Abstract
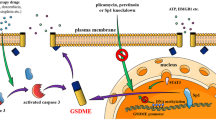
Although pyroptosis is critical for macrophages against pathogen infection, its role and mechanism in cancer cells remains unclear. PD-L1 has been detected in the nucleus, with unknown function. Here we show that PD-L1 switches TNFα-induced apoptosis to pyroptosis in cancer cells, resulting in tumour necrosis. Under hypoxia, p-Stat3 physically interacts with PD-L1 and facilitates its nuclear translocation, enhancing the transcription of the gasdermin C (GSDMC) gene. GSDMC is specifically cleaved by caspase-8 with TNFα treatment, generating a GSDMC N-terminal domain that forms pores on the cell membrane and induces pyroptosis. Nuclear PD-L1, caspase-8 and GSDMC are required for macrophage-derived TNFα-induced tumour necrosis in vivo. Moreover, high expression of GSDMC correlates with poor survival. Antibiotic chemotherapy drugs induce pyroptosis in breast cancer. These findings identify a non-immune checkpoint function of PD-L1 and provide an unexpected concept that GSDMC/caspase-8 mediates a non-canonical pyroptosis pathway in cancer cells, causing tumour necrosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The consensus caspase-8 recognition motifs were analysed using the CaspDB database (http://caspdb.sanfordburnham.org/). The p-Y705-Stat3-binding site in the GSDMC promoter was analysed using the GPMiner program (http://gpminer.mbc.nctu.edu.tw/). All other data supporting the findings of this study are available from the corresponding author on reasonable request. Source data are provided with this paper.
Change history
08 October 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41556-020-00599-1
References
Shi, J. et al. Cleavage of GSDMD by inflammatory caspases determines pyroptotic cell death. Nature 526, 660–665 (2015).
Kayagaki, N. et al. Caspase-11 cleaves gasdermin D for non-canonical inflammasome signalling. Nature 526, 666–671 (2015).
Liu, X. et al. Inflammasome-activated gasdermin D causes pyroptosis by forming membrane pores. Nature 535, 153–158 (2016).
Shi, J. et al. Inflammatory caspases are innate immune receptors for intracellular LPS. Nature 514, 187–192 (2014).
Ding, J. et al. Pore-forming activity and structural autoinhibition of the gasdermin family. Nature 535, 111–116 (2016).
Wang, Y. et al. Chemotherapy drugs induce pyroptosis through caspase-3 cleavage of a gasdermin. Nature 547, 99–103 (2017).
Rogers, C. et al. Cleavage of DFNA5 by caspase-3 during apoptosis mediates progression to secondary necrotic/pyroptotic cell death. Nat. Commun. 8, 14128 (2017).
Wang, Q. et al. A bioorthogonal system reveals antitumour immune function of pyroptosis. Nature 579, 421–426 (2020).
Zhang, Z. et al. Gasdermin E suppresses tumour growth by activating anti-tumour immunity. Nature 579, 415–420 (2020).
Zhou, Z. et al. Granzyme A from cytotoxic lymphocytes cleaves GSDMB to trigger pyroptosis in target cells. Science 368, eaaz7548 (2020).
Kovacs, S. B. & Miao, E. A. Gasdermins: effectors of pyroptosis. Trends Cell Biol. 27, 673–684 (2017).
Ribas, A. & Wolchok, J. D. Cancer immunotherapy using checkpoint blockade. Science 359, 1350–1355 (2018).
Powles, T. et al. MPDL3280A (anti-PD-L1) treatment leads to clinical activity in metastatic bladder cancer. Nature 515, 558–562 (2014).
Schmid, P. et al. Atezolizumab and nab-paclitaxel in advanced triple-negative breast cancer. N. Engl. J. Med. 379, 2108–2121 (2018).
Socinski, M. A. et al. Atezolizumab for first-line treatment of metastatic nonsquamous NSCLC. N. Engl. J. Med. 378, 2288–2301 (2018).
Satelli, A. et al. Potential role of nuclear PD-L1 expression in cell-surface vimentin positive circulating tumor cells as a prognostic marker in cancer patients. Sci. Rep. 6, 28910 (2016).
Ghebeh, H. et al. Doxorubicin downregulates cell surface B7-H1 expression and upregulates its nuclear expression in breast cancer cells: role of B7-H1 as an anti-apoptotic molecule. Breast Cancer Res. 12, R48 (2010).
