Abstract
Rho GTPases are central regulators of the cytoskeleton and, in humans, are controlled by 145 multidomain guanine nucleotide exchange factors (RhoGEFs) and GTPase-activating proteins (RhoGAPs). How Rho signalling patterns are established in dynamic cell spaces to control cellular morphogenesis is unclear. Through a family-wide characterization of substrate specificities, interactomes and localization, we reveal at the systems level how RhoGEFs and RhoGAPs contextualize and spatiotemporally control Rho signalling. These proteins are widely autoinhibited to allow local regulation, form complexes to jointly coordinate their networks and provide positional information for signalling. RhoGAPs are more promiscuous than RhoGEFs to confine Rho activity gradients. Our resource enabled us to uncover a multi-RhoGEF complex downstream of G-protein-coupled receptors controlling CDC42–RHOA crosstalk. Moreover, we show that integrin adhesions spatially segregate GEFs and GAPs to shape RAC1 activity zones in response to mechanical cues. This mechanism controls the protrusion and contraction dynamics fundamental to cell motility. Our systems analysis of Rho regulators is key to revealing emergent organization principles of Rho signalling.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All data collected and analysed in this study are available at http://the-rhome.com. The protein interactions from this publication (silver dataset) have been submitted to the IMEx (http://www.imexconsortium.org) consortium through IntAct86 and assigned the identifier IM-26436. The MS proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE87 partner repository with the dataset identifiers PXD010084 and PXD010144. cDNA sequences have been submitted to the European Nucleotide Archive (ENA) (accession numbers LS482294–LS482434). All data can also be found at Biostudies:S-BSST160 (ref. 88). Source data for Figs. 1, 2, 4, 6–8 and Extended Data Figs. 2, 4 and 5 are available online. All data supporting the findings of this study are also available from the corresponding authors upon reasonable request.
Code availability
The code used for the filtering of the interactome data is available at https://gitlab.ebi.ac.uk/petsalaki/the-rhome. The code for FRET analysis and FA localization analysis is available at https://github.com/paulmarkusmueller/Mueller_et_al_2020 or from the corresponding authors upon request.
References
Jaffe, A. B. & Hall, A. Rho GTPases: biochemistry and biology. Annu. Rev. Cell Dev. Biol. 21, 247–269 (2005).
Heasman, S. J. & Ridley, A. J. Mammalian Rho GTPases: new insights into their functions from in vivo studies. Nat. Rev. Mol. Cell Biol. 9, 690–701 (2008).
Porter, A. P., Papaioannou, A. & Malliri, A. Deregulation of Rho GTPases in cancer. Small GTPases 7, 123–138 (2016).
Bos, J. L., Rehmann, H. & Wittinghofer, A. GEFs and GAPs: critical elements in the control of small G proteins. Cell 129, 865–877 (2007).
García-Mata, R. & Burridge, K. Catching a GEF by its tail. Trends Cell Biol. 17, 36–43 (2007).
Rossman, K. L., Der, C. J. & Sondek, J. GEF means go: turning on RHO GTPases with guanine nucleotide-exchange factors. Nat. Rev. Mol. Cell Biol. 6, 167–180 (2005).
Tcherkezian, J. & Lamarche-Vane, N. Current knowledge of the large RhoGAP family of proteins. Biol. Cell 99, 67–86 (2007).
DerMardirossian, C. & Bokoch, G. M. GDIs: central regulatory molecules in Rho GTPase activation. Trends Cell Biol. 15, 356–363 (2005).
Garcia-Mata, R., Boulter, E. & Burridge, K. The ‘invisible hand’: regulation of RHO GTPases by RHOGDIs. Nat. Rev. Mol. Cell Biol. 12, 493–504 (2011).
Aspenström, P. Fast-cycling Rho GTPases. Small GTPases https://doi.org/10.1080/21541248.2017.1391365 (2018).
Pertz, O. Spatio-temporal Rho GTPase signaling—where are we now? J. Cell Sci. 123, 1841–1850 (2010).
Benink, H. A. & Bement, W. M. Concentric zones of active RhoA and Cdc42 around single cell wounds. J. Cell Biol. 168, 429–439 (2005).
Machacek, M. et al. Coordination of Rho GTPase activities during cell protrusion. Nature 461, 99–103 (2009).
