Abstract
Immunotherapies that target programmed cell death protein 1 (PD-1) and its ligand PD-L1 as well as cytotoxic T-lymphocyte-associated protein 4 (CTLA4) have shown impressive clinical outcomes for multiple tumours. However, only a subset of patients achieves durable responses, suggesting that the mechanisms of the immune checkpoint pathways are not completely understood. Here, we report that PD-L1 translocates from the plasma membrane into the nucleus through interactions with components of the endocytosis and nucleocytoplasmic transport pathways, regulated by p300-mediated acetylation and HDAC2-dependent deacetylation of PD-L1. Moreover, PD-L1 deficiency leads to compromised expression of multiple immune-response-related genes. Genetically or pharmacologically modulating PD-L1 acetylation blocks its nuclear translocation, reprograms the expression of immune-response-related genes and, as a consequence, enhances the anti-tumour response to PD-1 blockade. Thus, our results reveal an acetylation-dependent regulation of PD-L1 nuclear localization that governs immune-response gene expression, and thereby advocate targeting PD-L1 translocation to enhance the efficacy of PD-1/PD-L1 blockade.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The next-generation sequencing data generated in this study have been submitted to the Gene Expression Omnibus Database under the accession numbers GSE134510, GSE146557 and GSE146648. The data from MS analysis was deposited at the Japan Proteome Standard Repository/Database (JPOST) under the accession numbers JPST000666/PXD015191 and JPST000757/PXD017707, respectively. The human cancer data were derived from the TCGA Research Network (http://cancergenome.nih.gov/) and the Riaz2017_PD1 cohort68. The dataset derived from this resource that supports the findings of this study is available at GEPIA (http://gepia.cancer-pku.cn)66 and TIDE (http://tide.dfci.harvard.edu)67. All other data supporting the findings of this study are available from the corresponding authors on reasonable request. Source data are provided with this paper.
Code availability
Custom scripts used in the study are available at https://github.com/ejgkelvin/Nuclear_PD-L1_Acetylation.
References
Chen, D. S. & Mellman, I. Elements of cancer immunity and the cancer-immune set point. Nature 541, 321–330 (2017).
Baumeister, S. H., Freeman, G. J., Dranoff, G. & Sharpe, A. H. Coinhibitory pathways in immunotherapy for cancer. Annu. Rev. Immunol. 34, 539–573 (2016).
Ribas, A. & Wolchok, J. D. Cancer immunotherapy using checkpoint blockade. Science 359, 1350–1355 (2018).
Sharma, P., Hu-Lieskovan, S., Wargo, J. A. & Ribas, A. Primary, adaptive, and acquired resistance to cancer immunotherapy. Cell 168, 707–723 (2017).
Restifo, N. P., Smyth, M. J. & Snyder, A. Acquired resistance to immunotherapy and future challenges. Nat. Rev. Cancer 16, 121–126 (2016).
Zhang, J. et al. Cyclin D-CDK4 kinase destabilizes PD-L1 via cullin 3-SPOP to control cancer immune surveillance. Nature 553, 91–95 (2018).
Lim, S. O. et al. Deubiquitination and stabilization of PD-L1 by CSN5. Cancer Cell 30, 925–939 (2016).
Zhang, J., Dang, F., Ren, J. & Wei, W. Biochemical aspects of PD-L1 regulation in cancer immunotherapy. Trends Biochem. Sci. 43, 1014–1032 (2018).
Galluzzi, L., Chan, T. A., Kroemer, G., Wolchok, J. D. & Lopez-Soto, A. The hallmarks of successful anticancer immunotherapy. Sci. Transl. Med. 10, eaat7807 (2018).
Ansell, S. M. et al. PD-1 blockade with nivolumab in relapsed or refractory Hodgkin’s lymphoma. N. Engl. J. Med. 372, 311–319 (2015).
Inuzuka, H. et al. Acetylation-dependent regulation of Skp2 function. Cell 150, 179–193 (2012).
