Abstract
Epigenetic plasticity is a pivotal factor that drives metastasis. Here, we show that the promoter of the gene that encodes the ubiquitin ligase subunit FBXL7 is hypermethylated in advanced prostate and pancreatic cancers, correlating with decreased FBXL7 mRNA and protein levels. Low FBXL7 mRNA levels are predictive of poor survival in patients with pancreatic and prostatic cancers. FBXL7 mediates the ubiquitylation and proteasomal degradation of active c-SRC after its phosphorylation at Ser 104. The DNA-demethylating agent decitabine recovers FBXL7 expression and limits epithelial-to-mesenchymal transition and cell invasion in a c-SRC-dependent manner. In vivo, FBXL7-depleted cancer cells form tumours with a high metastatic burden. Silencing of c-SRC or treatment with the c-SRC inhibitor dasatinib together with FBXL7 depletion prevents metastases. Furthermore, decitabine reduces metastases derived from prostate and pancreatic cancer cells in a FBXL7-dependent manner. Collectively, this research implicates FBXL7 as a metastasis-suppressor gene and suggests therapeutic strategies to counteract metastatic dissemination of pancreatic and prostatic cancer cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The human pancreatic and prostate cancer data reported in Fig. 1a and Extended Data Fig. 1a were derived from the TCGA Research network http://cancergenome.nih.gov/ and the pan-cancer methylation database MethHC11. The datasets derived from the resources that support the findings of this study are reported in the source data. All other data supporting the findings are available from the corresponding authors on reasonable request. Source data are provided with this paper.
References
Lambert, A. W., Pattabiraman, D. R. & Weinberg, R. A. Emerging biological principles of metastasis. Cell 168, 670–691 (2017).
Harper, K. L. et al. Mechanism of early dissemination and metastasis in Her2+ mammary cancer. Nature 540, 588–592 (2016).
Hosseini, H. et al. Early dissemination seeds metastasis in breast cancer. Nature 540, 552–558 (2016).
Patel, S. A. & Vanharanta, S. Epigenetic determinants of metastasis. Mol. Oncol. 11, 79–96 (2016).
Pfeifer, G. P. Defining driver DNA methylation changes in human cancer. Int. J. Mol. Sci. 19, 1166 (2018).
Liu, L. et al. Association of tissue promoter methylation levels of APC, TGFbeta2, HOXD3 and RASSF1A with prostate cancer progression. Int. J. Cancer 129, 2454–2462 (2011).
Skrypek, N., Goossens, S., De Smedt, E., Vandamme, N. & Berx, G. Epithelial-to-mesenchymal transition: epigenetic reprogramming driving cellular plasticity. Trends Genet. 33, 943–959 (2017).
Skaar, J. R., Pagan, J. K. & Pagano, M. SCF ubiquitin ligase-targeted therapies. Nat. Rev. Drug Discov. 13, 889–903 (2014).
Wang, Z., Liu, P., Inuzuka, H. & Wei, W. Roles of F-box proteins in cancer. Nat. Rev. Cancer 14, 233–247 (2014).
Heo, J., Eki, R. & Abbas, T. Deregulation of F-box proteins and its consequence on cancer development, progression and metastasis. Semin. Cancer Biol. 36, 33–51 (2016).
Huang, W. Y. et al. MethHC: a database of DNA methylation and gene expression in human cancer. Nucleic Acids Res. 43, D856–D861 (2015).
Grawenda, A. M. & O’Neill, E. Clinical utility of RASSF1A methylation in human malignancies. Br. J. Cancer 113, 372–381 (2015).
Cunningham, D. & You, Z. In vitro and in vivo model systems used in prostate cancer research. J. Biol. Methods 2, e17 (2015).
Bao, J., Gur, G. & Yarden, Y. Src promotes destruction of c-Cbl: implications for oncogenic synergy between Src and growth factor receptors. Proc. Natl Acad. Sci. USA 100, 2438–2443 (2003).
Pettaway, C. A. et al. Selection of highly metastatic variants of different human prostatic carcinomas using orthotopic implantation in nude mice. Clin. Cancer Res. 2, 1627–1636 (1996).
