Abstract
Merkel cell carcinoma (MCC)—a neuroendocrine cancer of the skin—is caused by the integration of Merkel cell polyomavirus and persistent expression of large T antigen and small T antigen. We report that small T antigen in complex with MYCL and the EP400 complex activates the expression of LSD1 (KDM1A), RCOR2 and INSM1 to repress gene expression by the lineage transcription factor ATOH1. LSD1 inhibition reduces the growth of MCC in vitro and in vivo. Through a forward-genetics CRISPR–Cas9 screen, we identified an antagonistic relationship between LSD1 and the non-canonical BAF (ncBAF) chromatin remodelling complex. Changes in gene expression and chromatin accessibility caused by LSD1 inhibition were partially rescued by BRD9 inhibition, revealing that LSD1 and ncBAF antagonistically regulate an overlapping set of genes. Our work provides mechanistic insight into the dependence of MCC on LSD1 and a tumour suppressor role for ncBAF in cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
ATAC-Seq, ChIP-Seq and RNA-Seq data that support the findings of this study have been deposited in the Gene Expression Omnibus under accession codes GSE124856, GSE124857, GSE124861, GSE124864 and GSE140505. The mass spectrometry data are available at the ProteomeXchange Consortium under accession code PXD012516. All other data supporting the findings and computer codes implemented in this study are available from the corresponding author on reasonable request. Source data for Figs. 1–8 and Extended Data Figs. 1–3 and 7 are presented with the paper.
Change history
15 May 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41556-020-0533-9
References
Shuda, M. et al. T antigen mutations are a human tumor-specific signature for Merkel cell polyomavirus. Proc. Natl Acad. Sci. USA 105, 16272–16277 (2008).
Shuda, M., Kwun, H. J., Feng, H., Chang, Y. & Moore, P. S. Human Merkel cell polyomavirus small T antigen is an oncoprotein targeting the 4E-BP1 translation regulator. J. Clin. Invest. 121, 3623–3634 (2011).
Harms, P. W. et al. The distinctive mutational spectra of polyomavirus-negative Merkel cell carcinoma. Cancer Res. 75, 3720–3727 (2015).
Moll, I., Kuhn, C. & Moll, R. Cytokeratin 20 is a general marker of cutaneous Merkel cells while certain neuronal proteins are absent. J. Invest. Dermatol. 104, 910–915 (1995).
Cheng, J. et al. Merkel cell polyomavirus recruits MYCL to the EP400 complex to promote oncogenesis. PLoS Pathog. 13, e1006668 (2017).
Park, D. E. et al. Dual inhibition of MDM2 and MDM4 in virus-positive Merkel cell carcinoma enhances the p53 response. Proc. Natl Acad. Sci. USA 116, 1027–1032 (2018).
Shi, Y. et al. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119, 941–953 (2004).
Laurent, B. et al. A specific LSD1/KDM1A isoform regulates neuronal differentiation through H3K9 demethylation. Mol. Cell 57, 957–970 (2015).
Saleque, S., Kim, J., Rooke, H. M. & Orkin, S. H. Epigenetic regulation of hematopoietic differentiation by Gfi-1 and Gfi-1b is mediated by the cofactors CoREST and LSD1. Mol. Cell 27, 562–572 (2007).
Takagi, S. et al. LSD1 inhibitor T-3775440 inhibits SCLC cell proliferation by disrupting LSD1 interactions with SNAG domain proteins INSM1 and GFI1B. Cancer Res. 77, 4652–4662 (2017).
McGrath, J. P. et al. Pharmacological inhibition of the histone lysine demethylase KDM1A suppresses the growth of multiple acute myeloid leukemia subtypes. Cancer Res. 76, 1975–1988 (2016).
Mohammad, H. P. et al. A DNA hypomethylation signature predicts antitumor activity of LSD1 inhibitors in SCLC. Cancer Cell 28, 57–69 (2015).