Nihira, N. T. et al. Acetylation-dependent regulation of MDM2 E3 ligase activity dictates its oncogenic function. Sci. Signal. 10, eaai8026 (2017).
Song, H. et al. Acetylation of EGF receptor contributes to tumor cell resistance to histone deacetylase inhibitors. Biochem. Biophys. Res. Commun. 404, 68–73 (2011).
Lin, S. Y. et al. Nuclear localization of EGF receptor and its potential new role as a transcription factor. Nat. Cell Biol. 3, 802–808 (2001).
Gao, Y. S., Hubbert, C. C. & Yao, T. P. The microtubule-associated histone deacetylase 6 (HDAC6) regulates epidermal growth factor receptor (EGFR) endocytic trafficking and degradation. J. Biol. Chem. 285, 11219–11226 (2010).
Li, C. W. et al. Glycosylation and stabilization of programmed death ligand-1 suppresses T-cell activity. Nat. Commun. 7, 12632 (2016).
Mezzadra, R. et al. Identification of CMTM6 and CMTM4 as PD-L1 protein regulators. Nature 549, 106–110 (2017).
Horita, H., Law, A., Hong, S. & Middleton, K. Identifying regulatory posttranslational modifications of PD-L1: a focus on monoubiquitinaton. Neoplasia 19, 346–353 (2017).
Lasko, L. M. et al. Discovery of a selective catalytic p300/CBP inhibitor that targets lineage-specific tumours. Nature 550, 128–132 (2017).
Glozak, M. A., Sengupta, N., Zhang, X. & Seto, E. Acetylation and deacetylation of non-histone proteins. Gene 363, 15–23 (2005).
Pavlik, C. M. et al. Santacruzamate A, a potent and selective histone deacetylase inhibitor from the Panamanian marine cyanobacterium cf. Symploca sp. J. Nat. Prod. 76, 2026–2033 (2013).
von Kleist, L. et al. Role of the clathrin terminal domain in regulating coated pit dynamics revealed by small molecule inhibition. Cell 146, 471–484 (2011).
Schnitzer, J. E., Oh, P., Pinney, E. & Allard, J. Filipin-sensitive caveolae-mediated transport in endothelium: reduced transcytosis, scavenger endocytosis, and capillary permeability of select macromolecules. J. Cell Biol. 127, 1217–1232 (1994).
McMahon, H. T. & Boucrot, E. Molecular mechanism and physiological functions of clathrin-mediated endocytosis. Nat. Rev. Mol. Cell Biol. 12, 517–533 (2011).
Bonifacino, J. S. & Traub, L. M. Signals for sorting of transmembrane proteins to endosomes and lysosomes. Annu. Rev. Biochem. 72, 395–447 (2003).
Wang, H. et al. HIP1R targets PD-L1 to lysosomal degradation to alter T cell-mediated cytotoxicity. Nat. Chem. Biol. 15, 42–50 (2019).
Paczkowski, J. E., Richardson, B. C. & Fromme, J. C. Cargo adaptors: structures illuminate mechanisms regulating vesicle biogenesis. Trends Cell Biol. 25, 408–416 (2015).
Mattera, R., Boehm, M., Chaudhuri, R., Prabhu, Y. & Bonifacino, J. S. Conservation and diversification of dileucine signal recognition by adaptor protein (AP) complex variants. J. Biol. Chem. 286, 2022–2030 (2011).
Fazal, F., Minhajuddin, M., Bijli, K. M., McGrath, J. L. & Rahman, A. Evidence for actin cytoskeleton-dependent and -independent pathways for RelA/p65 nuclear translocation in endothelial cells. J. Biol. Chem. 282, 3940–3950 (2007).
Cortes-Reynosa, P., Robledo, T. & Salazar, E. P. Epidermal growth factor promotes epidermal growth factor receptor nuclear accumulation by a pathway dependent on cytoskeleton integrity in human breast cancer cells. Arch. Med. Res. 40, 331–338 (2009).