Mayer, E. L. & Krop, I. E. Advances in targeting SRC in the treatment of breast cancer and other solid malignancies. Clin. Cancer Res. 16, 3526–3532 (2010).
Summy, J. M. & Gallick, G. E. Src family kinases in tumor progression and metastasis. Cancer Metastasis Rev. 22, 337–358 (2003).
Bromann, P. A., Korkaya, H. & Courtneidge, S. A. The interplay between Src family kinases and receptor tyrosine kinases. Oncogene 23, 7957–7968 (2004).
Yokouchi, M. et al. Src-catalyzed phosphorylation of c-Cbl leads to the interdependent ubiquitination of both proteins. J. Biol. Chem. 276, 35185–35193 (2001).
Teckchandani, A. et al. Cullin 5 destabilizes Cas to inhibit Src-dependent cell transformation. J. Cell Sci. 127, 509–520 (2014).
Mertins, P. et al. Proteogenomics connects somatic mutations to signalling in breast cancer. Nature 534, 55–62 (2016).
Schreiber, T. B., Mausbacher, N., Keri, G., Cox, J. & Daub, H. An integrated phosphoproteomics work flow reveals extensive network regulation in early lysophosphatidic acid signaling. Mol. Cell. Proteom. 9, 1047–1062 (2010).
Trinidad, J. C. et al. Global identification and characterization of both O-GlcNAcylation and phosphorylation at the murine synapse. Mol. Cell. Proteom. 11, 215–229 (2012).
Kalluri, R. EMT: when epithelial cells decide to become mesenchymal-like cells. J. Clin. Invest 119, 1417–1419 (2009).
Guarino, M. Src signaling in cancer invasion. J. Cell. Physiol. 223, 14–26 (2010).
Lu, G. et al. Phosphorylation of ETS1 by Src family kinases prevents its recognition by the COP1 tumor suppressor. Cancer Cell 26, 222–234 (2014).
Thaper, D. et al. Targeting Lyn regulates snail family shuttling and inhibits metastasis. Oncogene 36, 3964–3975 (2017).
Mukhopadhyay, D. et al. Hypoxic induction of human vascular endothelial growth factor expression through c-Src activation. Nature 375, 577–581 (1995).
Kozlowski, J. M. et al. Metastatic behavior of human tumor cell lines grown in the nude mouse. Cancer Res. 44, 3522–3529 (1984).
Xie, D. et al. Role of DAB2IP in modulating epithelial-to-mesenchymal transition and prostate cancer metastasis. Proc. Natl Acad. Sci. USA 107, 2485–2490 (2010).
Seifert, L. et al. The necrosome promotes pancreatic oncogenesis via CXCL1 and mincle-induced immune suppression. Nature 532, 245–249 (2016).
Bellon, E., Gebauer, F., Tachezy, M., Izbicki, J. R. & Bockhorn, M. Pancreatic cancer and liver metastases: state of the art. Updates Surg. 68, 247–251 (2016).
Birnbaum, D. J. et al. Genome profiling of pancreatic adenocarcinoma. Genes Chromosomes Cancer 50, 456–465 (2011).
Vazquez-Mena, O. et al. Amplified genes may be overexpressed, unchanged, or downregulated in cervical cancer cell lines. PLoS ONE 7, e32667 (2012).
Iwakawa, R. et al. MYC amplification as a prognostic marker of early-stage lung adenocarcinoma identified by whole genome copy number analysis. Clin. Cancer Res. 17, 1481–1489 (2011).
Boelens, M. C. et al. Genomic aberrations in squamous cell lung carcinoma related to lymph node or distant metastasis. Lung Cancer 66, 372–378 (2009).
Coppola, D. Molecular prognostic markers in pancreatic cancer. Cancer Control 7, 421–427 (2000).
Criscuoli, M. L., Nguyen, M. & Eliceiri, B. P. Tumor metastasis but not tumor growth is dependent on Src-mediated vascular permeability. Blood 105, 1508–1514 (2005).
Zhang, X. H. et al. Latent bone metastasis in breast cancer tied to Src-dependent survival signals. Cancer Cell 16, 67–78 (2009).
Patel, A., Sabbineni, H., Clarke, A. & Somanath, P. R. Novel roles of Src in cancer cell epithelial-to-mesenchymal transition, vascular permeability, microinvasion and metastasis. Life Sci. 157, 52–61 (2016).