Shimizu, S. et al. Induction of apoptosis as well as necrosis by hypoxia and predominant prevention of apoptosis by Bcl-2 and Bcl-XL. Cancer Res. 56, 2161–2166 (1996).
Grivennikov, S. I., Greten, F. R. & Karin, M. Immunity, inflammation, and cancer. Cell 140, 883–899 (2010).
Scaffidi, P., Misteli, T. & Bianchi, M. E. Release of chromatin protein HMGB1 by necrotic cells triggers inflammation. Nature 418, 191–195 (2002).
Vakkila, J. & Lotze, M. T. Inflammation and necrosis promote tumour growth. Nat. Rev. Immunol. 4, 641–648 (2004).
Carswell, E. et al. An endotoxin-induced serum factor that causes necrosis of tumors. Proc. Natl Acad. Sci. USA 72, 3666–3670 (1975).
Noman, M. Z. et al. PD-L1 is a novel direct target of HIF-1α, and its blockade under hypoxia enhanced MDSC-mediated T cell activation. J. Exp. Med. 211, 781–790 (2014).
Pawlus, M. R., Wang, L. & Hu, C. J. STAT3 and HIF1α cooperatively activate HIF1 target genes in MDA-MB-231 and RCC4 cells. Oncogene 33, 1670–1679 (2014).
Yang, Y. et al. Exosomal PD-L1 harbors active defense function to suppress T cell killing of breast cancer cells and promote tumor growth. Cell Res. 28, 862–864 (2018).
Li, C.-W. et al. Glycosylation and stabilization of programmed death ligand-1 suppresses T-cell activity. Nat. Commun. 7, 12632 (2016).
Timmer, J. & Salvesen, G. Caspase substrates. Cell Death Differ. 14, 66–72 (2007).
Poręba, M., Stróżyk, A., Salvesen, G. S. & Drąg, M. Caspase substrates and inhibitors. Cold Spring Harb. Perspect. Biol. 5, a008680 (2013).
Salvesen, G. S. & Ashkenazi, A. Snapshot: caspases. Cell 147, 476–476.e1 (2011).
Brown, J. M. & Wilson, W. R. Exploiting tumour hypoxia in cancer treatment. Nat. Rev. Cancer 4, 437–447 (2004).
Murdoch, C., Muthana, M., Coffelt, S. B. & Lewis, C. E. The role of myeloid cells in the promotion of tumour angiogenesis. Nat. Rev. Cancer 8, 618–631 (2008).
Barkal, A. A. et al. Engagement of MHC class I by the inhibitory receptor LILRB1 suppresses macrophages and is a target of cancer immunotherapy. Nat. Immunol. 19, 76–84 (2018).
Karagiannis, G. S. et al. Neoadjuvant chemotherapy induces breast cancer metastasis through a TMEM-mediated mechanism. Sci. Transl. Med. 9, eaan0026 (2017).
Aglietti, R. A. & Dueber, E. C. Recent insights into the molecular mechanisms underlying pyroptosis and gasdermin family functions. Trends Immunol. 38, 261–271 (2017).
Zhang, M. et al. Programmed death-ligand 1 triggers PASMCs pyroptosis and pulmonary vascular fibrosis in pulmonary hypertension. J. Mol. Cell. Cardiol. 138, 23–33 (2020).
Ehrnhoefer, D. E. et al. p53 increases caspase-6 expression and activation in muscle tissue expressing mutant huntingtin. Hum. Mol. Genet. 23, 717–729 (2013).
Curiel, T. J. et al. Specific recruitment of regulatory T cells in ovarian carcinoma fosters immune privilege and predicts reduced survival. Nat. Med. 10, 942–949 (2004).
Lin, H. et al. Host expression of PD-L1 determines efficacy of PD-L1 pathway blockade-mediated tumor regression. J. Clin. Investig. 128, 805–815 (2018).
Tang, H. et al. PD-L1 on host cells is essential for PD-L1 blockade-mediated tumor regression. J. Clin. Investig. 128, 580–588 (2018).
Kumar, V. et al. CD45 phosphatase inhibits STAT3 transcription factor activity in myeloid cells and promotes tumor-associated macrophage differentiation. Immunity 44, 303–315 (2016).
Edwards, J. G. et al. Tumor necrosis correlates with angiogenesis and is a predictor of poor prognosis in malignant mesothelioma. Chest 124, 1916–1923 (2003).
Koukourakis, M. I. et al. Lactate dehydrogenase-5 (LDH-5) overexpression in non-small-cell lung cancer tissues is linked to tumour hypoxia, angiogenic factor production and poor prognosis. Br. J. Cancer 89, 877–885 (2003).