Elosegui-Artola, A. et al. Force triggers YAP nuclear entry by regulating transport across nuclear pores. Cell 171, 1397–1410 (2017).
Satelli, A. et al. Potential role of nuclear PD-L1 expression in cell-surface vimentin positive circulating tumor cells as a prognostic marker in cancer patients. Sci. Rep. 6, 28910 (2016).
Yu, Y. et al. Cancer-associated fibroblasts induce epithelial-mesenchymal transition of breast cancer cells through paracrine TGF-β signalling. Br. J. Cancer 110, 724–732 (2014).
Goldfarb, D. S., Corbett, A. H., Mason, D. A., Harreman, M. T. & Adam, S. A. Importin alpha: a multipurpose nuclear-transport receptor. Trends Cell Biol. 14, 505–514 (2004).
Wagstaff, K. M., Sivakumaran, H., Heaton, S. M., Harrich, D. & Jans, D. A. Ivermectin is a specific inhibitor of importin α/β-mediated nuclear import able to inhibit replication of HIV-1 and dengue virus. Biochem. J. 443, 851–856 (2012).
Isokane, M. et al. Plasma-membrane-anchored growth factor pro-amphiregulin binds A-type lamin and regulates global transcription. J. Cell Sci. 121, 3608–3618 (2008).
Hancock, M. L. et al. Insulin receptor associates with promoters genome-wide and regulates gene expression. Cell 177, 722–736 (2019).
Tu, X. et al. PD-L1 (B7-H1) competes with the RNA exosome to regulate the DNA damage response and can be targeted to sensitize to radiation or chemotherapy. Mol. Cell 74, 1215–1226 (2019).
Zou, W., Wolchok, J. D. & Chen, L. PD-L1 (B7-H1) and PD-1 pathway blockade for cancer therapy: mechanisms, response biomarkers, and combinations. Sci. Transl. Med. 8, 328rv324 (2016).
He, G. & Karin, M. NF-kappaB and STAT3—key players in liver inflammation and cancer. Cell Res. 21, 159–168 (2011).
Ceeraz, S., Nowak, E. C. & Noelle, R. J. B7 family checkpoint regulators in immune regulation and disease. Trends Immunol. 34, 556–563 (2013).
Bertrand, F. et al. TNFalpha blockade overcomes resistance to anti-PD-1 in experimental melanoma. Nat. Commun. 8, 2256 (2017).
Perez-Ruiz, E. et al. Prophylactic TNF blockade uncouples efficacy and toxicity in dual CTLA-4 and PD-1 immunotherapy. Nature 569, 428–432 (2019).
Azuma, T. et al. B7-H1 is a ubiquitous antiapoptotic receptor on cancer cells. Blood 111, 3635–3643 (2008).
Chang, C. H. et al. Metabolic competition in the tumor microenvironment is a driver of cancer progression. Cell 162, 1229–1241 (2015).
Patel, S. P. & Kurzrock, R. PD-L1 expression as a predictive biomarker in cancer immunotherapy. Mol. Cancer Ther. 14, 847–856 (2015).
Nelson-Rees, W. A., Daniels, D. W. & Flandermeyer, R. R. Cross-contamination of cells in culture. Science 212, 446–452 (1981).
Dai, X. et al. Prostate cancer-associated SPOP mutations confer resistance to BET inhibitors through stabilization of BRD4. Nat. Med. 23, 1063–1071 (2017).
Wei, W. et al. Degradation of the SCF component Skp2 in cell-cycle phase G1 by the anaphase-promoting complex. Nature 428, 194–198 (2004).
Mahoney, K. M. et al. PD-L1 antibodies to its cytoplasmic domain most clearly delineate cell membranes in immunohistochemical staining of tumor cells. Cancer Immunol. Res. 3, 1308–1315 (2015).