Dosch, A. R. et al. Src kinase inhibition restores E-cadherin expression in dasatinib-sensitive pancreatic cancer cells. Oncotarget 10, 1056–1069 (2019).
Irby, R. B. & Yeatman, T. J. Role of Src expression and activation in human cancer. Oncogene 19, 5636–5642 (2000).
Lutz, M. P. et al. Overexpression and activation of the tyrosine kinase Src in human pancreatic carcinoma. Biochem. Biophys. Res. Commun. 243, 503–508 (1998).
Tatarov, O. et al. SRC family kinase activity is up-regulated in hormone-refractory prostate cancer. Clin. Cancer Res. 15, 3540–3549 (2009).
Park, S. I. et al. Targeting SRC family kinases inhibits growth and lymph node metastases of prostate cancer in an orthotopic nude mouse model. Cancer Res. 68, 3323–3333 (2008).
Morton, J. P. et al. Dasatinib inhibits the development of metastases in a mouse model of pancreatic ductal adenocarcinoma. Gastroenterology 139, 292–303 (2010).
Brabletz, T. et al. Invasion and metastasis in colorectal cancer: epithelial-mesenchymal transition, mesenchymal-epithelial transition, stem cells and beta-catenin. Cells Tissues Organs 179, 56–65 (2005).
Liu, Y. et al. The proapoptotic F-box protein Fbxl7 regulates mitochondrial function by mediating the ubiquitylation and proteasomal degradation of survivin. J. Biol. Chem. 290, 11843–11852 (2015).
Coon, T. A., Glasser, J. R., Mallampalli, R. K. & Chen, B. B. Novel E3 ligase component FBXL7 ubiquitinates and degrades aurora A, causing mitotic arrest. Cell Cycle 11, 721–729 (2012).
Duan, S. et al. FBXO11 targets BCL6 for degradation and is inactivated in diffuse large B-cell lymphomas. Nature 481, 90–93 (2012).
Zambirinis, C. P. et al. TLR9 ligation in pancreatic stellate cells promotes tumorigenesis. J. Exp. Med. 212, 2077–2094 (2015).
Borowicz, S. et al. The soft agar colony formation assay. J. Vis. Exp. 92, e51998 (2014).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Schade, A. E. et al. Dasatinib, a small-molecule protein tyrosine kinase inhibitor, inhibits T-cell activation and proliferation. Blood 111, 1366–1377 (2008).
Qian, X. L. et al. Dasatinib inhibits c-src phosphorylation and prevents the proliferation of triple-negative breast cancer (tnbc) cells which overexpress syndecan-binding protein (SDCBP). PLoS ONE 12, e0171169 (2017).
Daley, D. et al. γδ T cells support pancreatic oncogenesis by restraining alphabeta T cell activation. Cell 166, e1415 (2016).
Xu, W., Doshi, A., Lei, M., Eck, M. J. & Harrison, S. C. Crystal structures of c-Src reveal features of its autoinhibitory mechanism. Mol. Cell 3, 629–638 (1999).
Acknowledgements
We thank A. Viale for reagents; and S. Courtneidge for reading the manuscript and discussion. M.P. is grateful to T. M. Thor for continuous support. This research was funded by grants from the National Institutes of Health to M.P. (GM136250 and CA76584) and G.M. (CA168611, CA215471 and CA206105) and by the Pancreatic Cancer Action network Translational Research Grant to G.M. M.P. is an Investigator with the Howard Hughes Medical Institute. The NYU Center for Biospecimen Research and Development, Immunohistochemistry Laboratory is supported in part by the Laura and Isaac Perlmutter Cancer Center Support Grant; and NIH/NCI P30CA016087 and the National Institutes of Health S10 Grants (NIH/ORIP S10OD01058 and S10OD018338).