Obeid, M. et al. Calreticulin exposure dictates the immunogenicity of cancer cell death. Nat. Med. 13, 54–61 (2007).
Krysko, D. V. et al. Immunogenic cell death and DAMPs in cancer therapy. Nat. Rev. Cancer 12, 860–875 (2012).
Galluzzi, L., Buque, A., Kepp, O., Zitvogel, L. & Kroemer, G. Immunological effects of conventional chemotherapy and targeted anticancer agents. Cancer Cell 28, 690–714 (2015).
Erkes, D. A. et al. Mutant BRAF and MEK inhibitors regulate the tumor immune microenvironment via pyroptosis. Cancer Discov. 10, 254–269 (2020).
Kosugi, S. et al. Six classes of nuclear localization signals specific to different binding grooves of importin α. J. Biol. Chem. 284, 478–485 (2009).
Bassi, C. et al. Nuclear PTEN controls DNA repair and sensitivity to genotoxic stress. Science 341, 395–399 (2013).
Kosugi, S., Hasebe, M., Tomita, M. & Yanagawa, H. Systematic identification of cell cycle-dependent yeast nucleocytoplasmic shuttling proteins by prediction of composite motifs. Proc. Natl Acad. Sci. USA 106, 10171–10176 (2009).
Pichler, A., Gast, A., Seeler, J. S., Dejean, A. & Melchior, F. The nucleoporin RanBP2 has SUMO1 E3 ligase activity. Cell 108, 109–120 (2002).
Lo, H. W. et al. Nuclear interaction of EGFR and STAT3 in the activation of the iNOS/NO pathway. Cancer Cell 7, 575–589 (2005).
Lin, S.-Y. et al. Nuclear localization of EGF receptor and its potential new role as a transcription factor. Nat. Cell Biol. 3, 802–808 (2001).
Shen, J. et al. EGFR modulates microRNA maturation in response to hypoxia through phosphorylation of AGO2. Nature 497, 383–387 (2013).
Chang, C. J. et al. EZH2 promotes expansion of breast tumor initiating cells through activation of RAF1-β-catenin signaling. Cancer Cell 19, 86–100 (2011).
Cha, T.-L. et al. Akt-mediated phosphorylation of EZH2 suppresses methylation of lysine 27 in histone H3. Science 310, 306–310 (2005).
Acknowledgements
We thank K. Dunner Jr. and acknowledge the National Institutes of Health Cancer Center (Support grant no. P30CA016672) for supporting the High Resolution Electron Microscopy Facility at The University of Texas MD Anderson Cancer Center for the electron microscopy analysis. This work was funded in part by the National Institutes of Health (grant nos P30CA016672 and R01 AI116722 (to M.-C.H.)); a National Breast Cancer Foundation Breast Cancer Research Foundation grant to M.-C.H.; the Patel Memorial Breast Cancer Endowment Fund; the Sister Institution Network Fund for collaboration between The University of Texas MD Anderson Cancer Center and China Medical University (to M.-C.H.); the YingTsai Young Scholar Award (grant no. CMU108-YTY02 to J.-M.H.); China Medical Univesity Start Up Fund (grant no. CMU108-N-02 to W.-J.W.); the NIH (grant nos R35 CA220430 and PO1 CA092584); the Cancer Prevention and Research Institute of Texas (grant no. RP180813); Robert A. Welch Chemistry Chair (to J.A.T.); Cancer Prevention and Research Institute of Texas (grant no. RP160710 to M.-C.H.); and Drug Development Center, Ministry of Education, Taiwan (grant no. 108-2311-B-241-001 to M.-C.H. and Y.C.).
Author information
Authors and Affiliations
Contributions
J.H. and M.-C.H. designed and conceived the study. J.H. and M.-C.H. wrote the manuscript. J.L.H. and K.H. contributed to the preparation of the manuscript. J.H., R.Z., W.X., Y. You, J.-M.H., Y.C., Y.-C.W., C.L., W.-J.W., B.K., Z.Y., Y. Yang, X.X., Y.L., C.-W.L., B.S. and J.A.T. performed experiments and analysed data. L.N. and C.-W.C. analysed the PD-L1 and GSDMC sequences. Y.W. provided patient tissue samples. M.-C.H. supervised the entire project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Establishment of stable cells in the MDA-MB-231 cell line.
