Abstract
Dynein is a microtubule motor that transports many different cargos in various cell types and contexts. How dynein is regulated to perform these activities with spatial and temporal precision remains unclear. Human dynein is regulated by autoinhibition, whereby intermolecular contacts limit motor activity. Whether this mechanism is conserved throughout evolution, whether it can be affected by extrinsic factors, and its role in regulating dynein function remain unclear. Here, we use a combination of negative stain electron microscopy, single-molecule assays, genetic, and cell biological techniques to show that autoinhibition is conserved in budding yeast, and plays a key role in coordinating in vivo dynein function. Moreover, we find that the Lissencephaly-related protein, LIS1 (Pac1 in yeast), plays an important role in regulating dynein autoinhibition. Our studies demonstrate that, rather than inhibiting dynein motility, Pac1/LIS1 promotes dynein activity by stabilizing the uninhibited conformation, which ensures appropriate dynein localization and activity in cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All of the yeast strains, datasets and raw video files that were generated during and/or analysed during the current study are available from the corresponding author upon reasonable request.
References
Schlager, M. A., Hoang, H. T., Urnavicius, L., Bullock, S. L. & Carter, A. P. In vitro reconstitution of a highly processive recombinant human dynein complex. EMBO J. 33, 1855–1868 (2014).
McKenney, R. J., Huynh, W., Tanenbaum, M. E., Bhabha, G. & Vale, R. D. Activation of cytoplasmic dynein motility by dynactin-cargo adapter complexes. Science 345, 337–341 (2014).
McKenney, R. J., Huynh, W., Vale, R. D. & Sirajuddin, M. Tyrosination of α-tubulin controls the initiation of processive dynein-dynactin motility. EMBO J. 35, 1175–1185 (2016).
Zhang, K. et al. Cryo-EM reveals how human cytoplasmic dynein is auto-inhibited and activated. Cell 169, 1303–1314 (2017).
Amos, L. A. Brain dynein crossbridges microtubules into bundles. J. Cell Sci. 93, 19–28 (1989).
Toropova, K., Mladenov, M. & Roberts, A. J. Intraflagellar transport dynein is autoinhibited by trapping of its mechanical and track-binding elements. Nat. Struct. Mol. Biol. 24, 461–468 (2017).
Jordan, M. A., Diener, D. R., Stepanek, L. & Pigino, G. The cryo-EM structure of intraflagellar transport trains reveals how dynein is inactivated to ensure unidirectional anterograde movement in cilia. Nat. Cell Biol. 20, 1250–1255 (2018).
Torisawa, T. et al. Autoinhibition and cooperative activation mechanisms of cytoplasmic dynein. Nat. Cell Biol. 16, 1118–1124 (2014).
Reck-Peterson, S. L. et al. Single-molecule analysis of dynein processivity and stepping behavior. Cell 126, 335–348 (2006).
Kardon, J. R., Reck-Peterson, S. L. & Vale, R. D. Regulation of the processivity and intracellular localization of Saccharomyces cerevisiae dynein by dynactin. Proc. Natl Acad. Sci. USA 106, 5669–5674 (2009).
Sasaki, S. et al. A LIS1/NUDEL/cytoplasmic dynein heavy chain complex in the developing and adult nervous system. Neuron 28, 681–696 (2000).
Wynshaw-Boris, A. & Gambello, M. J. LIS1 and dynein motor function in neuronal migration and development. Genes Dev. 15, 639–651 (2001).
Moughamian, A. J., Osborn, G. E., Lazarus, J. E., Maday, S. & Holzbaur, E. L. Ordered recruitment of dynactin to the microtubule plus-end is required for efficient initiation of retrograde axonal transport. J. Neurosci. 33, 13190–13203 (2013).
Raaijmakers, J. A., Tanenbaum, M. E. & Medema, R. H. Systematic dissection of dynein regulators in mitosis. J. Cell Biol. 201, 201–215 (2013).
Coquelle, F. M. et al. LIS1, CLIP-170’s key to the dynein/dynactin pathway. Mol. Cell. Biol. 22, 3089–3102 (2002).
Tsai, J. W., Bremner, K. H. & Vallee, R. B. Dual subcellular roles for LIS1 and dynein in radial neuronal migration in live brain tissue. Nat. Neurosci. 10, 970–979 (2007).
Tsai, J. W., Chen, Y., Kriegstein, A. R. & Vallee, R. B. LIS1 RNA interference blocks neural stem cell division, morphogenesis, and motility at multiple stages. J. Cell Biol. 170, 935–945 (2005).