Author information
Authors and Affiliations
Contributions
L.M. designed and performed most of the experiments and cowrote the manuscript. D.S. performed some of the biochemical experiments and generated the Fbxl7−/− FC1242 cells. E.K., W.W., A.M. and G.M. designed and performed the mouse experiments in the pancreatic model systems. E.K. also performed the FACS analyses. A.A.A. and P.K. analysed and scored immunohistochemistry and/or H&E staining. D.S., S.J. and A.T. performed the pan-cancer analysis on TCGA data. N.G. and S.G. performed some of the experiments on FBXL7 promoter methylation in prostate cancer tissues and cell lines. G.P. performed the ZEB1 immunohistochemistry. Y.-A.C., A.D., E.H. and J.-T.H. designed and performed the mouse experiments in the prostate model system. M.P. directed and coordinated the study, oversaw the results and cowrote the manuscript. All of the authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
M.P. has financial interests in CullGen Inc. and Kymera Therapeutics. M.P. is on the SAB of CullGen Inc. and Kymera Therapeutics. M.P. is a consultant for BeyondSpring Pharmaceuticals, CullGen Inc., Kymera Therapeutics, Exo Therapeutics and SEED Therapeutics. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
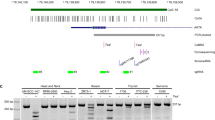
Extended Data Fig. 1 Genes encoding F-box proteins are not highly mutated in human cancers.
a, Heat map showing percentages of samples mutated in the indicated genes, across individual tumor types or pan-cancer, and ranked by their percent mutation rate in pan-cancer. Genes encoding F-box proteins are in black; different cancer genes are in blue. b, Scheme showing the CpG island within the FBXL7 promoter and the primers used to assess FBXL7 promoter methylation by methylation-specific PCR analysis. c, Total RNA extracts from the indicated pancreatic cells were analyzed for FBXL7 mRNA levels by qPCR. Mean ± s.d. is shown; n = 3 independent experiments; P values are from unpaired, two-tailed t-test. d, Total DNA extracts from H6c7, AsPC1, and COLO357 pancreatic cells were subjected to bisulfite modification and sequencing of a FBXL7 promoter region within the CpG island. The table shows the percentage of methylation of the FBXL7 promoter. e, Total mRNA extracts from the indicated prostate cells were analyzed for FBXL7 mRNA levels by qPCR. Mean ± s.d. is shown; n = 3 independent experiments; P values are from unpaired, two-tailed t-test. f, Total DNA extracts from PNT1A, LAPC4, PC-3, PC-3M, and PC-3M-LN4 prostate cells were subjected to bisulfite modification and sequencing of a FBXL7 promoter region within the CpG island. The table shows the percentage of methylation of the FBXL7 promoter. g, Co-expression of FBXL7 and AR mRNAs was analyzed in the prostate TCGA dataset (cBioPortal; n = 450). Two-tailed non-parametric Spearman correlation was used. The linear regression line is shown in red. h, Total mRNA extracts from AsPC1, PC-3, PC-3M and PC-3M-LN4 cells were analyzed for FBXL7 mRNA levels by qPCR upon addition of decitabine for the indicated times. Mean ± s.d. is shown; n = 3 independent experiments; P values are from unpaired, two-tailed t-test. i, PL45, AsPC1, PNT1A, LNCaP, and PC-3 cells were transfected with two different siRNAs to FBXL7 (each individually) or a non-targeting (NT) siRNA. Twenty-four hours after transfection, cells were harvested and plated in 96-well plates in triplicates. Cell proliferation was assessed at the indicated times by measuring absorbance (OD) at 590–650 nm. A representative experiment out of two, each performed in triplicate, is shown. Mean ± s.d. is shown.
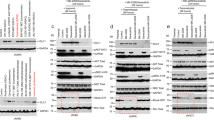
Extended Data Fig. 2 FBXL7 mediates the degradation of active c-SRC.