Bravo-Cordero, J. J. et al. A novel spatiotemporal RhoC activation pathway locally regulates cofilin activity at invadopodia. Curr. Biol. 21, 635–644 (2011).
Fritz, R. D. et al. A versatile toolkit to produce sensitive FRET biosensors to visualize signaling in time and space. Sci. Signal. 6, rs12 (2013).
Graessl, M. et al. An excitable Rho GTPase signaling network generates dynamic subcellular contraction patterns. J. Cell Biol. 216, 4271–4285 (2017).
Guilluy, C., Garcia-Mata, R. & Burridge, K. Rho protein crosstalk: another social network? Trends Cell Biol. 21, 718–726 (2011).
Devreotes, P. & Horwitz, A. R. Signaling networks that regulate cell migration. Cold Spring Harb. Perspect. Biol. 7, a005959 (2015).
Fritz, R. D. et al. SrGAP2-dependent integration of membrane geometry and slit-robo-repulsive cues regulates fibroblast contact inhibition of locomotion. Dev. Cell 35, 78–92 (2015).
Martin, K. et al. Spatio-temporal co-ordination of RhoA, Rac1 and Cdc42 activation during prototypical edge protrusion and retraction dynamics. Sci. Rep. 6, 21901 (2016).
Lawson, C. D. & Ridley, A. J. Rho GTPase signaling complexes in cell migration and invasion. J. Cell Biol. 217, 447–457 (2018).
Cherfils, J. & Zeghouf, M. Regulation of small GTPases by GEFs, GAPs, and GDIs. Physiol. Rev. 93, 269–309 (2013).
Eccles, R. L. et al. Bimodal antagonism of PKA signalling by ARHGAP36. Nat. Commun. 7, 12963 (2016).
Yagi, H. et al. A synthetic biology approach reveals a CXCR4–G13–Rho signaling axis driving transendothelial migration of metastatic breast cancer cells. Sci. Signal. 4, ra60 (2011).
Struckhoff, A. P. et al. PDZ-RhoGEF is essential for CXCR4-driven breast tumor cell motility through spatial regulation of RhoA. J. Cell Sci. 126, 4514–4526 (2013).
Sternweis, P. C. et al. Regulation of Rho guanine nucleotide exchange factors by G proteins. Adv. Protein Chem. 74, 189–228 (2007).
Diviani, D., Soderling, J. & Scott, J. D. AKAP-Lbc anchors protein kinase A and nucleates Gα12-selective Rho-mediated stress fiber formation. J. Biol. Chem. 276, 44247–44257 (2001).
Schliwa, M. Action of cytochalasin D on cytoskeletal networks. J. Cell Biol. 92, 79–91 (1982).
Pascual-Vargas, P. et al. RNAi screens for Rho GTPase regulators of cell shape and YAP/TAZ localisation in triple negative breast cancer. Sci. Data 4, 170018 (2017).
Foussard, H. et al. LRCH proteins: a novel family of cytoskeletal regulators. PLoS ONE 5, e12257 (2010).
Parsons, J. T., Horwitz, A. R. & Schwartz, M. A. Cell adhesion: integrating cytoskeletal dynamics and cellular tension. Nat. Rev. Mol. Cell Biol. 11, 633–643 (2010).
Winograd-Katz, S. E., Fässler, R., Geiger, B. & Legate, K. R. The integrin adhesome: from genes and proteins to human disease. Nat. Rev. Mol. Cell Biol. 15, 273–288 (2014).
Zaidel-Bar, R., Itzkovitz, S., Ma’ayan, A., Iyengar, R. & Geiger, B. Functional atlas of the integrin adhesome. Nat. Cell Biol. 9, 858–867 (2007).
Lundström, A. et al. Vilse, a conserved Rac/Cdc42 GAP mediating Robo repulsion in tracheal cells and axons. Genes Dev. 18, 2161–2171 (2004).
Wong, K. et al. Signal transduction in neuronal migration: roles of GTPase activating proteins and the small GTPase Cdc42 in the Slit–Robo pathway. Cell 107, 209–221 (2001).
Guilluy, C. et al. The Rho GEFs LARG and GEF-H1 regulate the mechanical response to force on integrins. Nat. Cell Biol. 13, 722–727 (2011).
