Abstract
Pioneer transcription factors such as OCT4 can target silent genes embedded in nucleosome-dense regions. How nucleosome interaction enables transcription factors to target chromatin and determine cell identity remains elusive. Here, we systematically dissect OCT4 to show that nucleosome binding is encoded within the DNA-binding domain and yet can be uncoupled from free-DNA binding. Furthermore, accelerating the binding kinetics of OCT4 to DNA enhances nucleosome binding. In cells, uncoupling nucleosome binding diminishes the ability of OCT4 to individually access closed chromatin, while more dynamic nucleosome binding results in expansive genome scanning within closed chromatin. However, both uncoupling and enhancing nucleosome binding are detrimental to inducing pluripotency from differentiated cells. Remarkably, stable interactions between OCT4 and nucleosomes are continuously required for maintaining the accessibility of pluripotency enhancers in stem cells. Our findings reveal how the affinity and residence time of OCT4–nucleosome complexes modulate chromatin accessibility during cell fate changes and maintenance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The cryo-EM map of the LIN28B nucleosome core particle has been deposited in Electron Microscopy Data Bank with database code EMD-12453. The atomic coordinates of the LIN28B NCP model have been deposited in Protein Data Bank with PDB ID: 7NL0. The human nucleosome core particle (PDB ID: 6IPU)51 and the OCT4-DBD–DNA crystal structures (PDB ID: 3L1P)52 were used for model building. All next-generation sequencing data generated as part of this study have been deposited in the Gene Expression Omnibus (GEO) under one super-series with the accession number GSE168142, which is composed of the subseries GSE167632 and GSE168141. Previously published ATAC-seq and MNase-seq data that were re-analysed here are available under accession codes GSE98124 (ref. 35) and GSM1004653 (ref. 13), respectively. Data were aligned to the Mus musculus genome assembly MGSCv37 (mm9) with accession PRJNA20689. Source data are provided with this paper. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
References
Iwafuchi-Doi, M. & Zaret, K. S. Pioneer transcription factors in cell reprogramming. Genes Dev. 28, 2679–2692 (2014).
King, H. W. & Klose, R. J. The pioneer factor OCT4 requires the chromatin remodeller BRG1 to support gene regulatory element function in mouse embryonic stem cells. eLife 6, e22631 (2017).
Gaspar-Maia, A., Alajem, A., E, Meshorer. & M, Ramalho-Santos. Open chromatin in pluripotency and reprogramming. Nat. Rev. Mol. Cell Biol. 12, 36–47 (2011).
Chen, X. et al. Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell 133, 1106–1117 (2008).
Soufi, A., Donahue, G. & Zaret, K. S. Facilitators and impediments of the pluripotency reprogramming factors’ initial engagement with the genome. Cell 151, 994–1004 (2012).
Knaupp, A. S. et al. Transient and permanent reconfiguration of chromatin and transcription factor occupancy drive reprogramming. Cell Stem Cell 21, 834–845.e6 (2017).
Li, D. et al. Chromatin accessibility dynamics during iPSC reprogramming. Cell Stem Cell 21, 819–833.e6 (2017).
Zviran, A. et al. Deterministic somatic cell reprogramming involves continuous transcriptional changes governed by Myc and epigenetic-driven modules. Cell Stem Cell 24, 328–341.e9 (2019).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Chronis, C. et al. Cooperative binding of transcription factors orchestrates reprogramming. Cell 168, 442–459.e20 (2017).
Chen, J. et al. Hierarchical Oct4 binding in concert with primed epigenetic rearrangements during somatic cell reprogramming. Cell Rep. 14, 1540–1554 (2016).
Soufi, A. et al. Pioneer transcription factors target partial DNA motifs on nucleosomes to initiate reprogramming. Cell 161, 555–568 (2015).
Teif, V. B. et al. Genome-wide nucleosome positioning during embryonic stem cell development. Nat. Struct. Mol. Biol. 19, 1185–1192 (2012).
Cirillo, L. A. et al. Opening of compacted chromatin by early developmental transcription factors HNF3 (FoxA) and GATA-4. Mol. Cell 9, 279–289 (2002).
Cirillo, L. A. et al. Binding of the winged-helix transcription factor HNF3 to a linker histone site on the nucleosome. EMBO J. 17, 244–254 (1998).
Cirillo, L. A. & Zaret, K. S. An early developmental transcription factor complex that is more stable on nucleosome core particles than on free DNA. Mol. Cell 4, 961–969 (1999).