a, HEK-293T cells were transfected with the indicated FLAG-tagged F-box proteins (FBPs) or an empty vector (EV). Twenty-four hours after transfection, cells were harvested and lysed. Whole-cell extracts (WCE) were subjected to immunoprecipitation (IP) with an anti-FLAG resin and immunoblotting. b, PNT1A cells were treated with MG132 during the last 3 h before lysis. Lysates were immunoprecipitated with either an antibody against c-SRC, an antibody to FBXL7, or nonspecific IgG, and immunoblotted. c, HEK-293T cells were transfected with FLAG-tagged wild-type FBXL7, FLAG-tagged FBXL7(ΔF), or an empty vector (EV). Twenty-four hours after transfection, whole-cell extracts (WCE) were subjected to immunoprecipitation (IP) with an anti-FLAG resin and immunoblotting. d, HEK-293T cells were transfected with FLAG-tagged wild-type c-SRC, FLAG-tagged c-SRC(Y530F), FLAG-tagged c-SRC(Y419F), or an empty vector (EV). Twenty-four hours after transfection, whole-cell extracts (WCE) were subjected to immunoprecipitation (IP) with an anti-FLAG resin and immunoblotting. e, DU145 cells were transfected with a pool of four siRNAs to FBXL7 or a non-targeting (NT) siRNA oligo for 48 h, and treated with cycloheximide (CHX) for the indicated times. Protein extracts were then immunoblotted as indicated. a–e, two independent experiments were performed with similar results.
Extended Data Fig. 3 FBXL7 binds c-SRC phosphorylated on Ser104.
a, Schematic representation of c-SRC mutants. Binding of c-SRC to FBXL7 is indicated with the symbol (+). b–d, HEK-293T cells were transfected with FLAG-tagged versions of either wild-type c-SRC or the indicated c-SRC mutants, or with an empty vector (EV). Twenty-four hours after transfection, whole-cell extracts (WCE) were subjected to immunoprecipitation (IP) with an anti-FLAG resin and immunoblotting. b–d, two independent experiments were performed with similar results. e, Lysates from HEK-293T cells were used in binding reactions with beads coupled to either a peptide or a phospho-peptide flanking the residue S104 in the c-SRC sequence (sequences shown on top of the panel). Bound proteins from three independent experiments were subjected to immunoblotting. f, Alignment of the region corresponding to amino acids 98–119 of human c-SRC in c-SRC orthologs. g, HEK-293T cells were transfected with FLAG-tagged FBXL7. Twenty-four hours after, whole-cell extracts were treated with λ-phosphatase for 4 h, then subjected to immunoprecipitation (IP) with an anti-FLAG resin and immunoblotting. h, Experiment was performed as in b–d. i, Surface structure of human c-SRC obtained from PDB [www.rcsb.org;57]. The amino acid S104 is highlighted in red. Its position shows that it is a solvent-exposed surface amino acid. j, HEK-293T cells were transfected with either one of two FLAG-tagged F-box proteins (FBPs), namely FBXL7 or FBXL1, or with an empty vector (EV). Twenty-four hours after transfection, whole-cell extracts (WCE) were subjected to immunoprecipitation (IP) with an anti-FLAG resin and immunoblotting. g, j: Two independent experiments were performed with similar results.
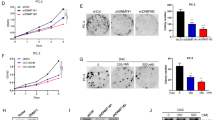
Extended Data Fig. 4 FBXL7 silencing promotes cell migration.
a, MCF-7 cells were transfected with two different siRNAs to FBXL7 or a non-targeting (NT) siRNA oligo (left panels). PL45 cells were transfected with an siRNA oligo to FBXL7 or NT siRNA (right panels). Twenty-four hours after transfection, whole-cell extracts were immunoblotted as indicated. b, PNT1A cells were transfected with an siRNA oligo to FBXL7 (FBXL7 si#1) or a non-targeting (NT) siRNA. Twenty-four hours after transfection, whole-cell extracts (WCE) were denatured and subjected to immunoprecipitation (IP) with anti-ETS1 antibody followed by immunoblotting. ETS1 phosphorylated on Tyr was detected with an anti-phospho-Tyr antibody (PY20). a, b: Two independent experiments were performed with similar results. c, d, PNT1A cells were transfected with an siRNA oligo to FBXL7 (FBXL7 si#1) or a non-targeting (NT) siRNA. Twenty-four hours after, cells were treated