Sugino, N. et al. A novel LSD1 inhibitor NCD38 ameliorates MDS-related leukemia with complex karyotype by attenuating leukemia programs via activating super-enhancers. Leukemia 31, 2303–2314 (2017).
Lee, C. et al. Lsd1 as a therapeutic target in Gfi1-activated medulloblastoma. Nat. Commun. 10, 332 (2019).
Kadoch, C. et al. Proteomic and bioinformatic analysis of mammalian SWI/SNF complexes identifies extensive roles in human malignancy. Nat. Genet. 45, 592–601 (2013).
Mashtalir, N. et al. Modular organization and assembly of SWI/SNF family chromatin remodeling complexes. Cell 175, 1272–1288.e20 (2018).
Alpsoy, A. & Dykhuizen, E. C. Glioma tumor suppressor candidate region gene 1 (GLTSCR1) and its paralog GLTSCR1-like form SWI/SNF chromatin remodeling subcomplexes. J. Biol. Chem. 293, 3892–3903 (2018).
Michel, B. C. et al. A non-canonical SWI/SNF complex is a synthetic lethal target in cancers driven by BAF complex perturbation. Nat. Cell Biol. 20, 1410–1420 (2018).
Inoue, D. et al. Spliceosomal disruption of the non-canonical BAF complex in cancer. Nature 574, 432–436 (2019).
Yang, P. et al. RCOR2 is a subunit of the LSD1 complex that regulates ESC property and substitutes for SOX2 in reprogramming somatic cells to pluripotency. Stem Cells 29, 791–801 (2011).
Rush, P. S. et al. Insulinoma-associated 1: a novel nuclear marker in Merkel cell carcinoma (cutaneous neuroendocrine carcinoma). J. Cutan. Pathol. 45, 129–135 (2018).
Florens, L. & Washburn, M. P. Proteomic analysis by multidimensional protein identification technology. Methods Mol. Biol. 328, 159–175 (2006).
Paoletti, A. C. et al. Quantitative proteomic analysis of distinct mammalian Mediator complexes using normalized spectral abundance factors. Proc. Natl Acad. Sci. USA 103, 18928–18933 (2006).
Berrios, C. et al. Merkel cell polyomavirus small T antigen promotes pro-glycolytic metabolic perturbations required for transformation. PLoS Pathog. 12, e1006020 (2016).
Sharma, R. et al. Bmp signaling maintains a mesoderm progenitor cell state in the mouse tailbud. Development 144, 2982–2993 (2017).
Wang, S. et al. Target analysis by integration of transcriptome and ChIP-Seq data with BETA. Nat. Protoc. 8, 2502–2515 (2013).
Zhu, L. J. et al. ChIPpeakAnno: a Bioconductor package to annotate ChIP-Seq and ChIP-chip data. BMC Bioinformatics 11, 237 (2010).
Wright, M. C. et al. Unipotent, Atoh1+ progenitors maintain the Merkel cell population in embryonic and adult mice. J. Cell Biol. 208, 367–379 (2015).
Paulo, J. & Gygi, S. P. Isobaric tag-based protein profiling of a nicotine-treated alpha7 nicotinic receptor-null human haploid cell line. Proteomics 18, e1700475 (2018).
Fagerberg, L. et al. Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol. Cell. Proteomics 13, 397–406 (2014).
Dhanjal, J. K., Radhakrishnan, N. & Sundar, D. Identifying synthetic lethal targets using CRISPR/Cas9 system. Methods 131, 66–73 (2017).
Lee, J.-E. et al. H3K4 mono- and di-methyltransferase MLL4 is required for enhancer activation during cell differentiation. eLife 2, e01503 (2013).
Remillard, D. et al. Degradation of the BAF complex factor BRD9 by heterobifunctional ligands. Angew. Chem. 56, 5738–5743 (2017).
Huelsken, J., Vogel, R., Erdmann, B., Cotsarelis, G. & Birchmeier, W. β-catenin controls hair follicle morphogenesis and stem cell differentiation in the skin. Cell 105, 533–545 (2001).