Dubash, A. D. et al. A novel role for Lsc/p115 RhoGEF and LARG in regulating RhoA activity downstream of adhesion to fibronectin. J. Cell Sci. 120, 3989–3998 (2007).
Lim, Y. et al. PyK2 and FAK connections to p190Rho guanine nucleotide exchange factor regulate RhoA activity, focal adhesion formation, and cell motility. J. Cell Biol. 180, 187–203 (2008).
Iwanicki, M. P. et al. FAK, PDZ-RhoGEF and ROCKII cooperate to regulate adhesion movement and trailing-edge retraction in fibroblasts. J. Cell Sci. 121, 895–905 (2008).
Kraynov, V. S. et al. Localized Rac activation dynamics visualized in living cells. Science 290, 333–337 (2000).
Rottner, K., Hall, A. & Small, J. V. Interplay between Rac and Rho in the control of substrate contact dynamics. Curr. Biol. 9, 640–648 (1999).
Lawson, C. D. & Burridge, K. The on-off relationship of Rho and Rac during integrin-mediated adhesion and cell migration. Small GTPases. 5, e27958 (2014).
Rojas, R. J. et al. Gαq directly activates p63RhoGEF and Trio via a conserved extension of the Dbl homology-associated Pleckstrin homology domain. J. Biol. Chem. 282, 29201–29210 (2007).
Canagarajah, B. et al. Structural mechanism for lipid activation of the Rac-specific GAP, beta2-chimaerin. Cell 119, 407–418 (2004).
Mitin, N. et al. Release of autoinhibition of ASEF by APC leads to CDC42 activation and tumor suppression. Nat. Struct. Mol. Biol. 14, 814–823 (2007).
Hall, A. Rho GTPases and the actin cytoskeleton. Science 279, 509–514 (1998).
Breitkreutz, A. et al. A global protein kinase and phosphatase interaction network in yeast. Science 328, 1043–1046 (2010).
St-Denis, N. et al. Phenotypic and interaction profiling of the human phosphatases identifies diverse mitotic regulators. Cell Rep. 17, 2488–2501 (2016).
Jaiswal, M., Dvorsky, R. & Ahmadian, M. R. Deciphering the molecular and functional basis of Dbl family proteins: a novel systematic approach toward classification of selective activation of the Rho family proteins. J. Biol. Chem. 288, 4486–4500 (2013).
Amin, E. et al. Deciphering the molecular and functional basis of RHOGAP family proteins: a systematic approach toward selective inactivation of RHO family proteins. J. Biol. Chem. 291, 20353–20371 (2016).
Nalbant, P., Hodgson, L., Kraynov, V., Toutchkine, A. & Hahn, K. M. Activation of endogenous Cdc42 visualized in living cells. Science 305, 1615–1619 (2004).
Smoly, I., Shemesh, N., Ziv-Ukelson, M., Ben-Zvi, A. & Yeger-Lotem, E. An asymmetrically balanced organization of kinases versus phosphatases across eukaryotes determines their distinct impacts. PLoS Comput. Biol. 13, e1005221 (2017).
Jaiswal, M., Fansa, E. K., Dvorsky, R. & Ahmadian, M. R. New insight into the molecular switch mechanism of human Rho family proteins: shifting a paradigm. Biol. Chem. 394, 89–95 (2013).
ten Klooster, J. P., Jaffer, Z. M., Chernoff, J. & Hordijk, P. L. Targeting and activation of Rac1 are mediated by the exchange factor beta-Pix. J. Cell Biol. 172, 759–769 (2006).
Chang, F., Lemmon, C. A., Park, D. & Romer, L. H. FAK potentiates Rac1 activation and localization to matrix adhesion sites: a role for βPIX. Mol. Biol. Cell 18, 253–264 (2007).
Friedl, P., Sahai, E., Weiss, S. & Yamada, K. M. New dimensions in cell migration. Nat. Rev. Mol. Cell Biol. 13, 743–747 (2012).
Kutys, M. L. & Yamada, K. M. An extracellular-matrix-specific GEF–GAP interaction regulates Rho GTPase crosstalk for 3D collagen migration. Nat. Cell Biol. 16, 909–917 (2014).
Hennig, A., Markwart, R., Esparza-Franco, M. A., Ladds, G. & Rubio, I. Ras activation revisited: role of GEF and GAP systems. Biol. Chem. 396, 831–848 (2015).