with either vehicle alone or two c-SRC kinase inhibitors [SU6656 or dasatinib] for an additional twenty-four hours, and either re-plated (c) or seeded on collagen type I-coated transwells (d). In c, after 18 h, a wound-healing assay was performed up to 48 h in presence or absence of SU6656 or dasatinib. The graph shows a representative experiment out of two, each performed in triplicate. Mean ± s.d. is shown; 24 h, *** P = 0.00093 (NT vs. FBXL7 siRNA), 48 h, *** P = 0.00032 (NT vs. FBXL7 siRNA). P values are from unpaired, two-tailed t-test. In d, after 5 h, cells that migrated on the bottom of the transwells were counted in 10 different fields/well. The graph shows a representative experiment out of two performed for each condition. Mean ± s.d. is shown. n = 10. P values are from unpaired, two-tailed t-test. e, PNT1A cells were transfected with an siRNA oligo to FBXL7 (FBXL7 si#1) or a non-targeting (NT) siRNA. Twenty-four hours after, cells were re-plated in 6-weel plates in a soft agar layer. After three weeks, the colonies were stained with nitro-blue-tetrazolium and photographed. PC-3 cancer cells were used as positive control. f, PNT1A cells treated as in c-d, were analyzed by immunoblotting. g, PC-3 prostate carcinoma cells were transfected with FLAG-tagged FBXL7 or an empty vector (EV). Twenty-four hours after transfection, cells were plated on 96-well plate in triplicates, allowed to adhere and, after 18 h, assayed for cell motility through a wound-healing assay. The graph shows quantification from two independent experiments. e–g, n = 3 independent experiments. In g, mean ± s.d. is shown. P values are from unpaired, two-tailed t-test.
Extended Data Fig. 5 FBXL7 protein expression inversely correlates with c-SRC protein expression in pancreatic and prostate cancer.
a, Representative immunohistochemistry staining images of normal (N) and tumor human pancreatic specimens at different disease stage. Levels of FBXL7 and c-SRC protein in consecutive tissue slides are shown. n = 5 (N) and n = 66 (tumor) independent specimens. Scale bar: 100 μm. b, Representative immunohistochemistry staining images of normal (N) and tumor prostate human specimens at different disease stage. Levels of FBXL7 and c-SRC protein in consecutive tissue slides are shown. n = 10 (N) and n = 84 (tumor) independent specimens. Scale bar: 100 μm.
Extended Data Fig. 6 Decrease in FBXL7 mRNA expression in prostate cancer is not associated with changes in c-SRC mRNA levels.
a–d, Oncomine analysis of FBXL7 and c-SRC mRNA levels in four independent collections of normal and tumor prostate human specimens. N, normal prostate; PCa, prostate carcinoma; PA, prostate adenocarcinoma. a, n = 28 (N), n = 94 (PCa); b, n = 29 (N), n = 155 (PCa), n = 1 (PA); c, n = 13 (N), n = 44 (PCa); d, n = 8 (N), n = 32 (PCa) independent specimens. Median, 10th and 90th percentile are shown in the bar graph. P values are from unpaired, two-tailed t-test, as reported in Oncomine (www.oncomine.org).
Extended Data Fig. 7 FBXL7 suppression promotes prostate cancer metastasis.
a, NOD/SCID-gamma mice were inoculated in the ventral prostate with 5 × 105 PC-3 cells stably transfected with a non-targeting shRNA (NT), shRNAs to FBXL7, or both shRNAs to FBXL7 and c-SRC. Orthotopically transplanted mice were treated with doxycycline to induce shRNA expression (n = 5, NT shRNA; n = 7, FBXL7 shRNA; n = 10, FBXL7/c-SRC shRNA mice). Panels show representative H&E and immunohistochemical staining (antibodies to Ki67 and CD31) of primary tumors. Scale bar: 100 μm. The right panel shows quantification of Ki67 positive cells. Mean ± s.d. is shown. n = 4. P values are from unpaired, two-tailed t-test. b, Representative anti-ZEB1 immunohistochemical staining of primary tumors from the experiment shown in a. n = 3. Scale bar: 10 μm. c, Representative immunohistochemical staining of spleen and coagulating gland (CG) sections from FBXL7 shRNA mice with an antibody specific for human mitochondria. Bottom panel, number of mice with specific micrometastases in different organs/tissues. Scale bar: 100 μm.