Iwafuchi-Doi, M. et al. The pioneer transcription factor FoxA maintains an accessible nucleosome configuration at enhancers for tissue-specific gene activation. Mol. Cell 62, 79–91 (2016).
Zhu, F. et al. The interaction landscape between transcription factors and the nucleosome. Nature 562, 76–81 (2018).
Fernandez Garcia, M. et al. Structural features of transcription factors associating with nucleosome binding. Mol. Cell 75, 921–932.e6 (2019).
Iwafuchi, M. et al. Gene network transitions in embryos depend upon interactions between a pioneer transcription factor and core histones. Nat. Genet. 52, 418–427 (2020).
Michael, A. K. et al. Mechanisms of OCT4–SOX2 motif readout on nucleosomes. Science 368, 1460–1465 (2020).
Tapia, N. et al. Dissecting the role of distinct OCT4–SOX2 heterodimer configurations in pluripotency. Sci. Rep. 5, 13533 (2015).
Nichols, J. et al. Formation of pluripotent stem cells in the mammalian embryo depends on the POU transcription factor Oct4. Cell 95, 379–391 (1998).
Ambrosetti, D. C., Basilico, C. & Dailey, L. Synergistic activation of the fibroblast growth factor 4 enhancer by Sox2 and Oct-3 depends on protein–protein interactions facilitated by a specific spatial arrangement of factor binding sites. Mol. Cell. Biol. 17, 6321–6329 (1997).
Avilion, A. A. et al. Multipotent cell lineages in early mouse development depend on SOX2 function. Genes Dev. 17, 126–140 (2003).
Remenyi, A. et al. Crystal structure of a POU/HMG/DNA ternary complex suggests differential assembly of Oct4 and Sox2 on two enhancers. Genes Dev. 17, 2048–2059 (2003).
Boyer, L. A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005).
Zaret, K. S. & Carroll, J. S. Pioneer transcription factors: establishing competence for gene expression. Genes Dev. 25, 2227–2241 (2011).
Kong, X. et al. Functional interplay between the RK motif and linker segment dictates Oct4–DNA recognition. Nucleic Acids Res. 43, 4381–4392 (2015).
Echigoya, K. et al. Nucleosome binding by the pioneer transcription factor OCT4. Sci. Rep. 10, 11832 (2020).
Jerabek, S. et al. Changing POU dimerization preferences converts Oct6 into a pluripotency inducer. EMBO Rep. 18, 319–333 (2017).
Huertas, J., MacCarthy, C. M., Schöler, H. R. & Cojocaru, V. Nucleosomal DNA dynamics mediate Oct4 pioneer factor binding. Biophysical J. 118, 2280–2296 (2020).
Zhou, K., Gaullier, G. & Luger, K. Nucleosome structure and dynamics are coming of age. Nat. Struct. Mol. Biol. 26, 3–13 (2019).
von Hippel, P. Protein–DNA recognition: new perspectives and underlying themes. Science 263, 769–770 (1994).
Benchetrit, H. et al. Direct induction of the three pre-implantation blastocyst cell types from fibroblasts. Cell Stem Cell 24, 983–994.e7 (2019).
Niwa, H., Masui, S., Chambers, I., Smith, A. G. & Miyazaki, J. Phenotypic complementation establishes requirements for specific POU domain and generic transactivation function of Oct-3/4 in embryonic stem cells. Mol. Cell. Biol. 22, 1526–1536 (2002).
Niwa, H., Miyazaki, J. & Smith, A. G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nat. Genet. 24, 372–376 (2000).
Smith, A. G. et al. Inhibition of pluripotential embryonic stem cell differentiation by purified polypeptides. Nature 336, 688–690 (1988).
Blomquist, P., Li, Q. & Wrange, Ö. The affinity of nuclear factor 1 for Its DNA site is drastically reduced by nucleosome organization irrespective of its rotational or translational position. J. Biol. Chem. 271, 153–159 (1996).
Taylor, I. C., Workman, J. L., Schuetz, T. J. & Kingston, R. E. Facilitated binding of GAL4 and heat shock factor to nucleosomal templates: differential function of DNA-binding domains. Genes Dev. 5, 1285–1298 (1991).
Mueller, B. et al. Widespread changes in nucleosome accessibility without changes in nucleosome occupancy during a rapid transcriptional induction. Genes Dev. 31, 451–462 (2017).