(a) Deletion of endogenous Stat3 by CRISPR–Cas9. KO, knockout. Blots are representative of three independent experiments. (b) Stable re-expression of Stat3-WT or Stat3-Y705F in Stat3-knockout MDA-MB-231 cells. Validation of p-Y705-Stat3 expression level by immunoblotting. Blots are representative of three independent experiments. (c) Analysis of the nuclear localization signal (NLS) and nuclear export signal (NES) in the human PD-L1 amino acid sequence compared with that of other species. Sequence alignment of PD-L1 from human, mouse, cow, rat, and dog. The alignment was generated and presented by the ClustalW2 algorithm and ESPript 3.0 (http://espript.ibcp.fr/ESPript/cgi-bin/ESPript.cgi). Identical residues are indicated by the dark red background and conserved residues are in red. The NLS and NES sequences of PD-L1 are conserved across species. (d-f) Stable re-expression of PD-L1-WT (wild-type PD-L1), PD-L1-NLS (NLS-mutated PD-L1) or PD-L1-NES (NES-mutated PD-L1) in the PD-L1–knockout MDA-MB-231 cell line. Immunoblotting of total PD-L1 expression level in stable transfectants (d). Blots are representative of three independent experiments. Representative images of PD-L1 distribution of three independent experiments was determined by confocal assay under hypoxia (e-f). Scale bar, 20 μm. Unprocessed blots are provided in Source Data Extended Data Fig. 1.
Extended Data Fig. 2 nPD-L1-mediated pyroptotic cell death in cell lines of multiple cancer types.
(a) Immunoblotting analysis of PD-L1 expression levels in liver cancer and lung cancer cell lines. Blots are representative of three independent experiments. (b) Hypoxia-induced pyroptotic cell death in PD-L1-positive cells of multiple cancer types treated with p-Stat3 inhibitor, HO-3867. Red arrows indicate the pyroptotic cells with big bubbles. The experiment was repeated three times with similar results. Scale bar, 20 μm. Unprocessed blots are provided in Source Data Extended Data Fig. 2.
Extended Data Fig. 3 nPD-L1 and p-Y705-Stat3 are required for hypoxia-induced GSDMC expression.
(a) Real-time quantitative polymerase chain reaction (RT–qPCR) analysis of mRNA levels of gasdermin family members in the indicated MDA-MB-231 stable transfectants. Data are shown as mean ± SD of n = 3 independent experiments. (b and c) GSDMC mRNA (b) and protein (c) levels in MDA-MB-231 cells under hypoxia. Data of (b) are shown as mean ± SD of n = 3 independent experiments. Blots are representative of three independent experiments. N, normoxia; H, hypoxia. (d) RT–qPCR analysis of mRNA levels of GSDMC in the indicated MDA-MB-231 stable transfectants. Data are shown as mean ± SD of n = 3 independent experiments. (e-g) MDA-MB-231 cells were cultured under hypoxia and treated with HO-3867 (20 μM) or ivermectin (25 μM). GSDMC mRNA (e) and protein (f) levels were analysed by RT–qPCR and immunoblotting, respectively. GSDMC promoter luciferase reporter plasmids were transfected to cells, and luciferase activity was measured in (g). Data of (e) and (g) are shown as mean ± SD of n = 3 independent experiments. Blots are representative of three independent experiments. P values of all statistical analysis were determined by two-sided Student’s t-test. Statistical source data and unprocessed blots are provided in Source Data Extended Data Fig. 3.
Extended Data Fig. 4 The p-Y705-Stat3 binding site (in red) in the nucleic acid sequence of GSDMC promoter and the expression level of GSDME in MDA-MB-231 cells.
(a) GSDMC promoter sequence was obtained from GeneCopoeia and analysed using the GPMiner program (http://gpminer.mbc.nctu.edu.tw/). Two putative p-Y705-Stat3-binding sites in the GSDMC promoter were predicted. p-Y705-Stat3-binding site was validated by screening with luciferase reporter assay. (b-d) Immunoblotting of GSDME in MDA-MB-231 cells. H226 cells as a positive control. Immunoblotting of the endogenous GSDME (b). Immunoblotting of GSDME in MDA-MB-231 cells under hypoxia (c). Immunoblotting of GSDME in MDA-MB-231 cells treated with daunorubicin, doxorubicin, epirubicin, and actinomycin D (d). The experiments of (b-d) were repeated three times with similar results respectively. Unprocessed blots are provided in Source Data Extended Data Fig. 4.
Extended Data Fig. 5 Caspase-8, but not caspase-6, is required for TNFα-induced pyroptosis under hypoxia.