Extended Data Fig. 8 Fbxl7 suppression promotes pancreatic cancer metastasis.
a, Schematic representation of the murine Fbxl7 genomic loci and gRNAs target locations. b, Fbxl7 was knocked-out in FC1242 cells using CRISPR technology. 80% confluent FC1242 cells were lysed and whole-cell extracts were immunoblotted. KO clones E11 and G10 are shown. Two independent experiments were performed with similar results. c, Top panels, FC1242 cells (parental and Fbxl7 KO, clone E11) were plated on transwells in the presence of 10% FBS in the outer chamber as chemoattractant. After 6 h, cells that migrated to the bottom of the transwells were counted in 10 different fields/well. Scale bar: 10 μm. The graph shows quantification from a representative experiment out of three independent experiments. n = 10. Middle graph, FC1242 cells (parental and Fbxl7 KO, clones E11 and G10) were harvested and plated in 96-well plates in triplicates. Cell proliferation was assessed by measuring absorbance (OD) at 590–650 nm at the indicated times. A representative experiment of two performed is shown. Bottom graph, FC1242 cells (parental and Fbxl7 KO, clone E11) were plated in a soft agar layer. After 3 weeks, colonies were stained and counted. The graph shows quantification from three independent experiments. Mean ± s.d. is shown. P values are from unpaired, two-tailed t-test. d, FC1242 mouse pancreatic adenocarcinoma cells were transfected with an siRNA oligo to Fbxl7 or a non-targeting siRNA oligo (NT). Twenty-four or forty-eight hours after transfection, whole-cell extracts were immunoblotted. Two independent experiments were performed. e, f, 80,000 Fbxl7 KO FC1242 cells were injected in the pancreas of 8-weeks old C57BL/6 mice. Eighteen days after injection, primary tumors, kidney, peritoneum, liver, epididymis and bladder were collected. Sections of primary tumors were stained with anti-ZEB1 and anti-ZEB2 antibodies (e). Sections of peritoneum, liver and kidney were subjected to H&E staining (f). N, normal tissue; T, tumor tissue. The table on the bottom shows the number of mice exhibiting macrometastases in specific tissues/total number of mice. Scale bars: 20 μm (e), 100 μm (f). g, AsPC1 cells stably expressing non-targeting (NT) shRNA or FBXL7 shRNA were analyzed by immunoblotting (Left panel; two independent experiments performed) or plated in soft agar layer and, three weeks after, colonies were counted. The graph in the right panel shows quantification from three independent experiments Mean ± s.d. is shown. P values are from unpaired, two-tailed t-test. h, 500,000 AsPC1 cells stably expressing NT or FBXL7 shRNA were injected in the tail of the pancreas of 8-weeks old B6.129S7-Rag1tm1Mom/J(002216) mice. Five weeks after injection, primary tumors were collected and sections analyzed by immunohistochemistry with an antibody to ZEB1 or ZEB2. n = 4. Scale bar: 20 μm.
Supplementary information
Supplementary Tables
Supplementary Tables 1–3.
Source data
Source Data Fig. 1
Unprocessed western blots and gels.
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Unprocessed western blots and gels.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Unprocessed western blots.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Unprocessed western blots.
Source Data Extended Data Fig. 8
Statistical source data.
Rights and permissions
About this article
Cite this article
Moro, L., Simoneschi, D., Kurz, E. et al. Epigenetic silencing of the ubiquitin ligase subunit FBXL7 impairs c-SRC degradation and promotes epithelial-to-mesenchymal transition and metastasis. Nat Cell Biol 22, 1130–1142 (2020). https://doi.org/10.1038/s41556-020-0560-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-0560-6
This article is cited by
-
Ubiquitination Process Mediates Prostate Cancer Development and Metastasis through Multiple Mechanisms
Cell Biochemistry and Biophysics (2024)
-
Emerging roles of deubiquitinating enzymes in actin cytoskeleton and tumor metastasis
Cellular Oncology (2024)
-
Hypoxia-mediated promotion of glucose metabolism in non-small cell lung cancer correlates with activation of the EZH2/FBXL7/PFKFB4 axis
Cell Death & Disease (2023)
-
Functional characterization of FBXL7 as a novel player in human cancers
Cell Death Discovery (2022)
-
Identification of a novel Calpain-2-SRC feed-back loop as necessity for β-Catenin accumulation and signaling activation in hepatocellular carcinoma
Oncogene (2022)