Chaudhri, A. et al. PD-L1 binds to B7-1 only in cis on the same cell surface. Cancer Immunol. Res. 6, 921–929 (2018).
Liang, S. C. et al. Regulation of PD-1, PD-L1, and PD-L2 expression during normal and autoimmune responses. Eur. J. Immunol. 33, 2706–2716 (2003).
Svendsen, S., Zimprich, C., McDougall, M. G., Klaubert, D. H. & Los, G. V. Spatial separation and bidirectional trafficking of proteins using a multi-functional reporter. BMC Cell Biol. 9, 17 (2008).
Zeng, H. et al. Systematic identification of Ctr9 regulome in ERα-positive breast cancer. BMC Genom. 17, 902 (2016).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Huang, W., Loganantharaj, R., Schroeder, B., Fargo, D. & Li, L. PAVIS: a tool for peak annotation and visualization. Bioinformatics 29, 3097–3099 (2013).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Tang, Z. et al. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 45, W98–W102 (2017).
Jiang, P. et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat. Med. 24, 1550–1558 (2018).
Riaz, N. et al. Tumor and microenvironment evolution during immunotherapy with nivolumab. Cell 171, 934–949 (2017).
Acknowledgements
We thank J. Guo, F. Dang and other members of the Wei laboratory for reading the manuscript, as well as members of the Wei, Freeman and Sicinski laboratories for helpful discussions. We thank staff at the Microscopy Resources on the North Quad (MicRoN) core at Harvard Medical School for helping with IF experiments. This research was supported in part by the NIH grants (R01CA177910 and R01GM094777, to W.W.; P50CA101942, to G.J.F.; R01CA236226 and R01CA202634, to P.S.; and R01CA236356, to W.X.); the Japan Society for Promotion of Science (JSPS) KAKENHI Grant (JP18H06157, to N.T.N). N.T.N. is supported by JSPS Research Fellowships for Young Scientists and the Osamu Hayaishi Memorial Scholarship for Study Abroad.
Author information
Authors and Affiliations
Contributions
Y.Gao, N.T.N. and X.B. designed and performed the experiments with assistance from J.Z., C.C., Y.F., Y.-H.H., L.M. A.K., X.D., S.S., Y.Geng, D.W., H.I., B.J.N. and L.L.; N.T.N., M.O., A.N. and J.L. performed the MS analysis. A.K., W.X. and N.T.C. performed the ChIP experiments. H.L., A.N. and M.O. analysed the data. C.C. and X.S.L. helped with the bioinformatics analysis. Y.M., P.S., G.J.F. and W.W. guided and supervised the study. N.T.N., Y.Gao, J.Z. and W.W. wrote the manuscript. All of the authors commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
G.J.F. is an inventor on patents covering the PD-1/PD-L1 pathway including US Patent Nos. 6,808,710; 7,038,013; 7,101,550; 7,105,328; 7,638,492; 7,700,301; 7,432,059; 7,709,214; and 7,722,868 and associated foreign patent issuances. These patents have been licensed non-exclusively to Roche, Merck, Bristol-Myers-Squibb, EMD-Serono, Boehringer-Ingelheim, AstraZeneca, Leica, Mayo Clinic, Dako and Novartis and several research reagent providers; and he receives royalties based on those licenses. G.J.F. was recently determined to be an inventor on US Patent Nos. 7,595,048; 8,168,179; 8,728,474; 9,067,999; 9,073,994; and 9,402,899 covering the PD-1/PD-L1 pathway and may in the future receive royalty payments based on licenses to those patents. G.J.F. has served on the advisory boards for Roche, Bristol-Myers-Squibb, Xios, Origimed, Triursus, iTeos, NextPoint, IgM and Jubilant. G.J.F. has equity in Nextpoint, Triursus, Xios, iTeos, IgM, and GV20.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Lysine 263 (K263) within the cytoplasmic domain of PD-L1 is acetylated.