Mieczkowski, J. et al. MNase titration reveals differences between nucleosome occupancy and chromatin accessibility. Nat. Commun. 7, 11485 (2016).
Voong, L. N. et al. Insights into nucleosome organization in mouse embryonic stem cells through chemical mapping. Cell 167, 1555–1570.e15 (2016).
Iurlaro, M. et al. Mammalian SWI/SNF continuously restores local accessibility to chromatin. Nat. Genet. 53, 279–287 (2021).
Schick, S. et al. Acute BAF perturbation causes immediate changes in chromatin accessibility. Nat. Genet. 53, 269–278 (2021).
Hockemeyer, D. et al. A drug-inducible system for direct reprogramming of human somatic cells to pluripotency. Cell Stem Cell 3, 346–353 (2008).
Kim, J. et al. An iPSC line from human pancreatic ductal adenocarcinoma undergoes early to invasive stages of pancreatic cancer progression. Cell Rep. 3, 2088–2099 (2013).
Dyer, P. N. et al. Reconstitution of nucleosome core particles from recombinant histones and DNA. Methods Enzymol. 375, 23–44 (2003).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Sharma, D. et al. PARP1 exhibits enhanced association and catalytic efficiency with γH2A.X–nucleosome. Nat. Commun. 10, 5751 (2019).
Esch, D. et al. A unique Oct4 interface is crucial for reprogramming to pluripotency. Nat. Cell Biol. 15, 295–301 (2013).
Acknowledgements
We thank D. O’Carroll, I. Chambers, W. Bickmore and K. Zaret for comments on the manuscript and insightful discussions. We thank S. Rosser, the Edinburgh Genome Foundry and the UK Centre for Mammalian Synthetic Biology for support with generating the synthetic DNA constructs. We thank Edinburgh Genomics for the sequencing service. We thank S. Thillainadarasan for assistance with making the OCT4 alanine-stretch mutant constructs. We thank M. Wilson for kindly providing the tailless H3, H4 and the acidic patch histone mutants and help with nucleosome reconstitution. We thank N. Lukoyanova for assistance with cryo-EM data collection. The work was supported by a MRC career development award (MR/N024028/1) to A.S. that supported A.S., B.O. and E.H.-P. The UK Research Councils’ Synthetic Biology for Growth programme and of the BBSRC, EPSRC and MRC (BB/M0180404/1) supported the work of G.A.R. The early detection committee CRUK-OHSU Project Award (C65925/A26986) to A.S. supported G.A.R. A Ph.D. scholarship from the Darwin Trust of Edinburgh supported M.R.O. The BBSRC-EASTBIO Ph.D. training programme (BB/M010996/1) supported I.G. A MRC career development award (MR/R008795/1) to P.J.R. supported the work of P.J.R. and M.S. Cryo-EM data for this investigation were collected at Birkbeck College, University of London, with financial support from the Wellcome Trust (202679/Z/16/Z and 206166/Z/17/Z).
Author information
Authors and Affiliations
Contributions
A.S. conceived the study and designed the experiments. G.A.R. carried out recombinant protein purification, EMSAs and nucleosome reconstitution with support from I.G. B.O. carried out virus production and iPSC reprogramming. G.A.R. and B.O carried out ChIP-seq and ATAC-seq with support from M.R.O. E.H.-P. carried out molecular cloning, MEF generation and pluripotency rescue with help from B.O. and G.A.R. A.S., G.A.R. and B.O. carried out bioinformatics analysis with support from M.R.O. I.G. generated nucleosomes for cryo-EM. M.S. and P.J.R. performed cryo-EM data collection, LIN28B nucleosome reconstruction, model building and refinement. A.S. prepared the manuscript with contribution from G.A.R., B.O., E.H.-P., M.R.O. and P.J.R.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Cell Biology thanks Alvaro Rada-Iglesias, Wolf Reik and the other, anonymous, reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Mutagenesis strategy to define the sub-domains of OCT4.
