR2 cells treated with vehicle or 4-OHT compared to a KPT cell line and a KPT;Lkb1−/− cell line. HSP90 is a sample processing control. e. RNA-sequencing reads mapping to the Lkb1 locus following six days of 4-OHT or vehicle treatment. f. Subcutaneous growth assay following injection of cell lines into recipient NSG mice. Tamoxifen or vehicle treatment was initiated on day 0. Mean tumor volume as measured by calipers of six tumors per condition +/− SEM is shown. g. Intravenous (i.v.) transplant assays. Left: Representative lung histology. Right: Change in % tumor area in LKB1-restored cells. Mean area of four mice per condition +/− SEM is shown. **p = 0.001, ***p = 0.0001, n.s. = not significant with a two-sided t-test. Scale bars represent 5 mm. h. Cumulative population doublings recorded over 12 days of 4-OHT treatment. Each cell line and condition was cultured and analyzed in triplicate. Mean +/− SEM is shown. **p = 0.0002 for LR1, **p = 0.0001 for LR2. i. Left: Representative image of clonogenic growth in LR1 cells. Right: % normalized area of cell growth. Each treatment group was cultured and analyzed in triplicate. Mean +/− SEM is shown. *p < 0.01, **p < 0.001, n.s. = not significant with a two-sided t-test. Scale bars represent 10 mm. p = 0.0001 for LR1 and p = 0.0059 for LR2. j. Heatmap of Pearson correlation matrix of log-normalized counts across all samples in the genome-scale CRISPR/Cas9 screen. k. Log2 fold enrichment of negative control sgRNAs and sgRNAs targeting Lkb1 at day 12 versus day 0.
Extended Data Fig. 2 LKB1 restoration drives widespread changes in chromatin accessibility in lung adenocarcinoma cells.
a. Schematic of preparing LKB1-deficient and LKB1-restored samples prior to ATAC-seq library preparation. Cell lines were treated with 4-OHT or vehicle for six days. b. Representative plot of aggregate signal around the transcription start site (TSS) for all ATAC-seq peaks in one vehicle-treated, LR1 replicate. This plot represents the signal-to-noise quantification of our ATAC-seq data. TSS enrichment scores greater than 10 indicate high quality ATAC-seq data. c. TSS enrichment scores for 16 ATAC-seq libraries with technical replicates. d. Differential accessibility across 178,783 ATAC-seq peaks following 4-OHT treatment in the LKB1-restorable (LR1 and LR2) and LKB1-unrestorable (LU1 and LU2) cell lines. The x-axis represents the log2 mean accessibility per peak and the y-axis represents the log2 fold change in accessibility following 4-OHT treatment. Colored points are significant (|log2 fold change | >0.5, FDR < 0.05). e. Percentage of differential peaks (|log2 fold change | >0.5, FDR < 0.05) across multiple ATAC-seq comparisons. f. Schematic of preparing samples for LKB1-restoration time-course. Cell lines were treated with 4-OHT for eight different time-points (0 hours, 6 hours, 12 hours, 24 hours, 36 hours, 48 hours, 4 days, and 6 days) in two cell lines (LR1 and LR2). g. and h. PCA (g) and k-means clustering (h) of 9,480 correlated, variable ATAC-seq peaks across the LKB1 restoration time-course in two cell lines (LR1 and LR2). Each row of the heatmap represents a z-score of log2 normalized accessibility across all samples within each cell line. i and j. SOX (i) and TEAD (j) motif accessibility changes (∆chromVAR deviation scores) across time in two cell lines (LR1 and LR2) treated with 4-OHT for the indicated time-points. Shaded area represents 95th percent confidence interval.
Extended Data Fig. 3 Inactivating chromatin modifiers only delays LKB1-induced chromatin changes.