Yi, J. Y. et al. High-resolution imaging reveals indirect coordination of opposite motors and a role for LIS1 in high-load axonal transport. J. Cell Biol. 195, 193–201 (2011).
Chapman, D. E. et al. Regulation of in vivo dynein force production by CDK5 and 14-3-3ε and KIAA0528. Nat. Commun. 10, 228 (2019).
Reddy, B. J. et al. Load-induced enhancement of dynein force production by LIS1–NudE in vivo and in vitro. Nat. Commun. 7, 12259 (2016).
McKenney, R. J., Vershinin, M., Kunwar, A., Vallee, R. B. & Gross, S. P. LIS1 and NudE induce a persistent dynein force-producing state. Cell 141, 304–314 (2010).
Yamada, M. et al. LIS1 and NDEL1 coordinate the plus-end-directed transport of cytoplasmic dynein. EMBO J. 27, 2471–2483 (2008).
Baumbach, J. et al. Lissencephaly-1 is a context-dependent regulator of the human dynein complex. eLife 6, e21768 (2017).
Gutierrez, P. A., Ackermann, B. E., Vershinin, M. & McKenney, R. J. Differential effects of the dynein-regulatory factor Lissencephaly-1 on processive dynein-dynactin motility. J. Biol. Chem. 292, 12245–12255 (2017).
Jha, R., Roostalu, J., Cade, N. I., Trokter, M. & Surrey, T. Combinatorial regulation of the balance between dynein microtubule end accumulation and initiation of directed motility. EMBO J. 36, 3387–3404 (2017).
Huang, J., Roberts, A. J., Leschziner, A. E. & Reck-Peterson, S. L. Lis1 acts as a “clutch” between the ATPase and microtubule-binding domains of the dynein motor. Cell 150, 975–986 (2012).
Toropova, K. et al. Lis1 regulates dynein by sterically blocking its mechanochemical cycle. eLife 3, e03372 (2014).
DeSantis, M. E. et al. Lis1 has two opposing modes of regulating cytoplasmic dynein. Cell 170, 1197–1208 (2017).
Markus, S. M. & Lee, W. L. Regulated offloading of cytoplasmic dynein from microtubule plus ends to the cortex. Dev. Cell 20, 639–651 (2011).
Dix, C. I. et al. Lissencephaly-1 promotes the recruitment of dynein and dynactin to transported mRNAs. J. Cell Biol. 202, 479–494 (2013).
Wang, S. et al. Nudel/NudE and Lis1 promote dynein and dynactin interaction in the context of spindle morphogenesis. Mol. Biol. Cell 24, 3522–3533 (2013).
Marzo, M. G. et al. Molecular basis for dyneinopathies reveals insight into dynein regulation and dysfunction. eLife 8, e47246 (2019).
Markus, S. M. & Lee, W. L. Microtubule-dependent path to the cell cortex for cytoplasmic dynein in mitotic spindle orientation. Bioarchitecture 1, 209–215 (2011).
Schmidt, H., Gleave, E. S. & Carter, A. P. Insights into dynein motor domain function from a 3.3 Å crystal structure. Nat. Struct. Mol. Biol. 19, 492–497 (2012).
Carter, A. P., Cho, C., Jin, L. & Vale, R. D. Crystal structure of the dynein motor domain. Science 331, 1159–1165 (2011).
Schmidt, H., Zalyte, R., Urnavicius, L. & Carter, A. P. Structure of human cytoplasmic dynein-2 primed for its power stroke. Nature 518, 435–438 (2015).
Gibbons, I. R. et al. The affinity of the dynein microtubule-binding domain is modulated by the conformation of its coiled-coil stalk. J. Biol. Chem. 280, 23960–23965 (2005).
Kon, T. et al. Helix sliding in the stalk coiled coil of dynein couples ATPase and microtubule binding. Nat. Struct. Mol. Biol. 16, 325–333 (2009).
Rao, L., Berger, F., Nicholas, M. P. & Gennerich, A. Molecular mechanism of cytoplasmic dynein tension sensing. Nat. Commun. 10, 3332 (2019).
Gennerich, A., Carter, A. P., Reck-Peterson, S. L. & Vale, R. D. Force-induced bidirectional stepping of cytoplasmic dynein. Cell 131, 952–965 (2007).