a, Schematic representation of OCT4 protein, indicating OCT4-DBD in white boxes and the linker region in black zigzag line. The α-helices (α1–8) defining the protein secondary structure are indicated by blue spirals. NT and CT regions are indicated by black lines. The deletion mutations spanning OCT4 domains are indicated above the curly brackets. Selected 5X alanine-stretch substitution are indicated by black boxes and numbered according to the equivalent deletion counterparts. b, The conservation plot (top), and the amino-acid consensus sequence (bottom) from multiple sequence alignment of OCT4 orthologs across mammals. c, Multiple sequence alignment of the 119 OCT4 derivatives, highlighting the deleted five a.a. regions as white boxes. The a.a. sequence is colour-coded following the colour scheme of the consensus sequence in (b). d, The generation of 118 out of 119 recombinant OCT4 deletion derivatives (del-1 to del-119) from bacteria as analyzed by SDS-PAGE and Coomassie staining. The respective OCT4 bands run at the expected size when compared to the sizes of the protein standards (M). The concentration of each OCT4 deletion derivative was measured from the intensity of the respective Coomassie-stained band relative to that of OCT4-WT run on the same gel. Apart from OCT4-WT, del-73, and del-79, all OCT4 recombinants were generated from one purification experiment. e, SDS-PAGE analysis showing Coomassie blue stained bands representing the recombinant OCT4-DBD with similar purity and concentration to the full-length counterpart. At least three biological replicates were used. f, The apparent dissociation constant (Kd) of OCT4-WT and mutants were determined from the binding curves in (Fig. 1d). ND indicates that Kd values were not determined due to non-saturated binding. Errors represent to the goodness of the fit to the experimental data.
Extended Data Fig. 2 The affinity of OCT4 deletion mutants for naked DNA versus nucleosomes.
a–d, EMSA gels showing the affinity of each recombinant OCT4 deletion mutant (118 in total) for Cy5-lablelled LIN28B-DNA (left half, black arrowheads) and LIN28B-nucleosome (right half, white arrowheads). No protein (-) and wildtype recombinant OCT4 (WT) were used as controls for each EMSA. The binding was measured at equilibrium using 2 nM of LIN28B-DNA or LIN28B-nucleosome incubated with 3 nM of each OCT4 deletion derivative. The number of OCT4 deletions are indicated above each EMSA. c, Asterisks indicate del-73 and del-75mutants showing enhanced affinity for DNA and nucleosomes. Dagger indicates del-79 mutant that can bind DNA but not nucleosomes. Double dagger indicates del-82–83 and del-86–87 OCT4 mutants with partial nucleosome uncoupling as the interaction of these mutants with DNA is also diminished. Apart from OCT4-WT, del-73, del-75 and del-79, which were repeated at least three times, the EMSA screen was carried out once.
Extended Data Fig. 3 OCT4 alanine-stretch substitution and deletion mutants display similar affinity for DNA and nucleosomes.
a, SDS-PAGE analysis showing Coomassie blue stained bands representing the selected OCT4 alanine stretch mutants shown in (Extended Data Fig. 1a). The fractions from the lysates (lys.), purified under denaturing conditions (pur.), and refolded (ref.) are shown for each mutant. The respective OCT4 bands (arrow) run at the expected size when compared to the sizes of the protein standards (M). b, c, EMSA showing the affinity of increasing amounts of recombinant deletion (top panels) and the equivalent alanine substitution (bottom panels) OCT4 mutants for Cy5-lablelled LIN28B-DNA (b, black arrowheads) and LIN28B-nucleosome (c, white arrowheads). OCT4 concentrations (nM) are indicated above each lane. d, e, same as (a) for OCT4 alanine point-mutants within (a.a. 236-240) and BRN2HD, BRN3HD hybrids (d), and delta 1-4 and ala-79 OCT4 mutants (e). The concentration of each OCT4 mutant was measured from the intensity of the respective Coomassie-stained band relative to that of OCT4-WT run on the same gel. a–e are representative images of at least three biological replicates.
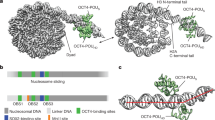
Extended Data Fig. 4 OCT4 configurations compatible with binding to the LIN28B-nucleosome.
a, Table summarizing cryo-EM imaging data collection and analysis to resolve the 3D structure of the LIN28B-nucleosome. b and d, Surface-filled representations of OCT4-DBD (green) bound to DNA (cartoon representation in grey) in the canonical (b) and MORE configuration (d). c, Surface-filled model of OCT4-DBD (orange) bound in the canonical configuration to the LIN28B-nucleosome at site 1 (pink), indicating incompatible binding due to steric clashes. e–g, Surface-filled models of OCT4-DBD in the MORE configuration bound to LIN28B-nucleosome at site 1 (e, magenta), site 2 (g, blue) and site 3 (f, green) to demonstrate that sites 1 and 3 are compatible (green ticks) with OCT4 nucleosome binding. Whereas OCT4 is prevented (red cross) from binding the LIN28B-nucleosome at site 2 due to steric clashes.