a, Immunoblot (IB) analysis of whole-cell lysates (WCL) and anti-PD-L1 immunoprecipitates (IPs) derived from MDA-MB-468, BT-549 and BT-20 cells. Immunoglobulin G (IgG) served as a negative control. b, Authentication results of the BT-20 cell line performed by ATCC. c, IB analysis of WCL and anti-Myc IPs derived from 293T cells transfected with HA-p300 and Myc-wild type (WT) PD-L1 or the deletion mutant of C-tail (amino acids (AA) 263-290). d, IB analysis of WCL derived from 293T cells transfected with HA-tag-inserted (HA-ins) or Myc-tagged wild-type (WT) or del. C-tail PD-L1 with or without 1 μg/ml tunicamycin treatment overnight. e, Predicted lysine acetylation sites by the Web Server for KAT-specific Acetylation Site Prediction (ASEB) analysis. f, A schematic diagram of the PD-L1 Lys263 acetylated peptide and non-acetylated peptide used for immunization to generate the anti-Ac-K263 PD-L1 antibody. g, Dot-blot testing of acetylated and non-acetylated peptides using indicated purified antibodies. h, IB analysis of WCL and anti-HA IPs derived from 293T cells transfected with HA-ins-PD-L1 WT or the K263R mutant. i, Mass-spectrometry detection of Lys263 acetylation using a synthetic peptide (AA 261 to 270) following in vitro acetylation assay. The blots and western blots in a, c, d, g and h were performed for n=2 independent experiments with similar results. Unprocessed immunoblots are shown in Source Data Extended Data Fig. 1.
Extended Data Fig. 2 HDAC2 mediates deacetylation of PD-L1.
a, IB analysis of WCL and anti-Flag IPs derived from 293T cells transfected with Myc-p300, HA-ins-PD-L1 and/or Flag-tagged deacetylases. b, IB analysis of WCL and Ni-NTA pull-down products from MDA-MB-231 WT and HDAC2 knockout (KO) cells transfected with His-Ub and treated with 10 μM MG-132 overnight. c, d, IB analysis of WCL derived from BT-549 CD274 KO cells transfected with HA-PD-L1 WT, K263R or K263Q mutants and treated with 150 μg/ml cycloheximide (CHX) for indicated hours (c). Signal intensity of PD-L1 protein was quantified by ImageJ as indicated (d). e, IB analysis of WCL and anti-Myc IPs derived from 293T cells transfected with indicated constructs. Western blots in a-c and e were performed for n=2 independent experiments with similar results. Unprocessed immunoblots are shown in Source Data Extended Data Fig. 2.
Extended Data Fig. 3 Lysine 263 (K263) acetylation regulates PD-L1 nuclear translocation.
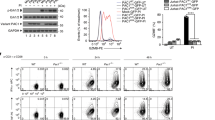
a, Immunofluorescence (IF) staining of human PD-L1 (clone 9A11) and DAPI of MDA-MB-231 WT and CD274 KO cells. Scale bars, 10 μm. b, c, Fractionation analysis using kit from Cell Signaling Technology (CST, #9038) for PD-L1 in human MDA-MB-436, Hs578T and BT-549 cells (b), as well as in mouse CT26, MC38, and B16F10 cells (c). d, Fractionation analysis using kits from Thermo Fisher Scientific™ (#78840) for PD-L1 in indicated cell lines. e Quantification of PD-L1 protein abundance of indicated compartments in MDA-MB-231 cells. Data were presented as mean ± s.d. (n=3 biologically independent samples). f, Fractionation analysis for PD-L1 from 293T cells transfected with mouse PD-L1. g, h, Fractionation analysis for PD-L1 in RAW264.7 cells stimulated with 1 μg/ml Lipopolysaccharide (LPS) for 16 hours (g) and in mouse embryonic fibroblasts (h). i, Z-stacks confocal microscopy images (3x close-up of the source picture) for IF study in Fig. 3d. PD-L1, yellow color and DAPI, blue. j, Fluorescence images of MDA-MB-231 CD274 KO cells transduced with Halo-PD-L1 (AF488) or its C-tail deletion mutant. Scale bars, 5 μm. k, IF staining of mouse PD-L1 (clone 5C5) in CT26 Cd274 KO cells transduced with mouse Cd274 WT, K262R or K262Q mutant lentivirus. Scale bars, 5 μm. l, IB analysis of WCL and anti-PD-L1 IPs derived from indicated fractions of MDA-MB-231 cells. m, Fractionation analysis for BT-549 cells treated with 50 μM HDAC2 inhibitor for 6 hrs. n, Fractionation analysis for PD-L1 in MDA-MB-231 WT or HDAC2 KO cells. The Western blots in b-d, f-h, l-n, and IF studies in a, j and k were performed for n=2 independent experiments with similar results. Unprocessed immunoblots are shown in Source Data Extended Data Fig. 3.