(a and b) MDA-MB-231 cells were cultured under normoxia (N) or hypoxia (H) and treated as indicated. LDH-released cell death was measured as shown in (a). Data are shown as mean ± SD of n = 3 independent experiments. P values were determined by two-sided Student’s t-test. Representative images of dying cell morphology of three independent experiments (b). Red arrows indicate cell swelling with big bubbles. Scale bar, 20 μm. Caspase-3i, caspase-3 inhibitor Z-DEVD-FMK; Caspase-6i, caspase-6 inhibitor Z-VEID-FMK; Caspase-8i, caspase-8 inhibitor Z-IETD-FMK (each at 10 μM). Statistical source data are provided in Source Data Extended Data Fig. 5.
Extended Data Fig. 6 The pyroptotic cell death in cell lines with different GSDMC and PD-L1 expression status.
(a) Immunoblotting of PD-L1 and GSDMC in Hs578T, BT549, MDA-MB-231 and MCF-7 cells. Blots are representative of three independent experiments. (b) LDH-released cell death in Hs578T, BT549, MDA-MB-231 and MCF-7 cells treated with TNFα under normoxia or hypoxia. Data are shown as mean ± SD of n = 3 independent experiments. P values were determined by two-sided Student’s t-test. Statistical source data and unprocessed blots are provided in Source Data Extended Data Fig. 6.
Extended Data Fig. 7 The N-terminal domain (1–365) of GSDMC is sufficient to induce pyroptosis.
(a) 3D structural modelling of GSDMC. The modelling was performed using SWISS-MODEL. The structural fold of GSDMC N- and C-terminal domains is in blue cartoon presentation. The linker region and the solvent-exposed loop region of the C-terminal domain of GSDMC are coloured in green with numbered Asp/Glu residues coloured in magenta. The structural figure was prepared using PyMOL Molecular Graphics System, Version 2.0 Schrödinger, LLC. (b) Confocal microscopy analysis of the distribution of GSDMC-FL, GSDMC (1¬–365), and GSDMD-N expressions in HeLa cells. GSDMD-N, N-terminal domain of GSDMD. Images are representative of three independent experiments. Scale bar, 20 μm. (c–d) HeLa cells were transfected with GSDMC-FL or GSDMC (1–365). Representative images of dying cell morphology of three independent experiments (c). Scale bar, 20 μm. Dying cells were stained with nucleic acid dye SYTOX green. Red arrows indicate cell swelling with large bubbles. LDH-released cell death is shown as mean ± SD of n = 3 independent experiments (d). P values were determined by two-sided Student’s t-test. (e) HEK293T cells were transfected with Flag-GSDMC-FL or Flag-GSDMC (1–365) for 16 h. Supernatants and cell pellets were collected and subjected to immunoblotting. The blot is representative of three independent experiments. (f) Cartoon diagram of GSDMC structure and the cleavage by caspase-8 across species. Statistical source data and unprocessed blots are provided in Source Data Extended Data Fig. 7.
Extended Data Fig. 8 Nuclear PD-L1 translocation in vivo causes poor survival in both immunocompetent and nude mice.
(a) IHC staining of HIF1α, GSDMC, PD-L1, and p-Y705-Stat3 in MDA-MB-231 xenografts in nude mice (n = 8). MDA-MB-231 cells were injected subcutaneously into nude mice. Four weeks after tumour cell injection, all mice were sacrificed and tumours excised for IHC staining. The expression levels and patterns of HIF1α, GSDMC, PD-L1, and p-Y705-Stat3 in normoxic and hypoxic area in tumours were analysed and indicated by green arrows. Scale bar, 100 μm. The experiment was repeated three times with similar results. (b) Survival analysis of 4T1 cells expressing PD-L1-WT or PD-L1-NLS in BALB/c nude mice. Data are shown of n = 10 mice. The experiment was repeated three times with similar results. Statistical significance for survival analysis was determined using a log-rank (Mantel–Cox) test. (c) Survival analysis of 4T1 cells expressing PD-L1-WT or PD-L1-NLS in immunocompetent BALB/c mice. Data are shown of n = 10 mice. The experiment was repeated three times with similar results. Statistical significance for survival analysis was determined using a log-rank (Mantel–Cox) test. Statistical source data are provided in Source Data Extended Data Fig. 8.
Extended Data Fig. 9 Chemotherapeutic drugs induce pyroptotic cell death by nPD-L1/GSDMC-mediated non-canonical pathway in breast tumour.