a. Schematic of generating single knock-out populations of chromatin modifiers identified in the CRISPR screen, treating with 4-OHT or vehicle for six days, and processing for ATAC-seq. b. Principle component analysis (PCA) of the top 10,000 variable ATAC-seq peaks across the indicated LR1;Cas9 knock-out populations treated with 4-OHT or vehicle. c. K-means clustered heatmap of differential peak accessibility (log2 fold change) for each genotype of LR1;Cas9 cells treated with 4-OHT for up to 48 hours compared to 0 hours. All peaks differential between sgSafe (0 hours 4-OHT) and sgSafe (48 hours 4-OHT) are shown. Each row represents the log2 fold change of each genotype and time-point versus the same genotype’s initial time-point (day 0). d. Log2 fold change in mean peak accessibility for all peaks in k-means cluster 3 (top) and cluster 4 (bottom) from (c) for the indicated genotype and 4-OHT time-points compared to 0 hours 4-OHT. N = 2 technical replicates per sgRNA population and time-point. Box-whisker plot; lower whisker is the lowest value greater than the 25% quantile minus 1.5 times the interquartile range (IQR), the lower hinge is the 25% quantile, the middle is the median, the upper hinge is the 75% quantile and the upper whisker is the largest value less than the 75% quantile plus 1.5 times the IQR.
Extended Data Fig. 4 SIK family members mediate LKB1-induced chromatin changes.
a. Schematic of generating single and multiple sgRNA knock-out cell lines and processing for ATAC-seq. LR1;Cas9 cells were treated with 4-OHT or vehicle for six days. b. Left: Heatmap of peak accessibility between each knock-out population treated with 4-OHT or vehicle. Each row represents a z-score of log2 normalized accessibility across all samples. Right: Transcription factor hypergeometric motif enrichment in each k-means cluster. c. Percent of differential ATAC-seq peaks (|log2 fold change | >0.5, FDR < 0.05) across LKB1-restorable cells treated with 4-OHT or vehicle. d. SOX (top) and FOXA (bottom) motif accessibility changes (∆chromVAR deviation scores normalized to vehicle-treated sgSafe) across LKB1-restorable knock-out populations treated with 4-OHT or vehicle. e. Heatmap of Pearson correlation matrix of log2-normalized accessibility (in counts per million (CPM)) across LKB1 downstream effector knock-out genotypes with and without LKB1 restoration in LR1;Cas9 cells. f. PCA of the top 10,000 variable ATAC-seq peaks across LR1;Cas9 knock-out populations treated with 4-OHT or vehicle. Principle components besides PC1 (70.6%) account for <4% of the variance in the dataset. N = 2 technical replicates per sgRNA population. g. SOX (top) and FOXA (bottom) motif accessibility changes (∆chromVAR deviation scores normalized to vehicle-treated sgSafe) across LKB1-restorable knock-out populations treated with 4-OHT or vehicle. Line represents average between two technical replicates. h. Heatmap of Pearson correlation matrix of log2-normalized accessibility (in counts per million (CPM)) across LKB1 downstream effector knock-out genotypes with and without LKB1 restoration in LR1;Cas9 cells. i and j. PCA of the top 10,000 variable ATAC-seq peaks across LR1;Cas9 knock-out populations treated with 4-OHT or vehicle. Principle components besides PC1 account for <4% of the variance in the dataset. N = 2 technical replicates per sgRNA population.
Extended Data Fig. 5 Loss of LKB1 partitions human lung adenocarcinoma primary tumors into two chromatin accessibility sub-types.
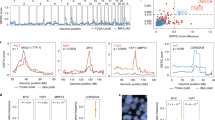
a. Enrichment of mutations in Chromatin Type 2 tumors compared to Chromatin Type 1 tumors. Genes are ranked according to -log10(FDR), with Rank 1 (LKB1) being the most significant (see Methods), as indicated on the y-axis. Points are colored by the number of mutations in the TCGA-LUAD ATAC-seq dataset (out of 21 samples). b. ChromVAR deviation scores for the indicated transcription factor motifs for samples in the TCGA-LUAD ATAC-seq dataset. *p < 0.1, **p < 0.005, ****p < 10−6 using a two-sided t-test. p = 1×10−7 for RUNX, p = 0.002 for FOXA, and p = 1×10−7. N = 13 biologically independent samples for Chromatin Type 1 and 8 biologically independent samples for Chromatin Type 2. Box-whisker plot; lower whisker is the lowest value greater than the 25% quantile minus 1.5 times the interquartile range (IQR), the lower hinge is the 25% quantile, the middle is the median, the upper hinge is the 75% quantile and the upper whisker is the largest value less than the 75% quantile plus 1.5 times the IQR.