Extended Data Fig. 5 OCT4 nucleosome binding enables specific access to closed chromatin.
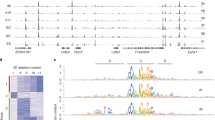
a, Western blot analysis showing antibody specificity and protein levels of OCT4-WT, del-73 and del-79 derivatives expressed in MEFs after lentiviral infection and Dox induction for 48 hr. GAPDH was used as a loading control. Protein sizes are indicated. Non-infected MEFs were used as a negative control. Data representative of 3 biological replicates. b, Agarose gel electrophoresis showing equivalent chromatin fragmentation used for ChIP-seq of OCT4-WT and mutants. Data representative of 3 biological replicates. c, Bar plots showing the distribution of OCT4 WT and mutants peaks relative to TSS as measured by GREAT analysis. d–f, The central enrichment of de novo motifs identified by DREME and TOMTOM analysis as measured by CentriMO analysis within the OCT4-WT closed sites (d), OCT4-del-73 open and closed sites (e), and the OCT4-del-79 open and closed sites (f). No motifs were identified within OCT4-WT open sites. Each motif enrichment is color coded (solid line) and controlled against background sequences (dotted line) that are 1 kb away from each OCT4 peak. A significant central enrichment is measured by CentriMO (p value) as indicated. Motif logos are indicated on top of each plot. Statistical analyses are provided in source data for Extended Data Fig. 5.
Extended Data Fig. 6 Pioneer function of OCT4 is essential for inducing pluripotency.
a, Whole-well images showing DAPI fluorescence (blue), Nanog immuno-fluorescence (red) and merged (magenta) on day 16 (4 days after Dox withdrawal) of reprogramming MEFs to iPSCs using OCT4 WT and mutants. MEFs infected with SKM (no OCT4) or OSKM in the absence of Dox (-Dox) were used as negative controls. 5 mm scale bars. Images are representative of 6 biological replicates. b, Bar plots showing the efficiency of reprogramming to pluripotency as quantified by the number of Nanog positive colonies (purple bars) versus DAPI-only colonies (Nanog -ve, blue bars) for OCT4-WT and mutants, each in combination with SKM. SKM + Dox and OSKM -/ + Dox were used as controls (bars 1-3). Averages of n = 6 independent biological replicates are shown (error bars indicate ± s.d.). Data points are represented in squares (Nanog +ve) and circles (Nanog -ve). c, Schematic showing the experimental workflow of reprogramming MEFs to iPSCs under increasing durations of ectopic OSKM expression using Dox. d, Representative whole-well images of 3 biological replicates showing DAPI fluorescence (blue), Nanog immuno-fluorescence (red) and merged (magenta) of MEFs ectopically expressing OCT4-WT, BRN2HD or BRN3HD hybrid mutants along with SKM for 6, 7, 8, 9, and 10 days. Dox-independent iPSC colonies were counted 4 days after Dox withdrawal. Representative bright field images of Dox-independent colonies (10 days induction) are shown in the left-hand panels. 5 mm scale bars. e, Line plots showing the reprogramming efficiency of OCT4-WT (circles), BRN2HD (squares) and BRN3HD (triangles) hybrid mutants during the different stages explained in (c and d). Averages of n = 3 independent biological replicates are shown (error bars indicate ± s.d.).
Extended Data Fig. 7 SOX2 can drive the pioneer deficient OCT4 to closed chromatin.