(a) Representative images of dying cell morphology by phase-contrast microscopy in MDA-MB-231 stable transfectants as indicated treated with daunorubicin, doxorubicin, epirubicin, and actinomycin D. The experiment was repeated three times with similar results. Scale bar, 20 μm. (b) LDH-released cell death in MDA-MB-231 stable transfectants as indicated treated with daunorubicin, doxorubicin, epirubicin, and actinomycin D. Data are shown as mean ± SD of n = 3 independent experiments. P values were determined by two-sided Student’s t-test. Statistical source data are provided in Source Data Extended Data Fig. 9.
Extended Data Fig. 10 A proposed model of nuclear PD-L1-mediated apoptosis-to-pyroptosis switch under hypoxia.
Under normoxia, caspase-3 can be activated by TNFα-induced caspase-8 to cause apoptotic cell death. However, hypoxia-activated p-Stat3 facilitates nPD-L1 translocation. nPD-L1 cooperates with p-Stat3 to transcriptionally activate GSDMC expression. GSDMC is specifically cleaved by macrophage-derived TNFα-activated caspase-8, which generates an N-terminal pore-forming fragment that causes pyroptotic cell death to induce tumour necrosis in breast cancer.
Supplementary information
Supplementary Video 1
MDA-MB-231–GFP cells were treated with TNFα plus CHX under normoxia. MDA-MB-231–GFP, MDA-MB-231 cells with stable expression of GFP protein.
Supplementary Video 2
MDA-MB-231–GFP cells were treated with TNFα plus CHX under hypoxia.
Supplementary Video 3
MDA-MB-231–GFP cells were co-treated with TNFα plus CHX and the Stat3 inhibitor HO-3867 (20 μM) under hypoxia.
Supplementary Video 4
MDA-MB-231–GFP cells were co-treated with TNFα plus CHX and caspase-8 inhibitor Z-IETD-FMK (10 μM) under hypoxia.
Supplementary Table 1
The final concentrations for TNFα, CHX and various stimuli.
Source data
Source Data Fig. 1
Statistical source data
Source Data Fig. 1
Unprocessed western blots
Source Data Fig. 2
Unprocessed western blots
Source Data Fig. 3
Statistical source data
Source Data Fig. 4
Statistical source data
Source Data Fig. 4
Unprocessed western blots
Source Data Fig. 5
Statistical source data
Source Data Fig. 5
Unprocessed western blots
Source Data Fig. 6
Statistical source data
Source Data Fig. 6
Unprocessed western blots and gels
Source Data Fig. 7
Statistical source data
Source Data Fig. 8
Statistical source data
Source Data Fig. 8
Unprocessed western blots
Source Data Extended Data Fig. 1
Unprocessed western blots
Source Data Extended Data Fig. 2
Unprocessed western blots
Source Data Extended Data Fig. 3
Statistical source data
Source Data Extended Data Fig. 3
Unprocessed western blots
Source Data Extended Data Fig. 4
Unprocessed western blots
Source Data Extended Data Fig. 5
Statistical source data
Source Data Extended Data Fig. 6
Statistical source data
Source Data Extended Data Fig. 6
Unprocessed western blots
Source Data Extended Data Fig. 7
Statistical source data
Source Data Extended Data Fig. 7
Unprocessed western blots
Source Data Extended Data Fig. 8
Statistical source data
Source Data Extended Data Fig. 9
Statistical source data
Rights and permissions
About this article
Cite this article
Hou, J., Zhao, R., Xia, W. et al. PD-L1-mediated gasdermin C expression switches apoptosis to pyroptosis in cancer cells and facilitates tumour necrosis. Nat Cell Biol 22, 1264–1275 (2020). https://doi.org/10.1038/s41556-020-0575-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-0575-z
This article is cited by
-
Melanoma biology and treatment: a review of novel regulated cell death-based approaches
Cancer Cell International (2024)
-
Pyroptosis: a double-edged sword in lung cancer and other respiratory diseases
Cell Communication and Signaling (2024)
-
Epigenetic regulation of diverse cell death modalities in cancer: a focus on pyroptosis, ferroptosis, cuproptosis, and disulfidptosis
Journal of Hematology & Oncology (2024)
-
Multiomics characterization of pyroptosis in the tumor microenvironment and therapeutic relevance in metastatic melanoma
BMC Medicine (2024)
-
NINJ1 mediates inflammatory cell death, PANoptosis, and lethality during infection conditions and heat stress
Nature Communications (2024)