Extended Data Fig. 6 Loss of LKB1 drives a unique chromatin accessibility state in human lung adenocarcinoma cell lines.
a. Hierarchical clustering of human lung cancer cell lines using the Euclidian distance within the first three principle components from Fig. 2d. b. ChromVAR deviation scores for the indicated transcription factor motifs in eight human lung cancer cell lines at baseline. *p < 0.1, **p < 0.005, ****p < 10-6 using a two-sided t-test. p = 0.066 for FOXA, p = 0.003 for SOX, p = 3.1 ×10−7 for NR4A, and p = 0.001 for RUNX. N = 4 biologically independent samples for each group. Box-whisker plot; lower whisker is the lowest value greater than the 25% quantile minus 1.5 times the interquartile range (IQR), the lower hinge is the 25% quantile, the middle is the median, the upper hinge is the 75% quantile and the upper whisker is the largest value less than the 75% quantile plus 1.5 times the IQR. c. Comparison of the changes in motif accessibility (∆ chromVAR deviation scores) across LKB1-wild-type and LKB1-mutant human lung cancer cell lines (y-axis) and Chromatin Type 1 and Type 2 tumors (x-axis). Dark grey or colored points are called significantly different (q < 0.05) across both comparisons. Light grey points are not significant. A selection of motif families and their associated motif logos are indicated. d. Differential accessibility across ATAC-seq peaks following LKB1 wild-type expression in eight human lung cancer cell lines. The x-axis represents the log2 fold change in accessibility following LKB1 restoration. LKB1-mutant and LKB1-wild-type status at baseline is indicated. Colored points are significant (|log2 fold change| >0.5, FDR < 0.05). e. LKB1-deficiency score by RNA-seq (using 16-gene signature from Kaufmann et al., 2017) compared to log10(number of differential ATAC-seq peaks + 1) following LKB1 expression in each indicated cell line. Pearson correlation indicated in top left. Shaded area represents 95th percent confidence interval. f and g. Relative chromVAR deviation scores for SOX (f) an NR4A (g) motifs in the indicated cell lines transduced with GFP, LKB1, or KEAP1. Scores are normalized based on the GFP control for each cell line. N = 2 technical replicates per cell line and overexpression condition. h. Percent of differential ATAC-seq peaks (|log2 fold change | >0.5, FDR < 0.05) in cells transduced to express KEAP1 compared to GFP.
Extended Data Fig. 7 Genotype-specific activation of SOX17 in LKB1-deficient metastatic cells.
a. Percent survival of KPT and KPT;Lkb1−/− mice compared to KT mice. b and c. Number of primary tumors observed in KPT and KPT;Lkb1−/− mice (b). Lung weights of KPT and KPT;Lkb1−/− mice (c). N = 7 biologically independent mice for each genotype. Box-whisker plot; lower whisker is the lowest value greater than the 25% quantile minus 1.5 times the interquartile range (IQR), the lower hinge is the 25% quantile, the middle is the median, the upper hinge is the 75% quantile and the upper whisker is the largest value less than the 75% quantile plus 1.5 times the IQR. n.s. = non-significant with a two-sided t-test. d. Metastatic rates of KPT (2/7 mice with metastases) and KPT;Lkb1−/− (7/7 mice with metastases). p-value = 0.00016 with a one-sided binomial test. e and f. Comparison of the changes in motif accessibility (∆chromVAR deviation scores) between murine LKB1-proficient (KPT) and LKB1-deficient (KPT;Lkb1−/−) metastases (y-axis) and between murine LKB1-restored and LKB1-deficient cells (x-axis; e) or Chromatin Type 1 tumors and Chromatin Type 2 tumors (x-axis; f). Dark grey or colored points are called significantly different (q < 0.05) across both comparisons. Light grey points are not significant. A selection of motif families and their associated motif logos are indicated. g. log2 fold change in mRNA expression (left) and accessibility within the gene body (right) of each NKX2 transcription factor compared to the average expression and accessibility in primary tumor samples. Asterisks indicate transcription factors with greater than log2fold change of −1 in both RNA and ATAC measurements. h. log2 fold change in mRNA expression (left) and accessibility within the gene body (right) of each SOX transcription factor compared to the average expression and accessibility in primary tumor samples. Asterisks indicate transcription factors with greater than log2fold change of 2 in both RNA and ATAC measurements.