Extended Data Fig. 4 Protein interacting network likely mediates PD-L1 nuclear-translocation process.
a, Results from mass spectrometry analysis were analyzed for GO term enrichment. Red stars denote pathways associated with protein translocation. n = 2 independent experiments with similar results. P values were calculated using hypergeometric test. b, IB of WCL and anti-HA IPs derived from 293T cells transfected with HA-ins-PD-L1 and mouse Hip1r-GFP, and treated with HDAC2 inhibitor for 6 hrs. c, IB of WCL and anti-Flag IPs derived from 293T cells transfected with PD-L1 WT or glycosylation-deficient 4NQ (N35, N192, N200 and N219) mutant. d, Fractionation analysis for PD-L1 from 293T cells transfected with WT or the glycosylation-deficient 4NQ mutant. e, Schematic diagram depicting the working model for endocytosis of PD-L1 from plasma membrane. f, Fractionation analysis for PD-L1 in Vimentin-low SKBR3 and BT-20 cells. g, Relative abundance of PD-L1 protein in each fraction was quantified and calculated for percentage. Statistics, two-tailed Student’s t-test. h, Fractionation analysis for PD-L1 in CT26 WT and Vim KO clones. i, IB of HCC1937 cells treated with 10 ng/ml Transforming Growth Factor-β1 (TGFβ1) for 14 days. j, Fractionation analysis for PD-L1 in HCC1937 cells treated with 10 ng/ml TGFβ1 for 14 days. k, Fractionation analysis for PD-L1 in MDA-MB-231 cells treated with vehicle or 25 μM Ivermectin (IVM) for 2 hrs. l, A schematic diagram to show the working model for nuclear translocation of PD-L1 from plasma membrane. Western blots in b-d, f, and h-k were performed for n=2 independent experiments with similar results. Unprocessed immunoblots are shown in Source Data Extended Data Fig. 4. Statistical source data are available in Statistical Source Data Extended Data Fig. 4.
Extended Data Fig. 5 Nuclear PD-L1 likely stimulates the gene expression of pro-inflammation pathways.