Guha, U. et al. Target-derived BMP signaling limits sensory neuron number and the extent of peripheral innervation in vivo. Development 131, 1175–1186 (2004).
Tu, W. B. et al. MYC interacts with the G9a histone methyltransferase to drive transcriptional repression and tumorigenesis. Cancer Cell 34, 579–595.e8 (2018).
Varlakhanova, N., Cotterman, R., Bradnam, K., Korf, I. & Knoepfler, P. S. Myc and Miz-1 have coordinate genomic functions including targeting Hox genes in human embryonic stem cells. Epigenetics Chromatin 4, 20 (2011).
Brien, G. L. et al. Targeted degradation of BRD9 reverses oncogenic gene expression in synovial sarcoma. eLife 7, e41305 (2018).
Sheng, W. et al. LSD1 ablation stimulates anti-tumor immunity and enables checkpoint blockade. Cell 174, 549–563.e19 (2018).
Nghiem, P. T. et al. PD-1 blockade with pembrolizumab in advanced Merkel-cell carcinoma. N. Engl. J. Med. 374, 2542–2552 (2016).
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-Seq data with DESeq2. Genome Biol. 15, 550 (2014).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Huang, D. Wa. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Zhang, W. & Wang, B. MAGeCKFlute: Integrative analysis pipeline for pooled CRISPR functional genetic screens. R package version 1.1.8 (2018).
Ianevski, A., He, L., Aittokallio, T. & Tang, J. SynergyFinder: a web application for analyzing drug combination dose–response matrix data. Bioinformatics 33, 2413–2415 (2017).
Yu, G., Wang, L.-G. G., Han, Y. & He, Q.-Y. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Carey, M. F., Peterson, C. L. & Smale, S. T. Dignam and Roeder nuclear extract preparation. Cold Spring Harb. Protoc. 2009, pdb.prot5330 (2009).
Cheng, J., Rozenblatt-Rosen, O., Paulson, K. G., Nghiem, P. & DeCaprio, J. A. Merkel cell polyomavirus large T antigen has growth-promoting and inhibitory activities. J. Virol. 87, 6118–6126 (2013).
Xu, T. et al. ProLuCID: an improved SEQUEST-like algorithm with enhanced sensitivity and specificity. J. Proteomics 129, 16–24 (2015).
Schmidt, D. et al. ChIP-Seq: using high-throughput sequencing to discover protein–DNA interactions. Methods 48, 240–248 (2009).
Mertins, P. et al. Reproducible workflow for multiplexed deep-scale proteome and phosphoproteome analysis of tumor tissues by liquid chromatography–mass spectrometry. Nat. Protoc. 13, 1632–1661 (2018).
Corces, M. R. et al. An improved ATAC-Seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Gaspar, J. M. NGmerge: merging paired-end reads via novel empirically-derived models of sequencing errors. BMC Bioinformatics 19, 536 (2018).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Yu, G., Wang, L.-G. G. & He, Q.-Y. Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Ross-Innes, C. S. et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481, 389–393 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. R137 (2008).
Dobin, A. et al. STAR: ultrafast universal RNA-Seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Supek, F., Bošnjak, M., Škunca, N. & Šmuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE 6, e21800 (2011).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Blighe, K., Rana, S. & Lewis, M. EnhancedVolcano: Publication-ready volcano plots with enhanced colouring and labeling. R package version 1.4.0 (2019).
Navarrete-Perea, J., Yu, Q., Gygi, S. P. & Paulo, J. A. Streamlined tandem mass tag (SL-TMT) protocol: an efficient strategy for quantitative (phospho)proteome profiling using tandem mass tag-synchronous precursor selection-MS3. J. Proteome Res. 17, 2226–2236 (2018).
Ting, L., Rad, R., Gygi, S. P. & Haas, W. MS3 eliminates ratio distortion in isobaric multiplexed quantitative proteomics. Nat. Methods 8, 937–940 (2011).