a, The distribution of de novo motifs identified by DRME and TOMTOM analysis as measured by CentriMO analysis within closed sites targeted by OCT4-WT, OCT4-del-73, and OCT4-del-79. The significance of central enrichment is measured by CentriMO (p value) as indicated. Statistical analysis are provided in source data Extended Data Fig. 7. b, Read-density heatmaps showing the ChIP-seq enrichment of OCT4 WT, del-73, and del-79 mutants when expressed alone in MEFs (blue) and MEFs ATAC-seq signal (red) spanning ± 1 kb from the centre of OCT4 unique peaks identified in early reprogramming (OSKM-48h). The analyzed sequences are rank-ordered from high to low according to the ATAC-seq reads within the central 300 bp of each peak. The ATAC-enriched (open) sequences are separated from the ATAC-depleted (closed) sequences and the percentage of each category is indicated. The total number of peaks (n) for each OCT4 derivative are shown to the right. The colour scales are indicated below. Input DNA is also shown. Line plots to show the average ChIP-seq or ATAC-seq enrichment are above each corresponding heatmap. c, Read-density heatmaps showing the ChIP-seq enrichment of SOX2, OCT4 WT, del-73, and del-79 mutants in early reprogramming (blue) and MEFs ATAC-seq signal (red) spanning ± 1 kb from the centre of SOX2 unique sites identified in OSKM-48h. The analyzed sequences are rank-ordered from high to low according to the ATAC-seq reads within the central 300 bp of each peak. The ATAC-enriched (open) sequences are separated from the ATAC-depleted (closed) sequences and the percentage of each category is indicated. The total number of peaks (n) and the colour scales are indicated. Input DNA is also shown. Line plots to show the average ChIP-seq or ATAC-seq enrichment are above each corresponding heatmap.
Extended Data Fig. 8 The conserved pioneer activity of OCT4 is crucial for maintaining pluripotency.
a, Bar plots showing the rescued AP positive ESC colonies in the presence of Dox and OCT4-WT or a del-79-derived mutant. Averages of n = 6 independent biological replicates are shown (error bars indicate ± s.d.). b, Whole well images showing AP positive ZHBTc4.1 ESCs in the absence of Dox (-Dox) or the presence of Dox (+Dox) either ectopically expressing OCT4 WT or one of the POUHD hybrids. Images are representative of 6 biological replicates. c, Bar plots showing the number of AP positive colonies in the presence of Dox for OCT4 WT, BRN2HD and BRN3HD as in (b). POUHD hybrids failed to rescue pluripotency compared to OCT4 WT (compare WT bars to BRN2HD and BRN3HD). Two-tailed t-test was used to assess statistical significance. **** indicates P < 0.0001. Averages of n = 6 independent biological replicates are shown (error bars indicate ± s.d.) as represented in (b). d, Western blot analysis showing the diminished levels of Oct4 after treating ZHBTc4.1 ESCs, ectopically expressing GFP, with Dox for 24 h and 48 h. GAPDH was used as loading control. Images are representative of 2 biological replicates. e, Representative Western blots from 2 biological replicates showing ZHBTc4.1 ESCs, ectopically expressing OCT4-WT or mutants after 24 h of Dox treatment. f, GO analysis of genes targeted by OCT4 at sites that closed down (red bars), remained open (blue bars) or remained closed (green bars) after treating ZHBTc4.1 ESCs, expressing ectopic OCT4-WT or mutants, with Dox for 24 h.
Supplementary information
Supplementary Information
Supplementary Methods: description of EMSA data analysis, cryo-EM and structural building and bioinformatics analysis.
Supplementary Tables
Supplementary Table 1: DNA sequences of OCT4 deletion mutant library; Supplementary Table 2: DNA sequences of additional OCT4 mutants; Supplementary Table 3: DNA oligonucleotide used for cloning.
Source data
Source Data Fig. 1
Unprocessed EMSA gels.
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Unprocessed EMSA gels.
Source Data Fig. 3
Unprocessed EMSA gels.
Source Data Fig. 4
Unprocessed EMSA gels.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Unprocessed whole-well images.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed EMSA gels.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Unprocessed whole-well images and western blots.
Source Data Extended Data Fig. 8
Statistical source data.
Rights and permissions
About this article
Cite this article
Roberts, G.A., Ozkan, B., Gachulincová, I. et al. Dissecting OCT4 defines the role of nucleosome binding in pluripotency. Nat Cell Biol 23, 834–845 (2021). https://doi.org/10.1038/s41556-021-00727-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-021-00727-5
This article is cited by
-
Structural mechanism of synergistic targeting of the CX3CR1 nucleosome by PU.1 and C/EBPα
Nature Structural & Molecular Biology (2024)
-
A pioneer factor locally opens compacted chromatin to enable targeted ATP-dependent nucleosome remodeling
Nature Structural & Molecular Biology (2023)
-
Histone modifications regulate pioneer transcription factor cooperativity
Nature (2023)
-
B1 SINE-binding ZFP266 impedes mouse iPSC generation through suppression of chromatin opening mediated by reprogramming factors
Nature Communications (2023)
-
Pioneer factors — key regulators of chromatin and gene expression
Nature Reviews Genetics (2023)