Moore, J. K., Li, J. & Cooper, J. A. Dynactin function in mitotic spindle positioning. Traffic 9, 510–527 (2008).
Markus, S. M., Punch, J. J. & Lee, W. L. Motor- and tail-dependent targeting of dynein to microtubule plus ends and the cell cortex. Curr. Biol. 19, 196–205 (2009).
Lammers, L. G. & Markus, S. M. The dynein cortical anchor Num1 activates dynein motility by relieving Pac1/LIS1-mediated inhibition. J. Cell Biol. 211, 309–322 (2015).
Lee, W. L., Oberle, J. R. & Cooper, J. A. The role of the lissencephaly protein Pac1 during nuclear migration in budding yeast. J. Cell Biol. 160, 355–364 (2003).
Yin, H., Pruyne, D., Huffaker, T. C. & Bretscher, A. Myosin V orientates the mitotic spindle in yeast. Nature 406, 1013–1015 (2000).
Hwang, E., Kusch, J., Barral, Y. & Huffaker, T. C. Spindle orientation in Saccharomyces cerevisiae depends on the transport of microtubule ends along polarized actin cables. J. Cell Biol. 161, 483–488 (2003).
Liakopoulos, D., Kusch, J., Grava, S., Vogel, J. & Barral, Y. Asymmetric loading of Kar9 onto spindle poles and microtubules ensures proper spindle alignment. Cell 112, 561–574 (2003).
Ecklund, K. H. et al. She1 affects dynein through direct interactions with the microtubule and the dynein microtubule-binding domain. Nat. Commun. 8, 2151 (2017).
Markus, S. M. et al. Quantitative analysis of Pac1/LIS1-mediated dynein targeting: implications for regulation of dynein activity in budding yeast. Cytoskeleton 68, 157–174 (2011).
Sheeman, B. et al. Determinants of S. cerevisiae dynein localization and activation: implications for the mechanism of spindle positioning. Curr. Biol. 13, 364–372 (2003).
Li, J., Lee, W. L. & Cooper, J. A. NudEL targets dynein to microtubule ends through LIS1. Nat. Cell. Biol. 7, 686–690 (2005).
Elshenawy, M. M. et al. Lis1 activates dynein motility by pairing it with dynactin. Nat. Cell Biol. https://doi.org/10.1038/s41556-020-0501-4 (2020).
Htet, Z. M. et al. Lis1 promotes the formation of activated cytoplasmic dynein-1 complexes. Nat. Cell Biol. https://doi.org/10.1038/s41556-020-0506-z (2020).
Urnavicius, L. et al. Cryo-EM shows how dynactin recruits two dyneins for faster movement. Nature 554, 202–206 (2018).
Qiu, R., Zhang, J. & Xiang, X. LIS1 regulates cargo-adapter-mediated activation of dynein by overcoming its autoinhibition in vivo. J. Cell Biol. 218, 3630–3646 (2019).
Curran, K. A. et al. Short synthetic terminators for improved heterologous gene expression in yeast. ACS Synth. Biol. 4, 824–832 (2015).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Poirier, K. et al. Mutations in TUBG1, DYNC1H1, KIF5C and KIF2A cause malformations of cortical development and microcephaly. Nat. Genet. 45, 639–647 (2013).
Scoto, M. et al. Novel mutations expand the clinical spectrum of DYNC1H1-associated spinal muscular atrophy. Neurology 84, 668–679 (2015).
Willemsen, M. H. et al. Mutations in DYNC1H1 cause severe intellectual disability with neuronal migration defects. J. Med. Genet. 49, 179–183 (2012).
Heil-Chapdelaine, R. A., Oberle, J. R. & Cooper, J. A. The cortical protein Num1p is essential for dynein-dependent interactions of microtubules with the cortex. J. Cell Biol. 151, 1337–1344 (2000).
Weissmann, F. et al. biGBac enables rapid gene assembly for the expression of large multisubunit protein complexes. Proc. Natl Acad. Sci. USA 113, E2564–E2569 (2016).
Mahamdeh, M., Simmert, S., Luchniak, A., Schaffer, E. & Howard, J. Label-free high-speed wide-field imaging of single microtubules using interference reflection microscopy. J. Microsc. 272, 60–66 (2018).
Thiede, C., Lakamper, S., Wessel, A. D., Kramer, S. & Schmidt, C. F. A chimeric kinesin-1 head/kinesin-5 tail motor switches between diffusive and processive motility. Biophys. J. 104, 432–441 (2013).