Elias, J. E. & Gygi, S. P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Acknowledgements
We are grateful to C. Kadoch (DFCI) and members of the Kadoch laboratory for helpful discussions relating to mSWI/SNF and specifically ncBAF complex biology and experimental approaches, Y. Shi (Boston Children’s Hospital) for providing valuable insights on LSD1, X. S. Liu (DFCI) for consultation about the CRISPR–Cas9 screen analysis, and S. Gygi (Harvard Medical School), J. Qi (DFCI) and M. Brown (DFCI) for sharing reagents and equipment. This work was supported in part by the US Public Health Service (grant F31CA213464 to D.E.P. and grants R35CA232128, R01CA63113, R01CA173023 and P01CA203655 to J.A.D.). J.A.D. received research funding from Constellation Pharmaceuticals. S.K.S., L.F. and M.P.W. were supported by the Stowers Institute for Medical Research. J.A.P. was supported by NIH NIGMS grant R01GM132129.
Author information
Authors and Affiliations
Contributions
D.E.P. and J.A.D. conceived of the study. D.E.P. and J.A.D. developed the methodology. D.E.P., M.Y.L. and L.F. developed the software. D.E.P. and J.C. validated the results. D.E.P., M.Y.L., L.F., C.C., J.P.M. and P.C.G. performed the formal analysis. D.E.P., J.C., J.P.M., M.Y.L., S.K.S. and M.L.T. performed the investigation. J.A.P., P.C.G., M.P.W., P.T. and J.A.D. provided resources. D.E.P. and J.A.D. wrote the original draft of the manuscript. D.E.P., J.C., J.P.M., M.Y.L., L.F. and J.A.D. reviewed and edited the manuscript. D.E.P. and M.Y.L. visualized the results. P.T., P.C.G., M.P.W. and J.A.D. supervised the study. P.T. and J.A.D. acquired funding.
Corresponding author
Ethics declarations
Competing interests
J.P.M. and P.T. are employees of Constellation Pharmaceuticals. J.A.D. received research funding from Constellation Pharmaceuticals. J.A.D. has served as a consultant to Merck & Co. and EMD Serono.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Merkel small T antigen transactivates LSD1 complex components.
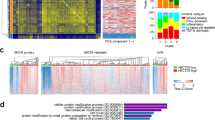
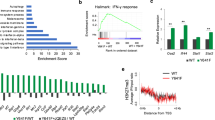
a, Integrated ChIP- and RNA-seq analysis of the MCV ST target genes predicts that MCV ST forms an activator complex. b, RNA-seq results show that EP400 depletion in MKL-1 leads to a reduction in mRNA levels of RCOR2, LSD1, INSM1, and additional components of the LSD1 complex. DESeq2 normalized counts were plotted. Differentially expressed genes were found by comparing each condition with DESeq2 and p-values were adjusted by Benjamini-Hochberg. c, Two independent shRNAs against EP400 decrease levels of EP400 and RCOR2 levels but increase levels of DLL1, HES1, LGR6, KRT17, KRT9, and CDH1 in MKL-1 cells. The RT-qPCR signals were normalized to each uninduced sample and the geomean of 36B (RPLP0) and 18 s rRNA. d, Depletion of EP400, MYCL, or MCV ST by shRNA leads to increased levels of genes involved in critical cancer and differentiation pathways in MKL-1 and WaGa MCC cell lines (n = 3 independent biological replicates used in each condition). The enrichment test was performed on hypergeometric distribution and the p-values were adjusted by FDR (Supplementary Tables 1–3). e, RNA-seq of MKL-1 cells expressing three independent shRNAs targeting EP400 or ST in MKL-1 and WaGa cells was performed. Mean expression was plotted against log fold change. f, The LSD1 and RCOR2 levels were significantly higher in PHEL ( + MCV LT and wild-type ST) and PH2L ( + MCV LT and the ST mutant E86S-E87S unable to bind the EP400) than in PHL (-MCV T antigens). The fold changes in the western blot signals from four replicate experiments were averaged. Data are shown as mean of n = 4 ± SD; two-sided t-test, *P < 0.05; **<0.005. g, A representative blot for Extended Data Fig 1g is shown. The experiment was performed four times. See Unprocessed Gels Extended Data Fig. 1.