Extended Data Fig. 8 LKB1-deficient primary tumors harbor sub-populations of SOX17 + cells.
a. Representative immunohistochemistry (IHC) against SOX17 and grading of SOX17 expression for LKB1-proficient KPT and LKB1-deficient KPT;Lkb1−/− samples. Images are annotated according to percent area of the tumor composed of SOX17 + cells. Negative (0%), low (<25%), medium (25–50%), and high (>50%). Scale bars represent 50uM. Images are representative of 117 KPT primary tumors, 203 KPT;Lkb1−/− primary tumors, 14 KPT metastases, and 8 KPT;Lkb1−/− metastases, as quantified in (b). b. Quantitation of SOX17 protein expression in LKB1-proficient KPT and LKB1-deficient KPT;Lkb1−/− primary tumors and metastases, graded according to (a). The number of samples analyzed for histology for each genotype and tumor type is indicated at the top. Overall 0% of LKB1-proficient primary tumors or metastases had SOX17 + cells, 31% of LKB1-deficient primary tumors had SOX17 + cells, and 100% of LKB1-deficient metastases had SOX17 + cells. c. Correlation of SOX17 mRNA expression (y-axis) and LKB1 mRNA expression (x-axis) in ten human lung adenocarcinoma samples that contain Type 1 metastatic cell clusters (H0 and H3; Laughney et al. 2020). Each point indicates the mean value of SOX17 or LKB1 expression for each sample +/− SEM for all single cells evaluated by scRNA-seq. Shaded area represents 95th percent confidence interval. d. SOX17 genome accessibility track of the average ATAC-seq signal from Chromatin Type 1 and Chromatin Type 2 tumors.
Extended Data Fig. 9 A subset of LKB1-deficient primary tumors harbor metastatic-like, SOX17 + sub-populations.
a. scATAC-seq quality control metrics. TSS enrichment (left, middle), insertion profiles (right), and number of fragments per cell (right inset) in seven primary tumors. N = 998 cells for 10 C, 3556 cells for 13B, 1467 cells for 13 A, 3373 cells for 15 A, 1310 cells for 15B, 2858 cells for 17 A, and 851 cells for 21 A. Box-whisker plot; lower whisker is the lowest value greater than the 25% quantile minus 1.5 times the interquartile range (IQR), the lower hinge is the 25% quantile, the middle is the median, the upper hinge is the 75% quantile and the upper whisker is the largest value less than the 75% quantile plus 1.5 times the IQR. b. UMAP of cells from seven primary tumors. c. Percent of cells from each cluster in each primary tumor. d. Comparison of the changes in motif accessibility (∆chromVAR deviation scores) between LKB1-deficient metastases and primary tumors (y-axis) versus the average difference between cluster 12 cells and cells in clusters 1–11 (x-axis). Dark grey or colored points are called significantly different (q < 0.05) across both comparisons. Light grey points are not significant. e and f. Average accessibility of peaks in each scATAC-seq cluster that are enriched in LKB1-deficient primary tumors compared to LKB1-deficient metastases (e) or enriched in LKB1-deficient metastases compared to LKB1-deficient primary tumors (f) and are overlapping with the scATAC-seq peakset. Error bars indicate +/− SEM. N = 2993 cells (Cluster 1), N = 1011 cells (Cluster 2), N = 508 cells (Cluster 3), N = 856 cells (Cluster 4), N = 408 cells (Cluster 5), N = 3435 cells (Cluster 6), N = 468 cells (Cluster 7), N = 1517 cells (Cluster 8), N = 1733 cells (Cluster 9), N = 119 cells (Cluster 11), N = 116 cells (Cluster 12). Box-whisker plot; lower whisker is the lowest value greater than the 25% quantile minus 1.5 times the interquartile range (IQR), the lower hinge is the 25% quantile, the middle is the median, the upper hinge is the 75% quantile and the upper whisker is the largest value less than the 75% quantile plus 1.5 times the IQR.