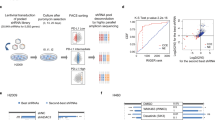
a, DNA binding assays of purified PD-L1 with biotinylated DNA in vitro. b, c, DNA binding assays of biotinylated DNA and 293T cells transfected with indicated constructs. d, DNA biding assays of transfected 293T cells treated with Acy957. e, Numbers of differentially expressed genes upon CD274 or Cd274 KO. f, Top 5 enriched immune response-related GO terms upon Cd274 KO in CT26 cells, analyzed by Fisher-exact test with Benjamini-Hochberg correction. g, GSEA signature upon CD274 KO in MDA-MB-231 cells. h, Heatmap display of interferon γ genes upon CD274 KO in MDA-MB-231 cells. i, Prediction analysis for transcription factors regulating down-regulated genes upon CD274 KO in MDA-MB-231 cells. j, GSEA signatures upon Cd274 KO in CT26 cells. k, l, GSEA signatures of pathways in CT26 Cd274 KO cells restored WT or K262Q mutant Cd274. m, RT-qPCR analysis of BT-549 CD274 KO cells transfected with CD274 WT or K263Q mutants. Data are shown as mean ± s.d. of n=3 independent experiments. Statistics, two-tailed Student’s t-test. n. Hierarchical clustering of ChIP-seq binding profiles and two replicates of PD-L1 binding profiles genome-wide in MDA-MB-231 cells. o. IB of WCL and anti-PD-L1 IPs derived from MDA-MB-231 cells. p, q, IB of WCL and IPs derived from 293T cells transfected with indicated constructs. r, Schematic diagram showing how nuclear PD-L1 enhances the immunotherapy response through affecting expression of immune-related genes. GSEA analyses in g and j-l were performed using Kolmogorov-Smirnov statistic. Biologically independent sequenced samples/group for f-j, n=4; for k and l, n=3. The blots and Western blots in a-d and o-q were performed for n=2 independent experiments with similar results. Unprocessed immunoblots are shown in Source Data Extended Data Fig. 5. Statistical source data are available in Statistical Source Data Extended Data Fig. 5.
Extended Data Fig. 6 PD-L1 expression levels correlate with and regulate immune-checkpoint genes.
a, RT-qPCR analysis of genes upon Cd274 KO in CT26 cells. b, IB of MDA-MB-231 cells transfected with control or CD274 siRNAs. c, d, IB (c) and RT-qPCR (d) analysis of MDA-MB-231 cells with CD274 knockdown by shRNAs. e, IB of WCL derived from breast cancer cell lines. f, g, Pearson correlation (two-tailed) analysis for PD-L1 mRNA (Z-score) with PD-L2 (f) or VISTA (g) in breast cancer cell lines (GSE36139). Red line, linear regression line. h, HDAC2 expression profiled by GEPIA. Tumour (T), red dots; normal tissues (N), green dots. i, Overall survival of patients with high (>70%, red curve) and low (<30%, blue curve) HDAC2 (i) or Vimentin (j) analyzed using Log-rank test by GEPIA. k, Progression-free survival (PFS) of melanoma patients (Riaz2017_PD1 cohort, PMID:29033130) treated with PD-1 mAb (Nivolumab) with high or low VIM expression analyzed using Kaplan-Meier curves by TIDE. Ipi_Naive, ipilimumab-naïve (n=25); Ipi_Prog, progressed on ipilimumab (n=26). l-p, RT-qPCR of MDA-MB-231 cells treated with vehicle or HDAC2 inhibitor. These genes are involved in Type I or III interferon pathways (l), STAT1/2 pathways (m), endogenous retrovirus ERVs (n), double-stranded pattern recognition receptors (o), antigen presenting and presentation via MHC class I (p). q, Schematic diagram to show a possible molecular mechanism of acquired PD-1/PD-L1 blockade resistance caused by nuclear PD-L1 (left), and the potential usage of HDAC2 inhibitor (right). Tumor abbreviations are shown in GEPIA. Western blots b-c and e were performed for n=2 independent experiments with similar results. PCR data a, d and l-p were shown as mean ± s.d. of n=3 independent experiments, analyzed by two-tailed Student’s t-test. Unprocessed immunoblots are shown in Source Data Extended Data Fig. 6. Statistical source data are available in Statistical Source Data Extended Data Fig. 6.