Extended Data Fig. 2 LSD1 inhibition reduces the growth of virus-positive MCC cell lines.
a-g, Virus-positive (a-d), but not virus-negative (e-g), MCC cell lines are sensitive to two independent LSD1 inhibitors (GSK2879552 and CPI-670242) in a dose-dependent manner. Relative viability was measured at 12 days of treatment by the CellTiter-Glo assay. Data are shown as mean ± SD and reflect three biological replicates.
Extended Data Fig. 3 Integrative ChIP-seq and RNA-seq LSD1 targetome analysis reveal that LSD1 regulates neuronal differentiation pathways in MCC.
a, Differential gene expression analysis was performed using DESeq2 and p-values were adjusted by Benjamini-Hochberg. b, c, RT-qPCR (b) and western blot (c) assessment of LSD1 target genes in MKL-1 cells of LSD1 inhibition (GSK-LSD1, 1 μM) in triplicate. For b, the signals were normalized to untreated samples and RPLP0 in each sample. See Unprocessed Gels Extended Data Fig. 3. d, Cells were treated with GSK-LSD1 (0.1 μM) for three days. Data are shown as mean of n = 3 ± SD; two-sided t-test, *P < 0.05; **<0.005. e, Multiplexed Isobaric Tag-Based Profiling of MKL-1 treated with GSK-LSD1 for eight days displays global changes in the proteome. The Venn diagrams show the numbers of genes identified in the RNA- and ChIP-seq-based targetome analysis (Fig. 3; LSD1 MCC targets: upregulated following LSD1i with LSD1 ChIP peaks; RNA down: downregulated following LSD1i) and the proteomics experiment (Supplementary Tables 5 and 11; Proteomics up: upregulated following LSD1i; Proteomics down: downregulated following LSD1i). f, The targetome analysis was performed by integrating ChIP-seq of LSD1 and RNA-seq of the virus-positive MCC cell lines in three independent biological replicates (Supplementary Table 5). Selected GOTERM biological processes (BP) and Cell compartments (CC) –log10 of Bonferroni adjusted p-values are shown. g, The LSD1 IP, followed by western blotting, indicates that LSD1 has reduced binding to INSM1 following LSD1 inhibition. The experiment was performed three times. # non-specific band. See Unprocessed Gels Extended Data Fig. 3. h, IP-western blotting shows INSM1 reduced interaction with LSD1. MKL-1 cells were treated with GSK-LSD1 (0.1 μM) for three days and processed for IP. The experiment was performed at least three times. See Unprocessed Gels Extended Data Fig. 3. i, RT-qPCR of MCC lines and primary human cells (HFF) were performed in triplicate60. Data are shown as mean of n = 3 ± SD; two-sided t-test, ***P < 0.0005. # - below detection level.
Extended Data Fig. 4 ChIP-seq of LSD1, RCOR2, and ATOH1 indicates shared DNA occupancy.
a, Integrated ChIP- and RNA-seq analysis35 reveals that genes perturbed by LSD1 inhibition are direct LSD1 targets. b, Tag heat maps of two replicates of LSD1 (LSD1_1 and LSD1_2), RCOR2 (RCOR2_1 and RCOR2_2), and ATOH1 (ATOH1_1 and ATOH1_2) ChIP-seq show that the factors bind to common regions close to the transcription start sites (TSSs + /- 4,000 bp). c, The heatmap displays correlations among LSD1, RCOR2, and ATOH1 ChIP-seq peaks. d, The SeqPos motif tool36 was used to determine the LSD1 binding motifs in MKL-1. The binding motifs of ATOH1, as well as OLIG2 and ASCL2, were enriched in the RCOR2 (RCOR2_1) ChIP-seq. e, GOTERM biological process pathway analysis of the two independent replicates of LSD1, RCOR2, and ATOH1 ChIP-seq revealed that the factors regulate genes involved in neuronal functions and developmental signaling. The enrichment test was performed on hypergeometric distribution and the p-values were adjusted by FDR.