Stellwagen, E., Prantner, J. D. & Stellwagen, N. C. Do zwitterions contribute to the ionic strength of a solution? Anal. Biochem. 373, 407–409 (2008).
Fraley, C. & Raftery, A. E. Bayesian regularization for normal mixture estimation and model-based clustering. J. Classif. 24, 155–181 (2007).
Redwine, W. B. et al. Structural basis for microtubule binding and release by dynein. Science 337, 1532–1536 (2012).
Alushin, G. M. et al. High-resolution microtubule structures reveal the structural transitions in αβ-tubulin upon GTP hydrolysis. Cell 157, 1117–1129 (2014).
Imai, H. et al. Direct observation shows superposition and large scale flexibility within cytoplasmic dynein motors moving along microtubules. Nat. Commun. 6, 8179 (2015).
Can, S., Lacey, S., Gur, M., Carter, A. P. & Yildiz, A. Directionality of dynein is controlled by the angle and length of its stalk. Nature 566, 407–410 (2019).
Acknowledgements
We thank S. Reck-Peterson for the 8His-ZZ-Pac1-SNAP-expressing yeast strain, and members of the Markus and DeLuca laboratories for discussions. Electron microscopy was performed at the University of Colorado, Boulder EM Services Core Facility in the MCDB Department, with the technical assistance of facility staff. This work utilized the RMACC Summit supercomputer, which is supported by the National Science Foundation (awards ACI-1532235 and ACI-1532236), the University of Colorado Boulder and Colorado State University. The RMACC Summit supercomputer is a joint effort of the University of Colorado Boulder and Colorado State University. We also thank E. Osborne-Nishimura, D. King and S. Bowerman for their assistance with using software on Summit. This research was funded by the NIH/NIGMS (GM118492, to S.M.M.).
Author information
Authors and Affiliations
Contributions
M.G.M. and S.M.M. designed the study. M.G.M. performed and analysed most of the assays, with support from S.M.M. and J.M.G. Electron microscopy was performed by Garry Morgan at the University of Colorado Boulder Electron Microscopy facility. Single-particle analysis was performed by S.M.M. with assistance from the EM facility. Reagents were generated by S.M.M., M.G.M. and J.M.G.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Representative raw EM image and quantitation of conformational states.
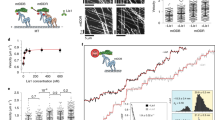
a, Representative EM image of negative stained yeast dynein complex (red arrow, phi particle conformation; green arrow, open, uninhibited conformation; yellow arrow, ambiguous). b, Quantitation of indicated conformational states from raw images (n = 435 particles).
Extended Data Fig. 2 Fluorescence intensity analysis of native and overexpressed single molecules of dynein.
Histogram of fluorescence intensity values for single molecules of motile dyneins, as indicated, along with accompanying Gaussian fits and modeled parameters (determined using the model-based clustering algorithm Mclust66). The percentages reflect the relative proportion of molecules that fall within each component (i.e., for mean 1, and mean 2). The two mean values for each likely represent single-labeled (mean 1) and dual-labeled (mean 2) dynein dimers, respectively. Red outlined region in panel D (“aggregates”), delineate particles with ~3-fold higher fluorescence intensity values than the single labeled complexes.
Extended Data Fig. 3 D2868K mutation has no effect on GST-dyneinMOTOR motility.
a, Schematic of the plasmid used to produce GST-dyneinMOTOR (wild-type and D2868K mutant). Restriction digest with ApaI (cuts within URA3 gene) targets the plasmid for homologous recombination into the ura3-1 locus as depicted. b, Cartoon representation of the minimal GST-dimerized dynein motor domain (amino acids 1219-4092 of the dynein heavy chain, Dyn1). (c and d) Plots depicting mean values (left) and all values (right) for velocity (c) and run length (d) of wild-type and D2868K GST-dyneinMOTOR, along with the standard error (n = 217 and 238 motors for each).