Extended Data Fig. 7 Targeting HDAC2 and inhibiting PD-L1 deacetylation can enhance immunotherapy efficacy.
a, b, Tumour growth (a) and survival curves (b) of nude mice bearing MC 38 tumors treated with control antibody, PD-1 mAb, HDAC2 inhibitor or combined therapy. c, TILs from treated MC38 syngeneic tumours (Control, n=6; PD-1 mAb, n=8; HDAC2i, n=6; Combined, n=8) after stimulation were analyzed for Interferon γ (IFNγ), IL-2 and IL-10. d, Immunofluorescence for PD-L1 and DAPI of MC38 syngeneic tumours treated as indicated. Scale bars, 10 μm. n=4 independent samples per group. e, f, Tumour growth (e) and survival curves (f) of BALB/c mice bearing tumor derived from CT26-Cd274 KO cells with re-introduced WT or K262Q Cd274, treated with control antibody or PD-1 mAb. Tumour volume was shown as mean ± s.d. Statistics in e, two-tailed Student’s t-test. g, Tumour growth of MC38/K262Q Cd274 tumour-bearing C57BL/6 mice treated as indicated. h, A schematic diagram of molecular mechanism underling nuclear translocation of PD-L1 and its contradictory functions in immune response. PD-L1 deacetylated by HDAC2 is translocated into the nucleus via interacting with various key regulatory proteins for endocytosis and nuclear translocation, then transactivates immune responsive in the nucleus to impact tumour sensitivity to PD-1 blockage (the lower left panel with yellow background), as well as controlling various immune checkpoint gene expression to possibly confer resistance to PD-1 blockage treatment (the lower right panel with gray background). Thus, HDAC2 inhibitor will reduce PD-L1 nuclear localization to prevent the emerging resistance to PD-1 blockade treatment. P values in b and f were calculated using Gehan-Breslow-Wilcoxo test, two-sided. Statistical source data are available in Statistical Source Data Extended Data Fig. 7.
Supplementary information
Supplementary Tables
Supplementary Table 1: list of PD-L1-interacting proteins detected by anti-Flag IPs coupled with MS analysis (IP–MS). Supplementary Table 2: list of PD-L1-interacting proteins detected by anti-HA IP–MS analysis. Supplementary Table 3: selected list of transport and cytoskeleton proteins that interact with PD-L1, as detected by IP–MS. Supplementary Table 4: list of differentially expressed genes after CD274 KO in MDA-MB-231 cells. Supplementary Table 5: list of differentially expressed genes after Cd274 KO in CT26 cells. Supplementary Table 6: list of the top enriched de novo binding motifs revealed by PD-L1 ChIP–seq assay. Supplementary Table 7: list of top enriched known binding motifs revealed by PD-L1 ChIP–seq assay. Supplementary Table 8: cell line information. Supplementary Table 9: primers for RT–qPCR.
Source data
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 2
Unprocessed western blots.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Unprocessed western blots.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 4.
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 5.
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 6.
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
About this article
Cite this article
Gao, Y., Nihira, N.T., Bu, X. et al. Acetylation-dependent regulation of PD-L1 nuclear translocation dictates the efficacy of anti-PD-1 immunotherapy. Nat Cell Biol 22, 1064–1075 (2020). https://doi.org/10.1038/s41556-020-0562-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-0562-4
This article is cited by
-
CircPPAP2B controls metastasis of clear cell renal cell carcinoma via HNRNPC-dependent alternative splicing and targeting the miR-182-5p/CYP1B1 axis
Molecular Cancer (2024)
-
Hypoxia-activated XBP1s recruits HDAC2-EZH2 to engage epigenetic suppression of ΔNp63α expression and promote breast cancer metastasis independent of HIF1α
Cell Death & Differentiation (2024)
-
Dietary factors and their influence on immunotherapy strategies in oncology: a comprehensive review
Cell Death & Disease (2024)
-
The role of the immunosuppressive PD-1/PD-L1 checkpoint pathway in the aging process and age-related diseases
Journal of Molecular Medicine (2024)
-
CTLA4 DNA methylation is associated with CTLA-4 expression and predicts response to immunotherapy in head and neck squamous cell carcinoma
Clinical Epigenetics (2023)