Extended Data Fig. 10 SOX17 regulates chromatin accessibility state and growth in metastatic, LKB1-deficient cells.
a. Sox17 genome accessibility track (left) and mean mRNA expression (right) following 4-OHT or vehicle. Significantly differential ATAC-seq peaks in grey (log2 fold change < −0.5, FDR < 0.05). Sox17 also has significantly decreased mRNA expression (log2 fold change < −1, FDR < 0.05). b. GREAT GO term enrichment of genes nearby the differential peaks that contain SOX binding motifs. c. Sox17 genome accessibility track of an LKB1-restorable cell line (LR1;Cas9) transduced with the indicated sgRNAs and treated with 4-OHT or vehicle. d. Relative Sox17 mRNA expression in LR1;Cas9 cells transduced with sgSafe or sgSik1-3 and treated with either vehicle or 4-OHT. Mean values +/− SEM. N = 3 biologically independent samples examined over 2 experiments. e. Expression of SOX17 and/or LKB1 by immunoblot in LR2;Cas9 cells transduced with non-targeting (sgNT#1 and sgNT#2) or Sox17-targeting sgRNAs (sgSox17#1 and sgSox17#2) (top) or LR2;Cas9 cells transduced with BFP-overexpressing (control) or Sox17-overexpressing constructs and treated with vehicle or 4-OHT. HSP90 is a sample processing control. f. Heatmap of relative log2fold changes of the indicated genotypes of LR2;Cas9 cells. The top 10,000 consistent, variable ATAC-seq peaks following LKB1 restoration in both sgSafe and BFP transduced cells are shown. Clusters 3 and 4 from the Sox17 knock-out experiment are shown independently for emphasis in Fig. 5d. g and h. Lung weight following injection of LR2;Cas9 cells treated with vehicle or 4-OHT after Sox17 knock-out (g) or Sox17 overexpression (h). *p < 0.05, **p < 0.005, ***p < 0.0005 with a two-sided t-test. N = 3 biologically independent mice evaluated for LKB1-deficient (sgSafe) and 4 biologically independent mice for all other conditions. p = 0.0481 for sgSafe vs. sgSox17-1 (LKB1-deficient), p = 0.0184 for sgSafe vs. sgSox17-2 (LKB1-deficient), p = 0.0008 for BFP-vehicle vs. BFP-4-OHT, and p = 0.001 for BFP-4-OHT vs. Sox17-4-OHT. i. Schematic of intrasplenic (i.s.) injections into immunocompromised NSG mice. j. Representative fluorescent tdTomato+ images of the left lateral lobe of the liver. Scale bars represent 5 mm. k. Log10 (number of liver metastases) following intrasplenic injection of cells. Condition +/− SEM is shown. p = 0.055 with a two-sided t-test. N = 9 mice BFP, N = 8 mice for Sox17.
Supplementary information
Supplementary Information
Information on how to perform more detailed ATAC–seq analyses.
Supplementary Table 1
Enriched sgRNAs and gene targets in LKB1-restored cells compared with LKB1-deficient cells from a genome-scale CRISPR–Cas9 screen. P values are calculated from a negative binomial model using the MAGeCK algorithm.
Supplementary Table 2
List of all ATAC–seq and scATAC–seq samples processed in this study with related quality control information.
Supplementary Table 3
Gene expression changes in LKB1-restorable and LKB1-unrestorable cell lines after treatment with 4-OHT or vehicle.
Supplementary Table 4
sgRNA spacer sequences used in this study.
Supplementary Table 5
List of all KPT and KPT;Lkb1−/− mouse samples processed for ATAC–seq in this study.
Supplementary Table 6
Gene expression of LKB1-proficient KPT and LKB1-deficient KPT;Lkb1−/− mouse primary tumours and metastases.
Source data
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Source Data Extended Data Fig. 10.
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Pierce, S.E., Granja, J.M., Corces, M.R. et al. LKB1 inactivation modulates chromatin accessibility to drive metastatic progression. Nat Cell Biol 23, 915–924 (2021). https://doi.org/10.1038/s41556-021-00728-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-021-00728-4
This article is cited by
-
Roflumilast inhibits tumor growth and migration in STK11/LKB1 deficient pancreatic cancer
Cell Death Discovery (2024)
-
Oncogenic enhancers prime quiescent metastatic cells to escape NK immune surveillance by eliciting transcriptional memory
Nature Communications (2024)
-
Oxidative stress-triggered Wnt signaling perturbation characterizes the tipping point of lung adeno-to-squamous transdifferentiation
Signal Transduction and Targeted Therapy (2023)
-
Dissecting metastasis using preclinical models and methods
Nature Reviews Cancer (2023)
-
Cancer cell plasticity during tumor progression, metastasis and response to therapy
Nature Cancer (2023)