Rocks, O. et al. The palmitoylation machinery is a spatially organizing system for peripheral membrane proteins. Cell 141, 458–471 (2010).
Lorentzen, A., Kinkhabwala, A., Rocks, O., Vartak, N. & Bastiaens, P. I. H. Regulation of Ras localization by acylation enables a mode of intracellular signal propagation. Sci. Signal. 3, ra68 (2010).
Schmick, M. et al. KRas localizes to the plasma membrane by spatial cycles of solubilization, trapping and vesicular transport. Cell 157, 459–471 (2014).
Sterpetti, P. et al. Activation of the Lbc Rho exchange factor proto-oncogene by truncation of an extended C terminus that regulates transformation and targeting. Mol. Cell. Biol. 19, 1334–1345 (1999).
Chikumi, H. et al. Homo- and hetero-oligomerization of PDZ-RhoGEF, LARG and p115RhoGEF by their C-terminal region regulates their in vivo Rho GEF activity and transforming potential. Oncogene 23, 233–240 (2004).
Banerjee, J., Fischer, C. C. & Wedegaertner, P. B. The amino acid motif L/IIxxFE defines a novel actin-binding sequence in PDZ-RhoGEF. Biochemistry 48, 8032–8043 (2009).
Kirchner, J., Kam, Z., Tzur, G., Bershadsky, A. D. & Geiger, B. Live-cell monitoring of tyrosine phosphorylation in focal adhesions following microtubule disruption. J. Cell Sci. 116, 975–986 (2003).
Finn, R. D. et al. The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 44, D279–D285 (2016).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Colwill, K. et al. Modification of the Creator recombination system for proteomics applications—improved expression by addition of splice sites. BMC Biotechnol. 6, 13 (2006).
Ethier, M., Hou, W., Duewel, H. S. & Figeys, D. The proteomic reactor: a microfluidic device for processing minute amounts of protein prior to mass spectrometry analysis. J. Proteome Res. 5, 2754–2759 (2006).
O’Leary, N. A. et al. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 44, D733–D745 (2016).
Liu, G. et al. ProHits: integrated software for mass spectrometry-based interaction proteomics. Nat. Biotechnol. 28, 1015–1017 (2010).
Hubner, N. C. et al. Quantitative proteomics combined with BAC TransgeneOmics reveals in vivo protein interactions. J. Cell Biol. 189, 739–754 (2010).
Rappsilber, J., Ishihama, Y. & Mann, M. Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal. Chem. 75, 663–670 (2003).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell. Proteomics 13, 2513–2526 (2014).
Schaefer, M. H. et al. HIPPIE: integrating protein interaction networks with experiment based quality scores. PLoS ONE 7, e31826 (2012).
Mellacheruvu, D. et al. The CRAPome: a contaminant repository for affinity purification-mass spectrometry data. Nat. Methods 10, 730–736 (2013).
Wenger, C. D., Phanstiel, D. H., Lee, M. V., Bailey, D. J. & Coon, J. J. COMPASS: a suite of pre- and post-search proteomics software tools for OMSSA. Proteomics 11, 1064–1074 (2011).
Choi, H. et al. SAINT: probabilistic scoring of affinity purification-mass spectrometry data. Nat. Methods 8, 70–73 (2011).
Yu, G. & He, Q.-Y. ReactomePA: an R/Bioconductor package for reactome pathway analysis and visualization. Mol. Biosyst. 12, 477–479 (2016).
Murtagh, F. & Legendre, P. Ward’s hierarchical agglomerative clustering method: which algorithms implement Ward’s Criterion? J. Classification 31, 274–295 (2014).
Huttlin, E. L. et al. Architecture of the human interactome defines protein communities and disease networks. Nature 545, 505–509 (2017).
Hermjakob, H. IntAct: an open source molecular interaction database. Nucleic Acids Res. 32, 452D–455D (2004).
Wells, C. D., Gutowski, S., Bollag, G. & Sternweis, P. C. Identification of potential mechanisms for regulation of p115 RhoGEF through analysis of endogenous and mutant forms of the exchange factor. J. Biol. Chem. 276, 28897–28905 (2001).
Simpson, J. C., Wellenreuther, R., Poustka, A., Pepperkok, R. & Wiemann, S. Systematic subcellular localization of novel proteins identified by large-scale cDNA sequencing. EMBO Rep. 1, 287–292 (2000).