Extended Data Fig. 4 Uninhibited dynein mutant exhibits only moderate increase in microtubule landing activity.
a, Plots depicting relative microtubule landing rate of full-length wild-type (WT) and D2868K (DK) dynein, as measured from single molecule motility experiments (mean ± standard deviation; n = 554 wild-type motors from 1532 µm of microtubules, and 553 D2868K motors from 1177 µm of microtubules; 3 independent experiments were quantitated for each). Diamonds represent mean normalized values obtained from each independent replicate experiment. Briefly, equivalent concentrations of full length wild-type or D2868K dynein were added to imaging chambers (after taking into account relative differences in labeling efficiencies, as determined from fluorescent scans of protein gels), and the number of moving motors were quantitated. Statistical significance was determined using a two-tailed Welch’s t test. (b and c) Representative gel (b; Sypro Ruby-stained) and quantitation (c) of microtubule co-sedimentation assay with full-length wild-type (WT) and D2868K (DK) dynein done in the absence and presence of ATP (mean ± standard deviation; n = 2 independent experiments; diamonds represent values obtained from each replicate). Relative microtubule binding was determined by measuring background-corrected band intensities of each, and subtracting any non-specific microtubule-independent pelleting (as determined from experiment performed in the absence of microtubules).
Extended Data Fig. 5 Synthetic interactions between dynein mutants and Kar9.
(a–c) Serial dilutions of cells with indicated genotype were plated on rich media (YPA supplemented with 2% glucose) and grown at 30 °C for 2-4 days (a, extended incubation of plates shown in Fig. 4e were grown for 4 days; all others were incubated for 2 days). Note the severe growth defects in dyn1HL3 kar9∆ cells (in panel c), suggesting that dyneinHL3 is not active in cells. Note that similar results were obtained from 2 independent replicates.
Extended Data Fig. 6 Pac1-microtubule binding behavior, and its contribution to dynein activity.
(a and b) Representative images (a) and intensity scatter plots (b; bars depict mean ± standard deviation) of microtubule-bound Pac1 in different buffers. Pac1-SNAP647 diluted in motility buffer (50 nM, dimer concentration) with indicated salts was introduced into a chambers with coverglass-adhered microtubules, and images were acquired (yellow and magenta circles represent data acquired from each independent experiments; n = 38, 49, 41, 49, and 45 microtubules that span 911 µm, 1074 µm, 1077 µm, 906 µm, 1017 µm in length for each condition, left to right). (c) Pac1-microtubule binding is reduced after enzymatic removal of the unstructured tubulin carboxy-terminal tails (see Methods; similar results were obtained from 2 independent experiments). (d–g) Addition of cell extracts reduces Pac1-microtubule binding, and attenuates Pac1-mediated dynein velocity reduction. Representative fluorescence images of Pac1-SNAP647 on microtubules (d; Pac1-SNAP647 shown as a heat map) and scatter plots depicting intensity values (e; bars depict mean ± standard deviation; n= 48 and 47 microtubules that span 890 µm and 841 µm in length for each condition, left to right; similar results were obtained from 2 independent experiments). (f and g) Plots depicting the motility properties for GST-dyneinMOTOR in the absence and presence of 25 nM Pac1 (dimer concentration) in low ionic strength buffer (50 mM potassium acetate) in the presence of cell extracts (0.96 mg/ml final; 275 and 258 motors, left to right, from 2 independent experiments were quantitated). Note the small Pac1-mediated GST-dyneinMOTOR velocity reduction in the presence of cell extracts (22.1%, compared to 69.5% in the absence of extracts). (h) Additional representative kymograph of GST-dyneinMOTOR comigrating with Pac1 in buffer with 150 mM KCl (see Fig. 6d and Extended Data Fig. 7E – G for quantitation and statistics). Note the diffusive behavior of Pac1 on microtubules. Scale bars in panels a, c and d, 4 µm.
Extended Data Fig. 7 Assessment of dynein-Pac1 stoichiometry, structural analysis of the Pac1-dynein-microtubule complex, and additional motility plots.
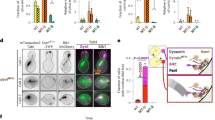
(a and b) Representative kymographs (a) and quantitation (b) of dynein–Pac1/2×Pac1 complex motility (in motility buffer supplemented with 120 mM potassium acetate; n = 870 dynein molecules from 3 independent experiments; mean values with standard error are shown, along all datapoints for middle and right plots; similar results were obtained from each replicate). Statistical significance was determined by calculating Z scores (left; ***, p < 0.0001), using a two-tailed Mann-Whitney test (middle; p = 0.6068), or with a two-tailed Welch’s t test (right; p = 0.6581). Note the estimated fraction of dynein-2xPac1 complexes (5.7%; see main text) is less than what would be expected if there was no cooperativity for Pac1-dynein binding (i.e., the product of the probabilities of two single, independent binding events, 10.8%). (c and d) Structural and cartoon model of a microtubule and Pac1-bound dynein monomer (generated with pdbs 4RH736, 3J1T67, 5VH928, and 3J6G68). Note the close proximity of Pac1 to the microtubule surface, the latter of which is lacking the unstructured E-hooks. (d) Cryo-EM data reveals the dynein-microtubule angle varies due to a hinge point within the MTBD, and can be much steeper than that shown in panel A69,70 (Θ ≥ 15-20°, with average = 55°). Cartoons depict range of angles sampled by dynein on microtubules, and thus the distances between Pac1 and the microtubule. (e - g) Non-normalized plots of mean values (e and f) and all data points (g; see Fig. 6d for n values) showing the relationship between Pac1-mediated dynein velocity reduction and Pac1-microtubule binding (for panels b left, e, and f, diamonds represent mean values obtained from each independent replicate experiment; for panel g, mean values and standard deviations are depicted with red lines).