Extended Data Fig. 5 Multiplexed Isobaric Tag-Based Profiling of MCC-derived xenografts reveals that LSD1 inhibition perturbs neuronal gene expression in human and mouse tissues.
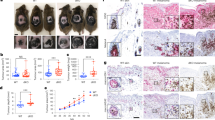
a-b, The multiplexed Isobaric Tag-Based Profiling of the MKL-1 virus-positive MCC cell line treated with GSK-LSD1 for eight days identified genes that are differentially expressed during LSD1 inhibition. n = 5 independent biological replicates were used for analysis. Selected GOTERM biological process and cellular compartment terms are shown for the upregulated (a) and downregulated (b) proteins. c, Eight mice were injected with MKL-1 or WaGa MCC cells, and when the tumor size reached 150 mm3, two of each four mice for each MCC model were treated with CPI-670242 (40 mg/kg) orally once a week for 22 days. The tumors were harvested for the TMT-10 plex isobaric tag-labeling quantitative mass spectrometry experiment. c, The heatmap shows the relative abundance of 30 most upregulated and 30 most downregulated mouse proteins after the treatment.
Extended Data Fig. 6 Multiplexed Isobaric Tag-Based Profiling of MCC-derived xenografts profiles global changes in tumor proteomes.
The volcano plots display global protein expression changes of MKL-1 human proteins (a), WaGa human proteins (b), MKL-1 mouse proteins (c), and WaGa mouse proteins (d) with or without the LSD1 inhibitor. Tukey’s Honest Significance Test was performed post-hoc on-peptide level linear models for each protein and the p-values were adjusted by Benjamini-Hochberg. Each protein is plotted based on its log2 fold change against -log10 of adjusted p-values. The five most upregulated and five most downregulated proteins in each plot are labeled.
Extended Data Fig. 7 CRISPR screen shows that LSD1 inhibition creates positive pressure for deleting ncBAF complex components.
a, Gene ranks based on the differences of the beta (selection) scores between the treatment (GSK-LSD1, 20 days, IC30–1.5 nM) and control (DMSO 20 days) screens show the positively and negatively selected mSWI/SNF components genes including the ncBAF complex components BRD9, GLTSCR1, SMARCA2, SMARCD1, and SMARCC1 (Supplementary Tables 18–20). The rank list contains all the previously reported mSWI/SNF components. b, BRD9 binds to SMARCA4 (BRG1) and GLTSCR1 (BICRA) in MKL-1. IP using a BRD9 antibody followed by western blotting was performed to determine interactions among BRD9, GLTSCR1, and SMARCA4. The experiment was performed at least three times. See Unprocessed Gels Extended Data Fig. 7. c, BRD9 degradation restores the loss of cell viability caused by LSD1 inhibition in MCC. MKL-1 cells were treated with varying doses of the dBRD9 and LSD1 inhibitor (GSK-LSD1) for six days. The XTT assay measured relative cell viability. d, dBRD9 degrades BRD9 efficiently. MKL-1 cells were treated with GSK-LSD1 (0.1 μM), dBRD9 (0.1 μM), or both for three days and harvested for western blotting. The experiment was performed at least three times. See Unprocessed Gels Extended Data Fig. 7. e, The PCA plot shows that the degradation of BRD9 by dBRD9 partially rescues the global gene expression changes caused by LSD1 inhibition. n = 2 independent biological replicates were used for analysis. f, BRD9 depletion by shRNAs rescues gene expression changes caused by LSD1 depletion. MKL-1 cells were transduced with an LSD1-targeting shRNA either with a control shRNA or two distinct BRD9-targeting shRNAs for six days and harvested for western blotting. The experiment was performed at least three times. See Unprocessed Gels Extended Data Fig. 7.