Orchard, S. et al. The MIntAct project—IntAct as a common curation platform for 11 molecular interaction databases. Nucleic Acids Res. 42, D358–D363 (2013).
Vizcaíno, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44, D447–D456 (2016).
McEntyre, J., Sarkans, U. & Brazma, A. The BioStudies database. Mol. Syst. Biol. 11, 847 (2015).
Acknowledgements
This work is dedicated to the memory of Tony Pawson, without whom this study would not have been initiated. We thank O. Daumke, R. Hodge, D. Panakova and P. Bieling for critically reading the manuscript, R. D. Fritz and K. Rottner for helpful discussions and F.U.P. Cramer for advice. We thank I. Laue, D. Heidler, H. Naumann and the rest of the Rocks Lab and the MDC Advanced Light Microscopy Facility for technical assistance. This work was supported by the Human Frontier Science Program (LT000759/2008-L) and Helmholtz Young Investigator Program VH-NG-737 (to O.R.), the CIHR Post-doctoral fellowship award (to R.D.B.), the Cancer Research UK (CRUK) Programme Foundation Award (C37275/A20146) and the Stand Up to Cancer campaign for Cancer Research UK (to C. Bakal), and Genome Canada through Ontario Genomics, the Ontario Government (ORF GL2-025) and the Terry Fox Research Institute (to T.P.). Proteomics at the Network Biology Collaborative Centre at the Lunenfeld–Tanenbaum Research Institute was supported by Genome Canada and Ontario Genomics (OGI-139). F.P.R. and E.P. were supported by the Canada Excellence Research Chairs Program, the Krembil Foundation, the Avon Foundation and by the NIH/NHGRI Center of Excellence in Genomic Science program (HG004233).
Author information
Authors and Affiliations
Contributions
P.M.M., R.D.B., T.P., E.P. and O.R. conceived the study. Methodologies were developed by the following individuals: P.M.M. (FRET and TIRF screen); P.M.M., C. Barth and O.R. (FA segregation); O.R. and C.W. (confocal screen); R.D.B., K.M.A., V.N., M.S.-C., M.T., O. Popp and B.L. (MS); J.R. (PLEKHG4B study and cytochalasin D screen); J.v.U. (FA biosensor analysis); P.M.M., L.E.H., R.D.B., O.R., C. Bakal and E.P. (computational methods); S.Z., L.S., L.B., G.M., T.R., K.F., J.R., P.T. and O.R. (library). E.P., P.M.M., J.R., R.D.B. and O.R. performed the data analyses. E.P., G.G. and P.M.M. developed the software. R.L.E., L.B., M.T.C., C. Barth, R.-W.W., P.P.-V. and P.T. performed other investigations. E.P., P.M.M., R.D.B., J.R. and O.R. curated the data. O. Pertz and K.C. provided reagents. O.R. wrote the paper (with contributions from E.P. and P.M.M). E.P., F.P.R., P.M., A.-C.G. and K.C. were the project administrators, and O.R. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Domain architecture of the human RhoGEF and RhoGAP proteins.
The canonical isoforms of all 145 human RhoGEFs and RhoGAPs containing all distinguishable domains, were collected from UniProt and clustered by multiple sequence alignment using ClustalOmega. The resulting dendrogram and domain structures as predicted by SMART and Pfam (or by Prosite for ARHGEF37 and ARHGEF38) were assembled using iTol. Not all domain families are listed, non-selected ones are indicated as ‘other’. IGc2, IG and IG_like were summarized as IG. OBSCN is downscaled by a factor of 0.5. Four cDNAs are not included in our collection: OBSCN (~8000a.a.), as well as ARHGEF33, ARHGEF37 and ARHGEF38 which were originally not predicted as RhoGEFs.