Supplementary information
Supplementary Table
A list of yeast strains used throughout the study.
Supplementary Video 1
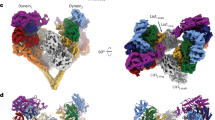
Three-dimensional classification of the full-length yeast dynein phi particle reveals similarities to human dynein. Video of the 3D class average of the autoinhibited yeast dynein conformation. The high-resolution cryo-EM structure of the human dynein phi particle (PDB 5NVU1) was manually docked into the EM density map. Note that the atomic structures of the two tail domains have been slightly rotated with respect to the motor domains to better fit the 3D model, and that the atomic structures of both TcTEX and Robl have been eliminated due to their absence from the yeast dynein complex.
Supplementary Video 2
Uninhibited dynein mutants can bypass the need for Pac1 to move the mitotic spindle. The mitotic spindle (GFP–Tub1) is observed in pac1∆ kar9∆ cells expressing either wild-type (top), K1475E (middle) or D2868K (bottom) dynein. Each frame is a maximum-intensity projection of a 2.4 μm z stack of confocal images collected at intervals of 10 s. The appearance of green circles in the top-left corner of each video corresponds to frames in which dynein-mediated spindle movements are apparent. Note the presence of two spindles in the wild-type cell. Quantification and n values are provided in Fig. 4f.
Supplementary Video 3
Proposed mechanism for autoinhibition restricting dynein processivity. In combination with the tail-dependence of the autoinhibited conformation, our observation of 2D classes with apparent tail–tail contacts — but not motor–motor contacts (Fig. 1, classes vii and viii) — indicates that these contacts may initiate the transition to the autoinhibited state. Subsequent to adopting the autoinhibited conformation, we hypothesize that dynein dissociates from the microtubule.
Source data
Source Data Fig. 1
Uncropped gel from Fig. 1c.
Source Data Fig. 2
Statistical source data for Fig. 2.
Source Data Fig. 3
Statistical source data for Fig. 3.
Source Data Fig. 4
Statistical source data for Fig. 4.
Source Data Fig. 4
Uncropped gels from Fig. 4c.
Source Data Fig. 5
Statistical source data for Fig. 5.
Source Data Extended Data Fig. 2
Statistical source data for Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Statistical source data for Extended Data Fig. 3.
Source Data Extended Data Fig. 4
Statistical source data for Extended Data Fig. 4.
Source Data Extended Data Fig. 4
Uncropped gels from Extended Data Fig. 4b.
Source Data Extended Data Fig. 6
Statistical source data for Extended Data Fig. 6.
Source Data Extended Data Fig. 7
Statistical source data for Extended Data Fig. 7.
Rights and permissions
About this article
Cite this article
Marzo, M.G., Griswold, J.M. & Markus, S.M. Pac1/LIS1 stabilizes an uninhibited conformation of dynein to coordinate its localization and activity. Nat Cell Biol 22, 559–569 (2020). https://doi.org/10.1038/s41556-020-0492-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-0492-1
This article is cited by
-
LIS1 (Pac1) binding slows dissociation of dynein from microtubules
Nature Chemical Biology (2024)
-
Lis1 slows force-induced detachment of cytoplasmic dynein from microtubules
Nature Chemical Biology (2024)
-
TRAK adaptors regulate the recruitment and activation of dynein and kinesin in mitochondrial transport
Nature Communications (2023)
-
Lis1 relieves cytoplasmic dynein-1 autoinhibition by acting as a molecular wedge
Nature Structural & Molecular Biology (2023)
-
Nde1 promotes Lis1-mediated activation of dynein
Nature Communications (2023)