Extended Data Fig. 8 Heatmaps of gene expression changes following LSD1 and BRD9 inhibition indicate that LSD1 and BRD9 regulate an overlapping set of genes in an antagonistic manner.
a, b, RNA-seq was performed with n = 2 biologically independent replicates of MKL-1 cells treated with DMSO, GSK-LSD1 (LSD1i, 0.1 μM), dBRD9 (0.1 μM), or both GSK-LSD1 and dBRD9 for six days. a, Top 50 most differentially expressed genes between LSD1i and LSD1i + dBRD9 are shown. Differential gene expression analysis was performed using DESeq2 and p-values were adjusted by Benjamini-Hochberg. b, All differentially expressed genes (3392 genes; FDR < 0.05) across the conditions are shown.
Extended Data Fig. 9 ATAC-seq of MKL-1 cells treated with LSD1 and BRD9 inhibitors suggests that BRD9 is required to de-repress a subset of LSD1 target genes.
a, Tag heat maps of ATAC-seq peaks show open chromatin regions localize close to the transcription start sites (TSSs, +/- 2000 bp). Combined peaks from two replicates of ATAC-seq of MKL-1 cells treated with DMSO (Control), dBRD9 (0.1 µM), GSK-LSD1 (LSD1i, 0.1 µM), or both dBRD9 and LSD1i (LSD1i_dBRD9) are shown. b–d, The differentially enriched ATAC-seq peaks between Control vs. LSD1i (b), Control vs. dBRD9 (c), and LSD1i vs. LSD1i + dBRD9 (d) are shown with the peak abundance (log concentration) and log fold change in peak scores. The differentially enriched peaks were called using the Diffbind R package with the Wald test and FDR p-value correction. e, ATAC-seq peaks in the promoters of LSD1 target genes ID1, SYT4, and CALB2 with the mentioned conditions are shown. f, Model: Merkel cell polyomavirus activates LSD1-mediated blockade of non-canonical BAF to regulate transformation and tumorigenesis.
Supplementary information
Supplementary Tables
Supplementary Tables 1–28.
Source data
Source Data Fig. 1
Source data
Source Data Fig. 1
Unprocessed western blots for Fig. 1h and 1i
Source Data Fig. 2
Source data
Source Data Fig. 2
Unprocessed western blots for Fig. 2d
Source Data Fig. 3
Source data
Source Data Fig. 3
Unprocessed western blots for Fig. 3e, Fig. 3f, Fig. 3g,h
Source Data Fig. 4
Source data
Source Data Fig. 5
Source data
Source Data Fig. 6
Source data
Source Data Fig. 7
Source data
Source Data Fig. 8
Source data
Source Data Extended Data Fig. 1
Source data
Source Data Extended Data Fig. 1
Unprocessed gels for ED Fig. 1g
Source Data Extended Data Fig. 2
Source data
Source Data Extended Data Fig. 3
Source data
Source Data Extended Data Fig. 3
Unprocessed gels for ED Fig. 3c,g,h
Source Data Extended Data Fig. 7
Unprocessed gels for ED Fig. 7b,d,f
Rights and permissions
About this article
Cite this article
Park, D.E., Cheng, J., McGrath, J.P. et al. Merkel cell polyomavirus activates LSD1-mediated blockade of non-canonical BAF to regulate transformation and tumorigenesis. Nat Cell Biol 22, 603–615 (2020). https://doi.org/10.1038/s41556-020-0503-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-0503-2
This article is cited by
-
3D genome mapping identifies subgroup-specific chromosome conformations and tumor-dependency genes in ependymoma
Nature Communications (2023)
-
CRISPR/Cas9 application in cancer therapy: a pioneering genome editing tool
Cellular & Molecular Biology Letters (2022)
-
Identification of a chromatin regulator signature and potential candidate drugs for bladder cancer
Hereditas (2022)
-
DNA-methylation patterns imply a common cellular origin of virus- and UV-associated Merkel cell carcinoma
Oncogene (2022)
-
Proteomic characterization of post-translational modifications in drug discovery
Acta Pharmacologica Sinica (2022)