Extended Data Fig. 2 Establishment of the FRET-based RhoGEF/RhoGAP activity screen.
a, RhoGDI restores basal activity levels of overexpressed Rho sensors, facilitating RhoGEF assays. Data represents mean ± SD normalized to RhoGDI/sensor vector ratio 0, n=5 FOV of one sample, experiment repeated once with similar results. b, RhoGDI depletion increases the basal activity of the Rho biosensors, facilitating RhoGAP assays. Data represents mean ± SD normalized to WT, n=4 independent experiments. c, Western blot showing shRNA-mediated RhoGDI-knockdown in cell lines used in (b) and (d) compared to virus and non-infected (WT) controls. Representative example out of three experiments with similar results. d, Biosensor response to RhoGAPs is more pronounced in RhoGDI-depleted cells. Cell lines as in (c) were transfected with Rho sensors and RhoGAPs or mCherry (control). Data represents mean ± SD normalized to control of each cell line, n=4 independent experiments. e, FRET ratio is stable across a wide range of sensor expression levels. Cells were transfected with indicated amounts of biosensor vector. mVenus intensity represents relative sensor expression levels. Data represents mean ± SD, n=3 independent experiments. p-values were calculated by unpaired two-sided Student’s t-tests between each of the samples under the line. f and g, Minimal RhoGEF or RhoGAP levels are sufficient to alter the Rho GTPase activity state. HEK293T (f) or RhoGDI-shRNA2 HEK293T (g) cells were cotransfected with Rho sensors and increasing amounts of mCherry-tagged MCF2 (together with 80 ng RhoGDI where indicated, or 80 ng mCherry) or increasing amounts of mCherry-ARHGAP1 or mCherry-ARHGAP22, respectively. mCherry intensities represent relative RhoGEF or RhoGAP expression levels. Data represents mean ± SD normalized to the 0 ng MCF2+RhoGDI sample or normalized to the 0 ng RhoGAP sample, respectively, n=5 FOV of one sample, experiment repeated once with similar results. h, The FRET assay detects only catalytically active RhoGAPs (WT) but not GAP-deficient RK-mutants (ARHGAP4-R543K, ARHGAP11A-R87K, ARHGAP40-R311K, FAM13A-R81K, SYDE2-R854K). Data represents mean ± SD normalized to WT control (mCherry), n=3 independent experiments. i, G-LISA pulldown assay data confirming the substrate specificities found in Fig. 1 for a subset of regulators. RK: ARHGAP23-R986K in lysates. Data represents mean ± SD normalized to YFP, n=3 independent experiments. j, Error-propagation in pulldown assays due to fast GTP hydrolysis. Lysates of transfected HEK293T cells were either processed as fast as possible or with the maximum allowed time according to manual, before processing by Cdc42 GLISA assay. Data represents mean ± SD normalized to YFP, n=3 independent experiments. (a-h) All p-values were calculated by unpaired two-sided Student’s t-test against WT, CONTROL, YFP or 0 ng vector transfected or as indicated by lines and ranked as ***p<0.001, **p<0.01, *p<0.05, n.s.=not significant. Source data including p-values is provided in Source Data Extended Data Fig. 2.
Extended Data Fig. 3 Reactome pathways enrichment analysis and GO Terms enrichment analysis.
a, Pathway enrichment analysis was performed using the ReactomePA function from Bioconductor. Clustering was done using the hclust function in R. b, GO Terms enrichment analysis was performed using the Funcassociate 3.0 web server (http://llama.mshri.on.ca/funcassociate/). Edges between GO terms were calculated using vectors of the genes included in the term and our set and calculating the Jaccard index. Highly redundant nodes were reduced manually to the most informative one for improved visualization (e.g. among the nodes: ‘Cell process’, ‘regulation of cell process’, ‘positive regulation of cell process’, only ‘Cell process’ is kept). Results were visualized using Cytoscape. Our 1292 interactions were used for this enrichment and as background the entire human proteome.
Extended Data Fig. 4 Validation of interactions between RhoGEFs and RhoGAPs identified by mass spectrometry.
To assess the quality of the network dataset, lysates of HEK293T cells transfected with the indicated YFP- and FLAG-tagged regulators or controls were immunoprecipitated (IP) as indicated using either a FLAG or GFP antibody and subsequently immunoblotted using a corresponding GFP or FLAG antibody. Protein bands were detected either by chemiluminescence or using a gel imaging system. 22 out of 26 RhoGEF/RhoGAP pairs tested were successfully validated in two independent repeats. See Source Data_Extended Data Fig. 4 for unprocessed blots.
Extended Data Fig. 5 Characterization of the PLEKHG4B/ARHGEF11/ARHGEF12 multi-RhoGEF complex.
a, PLEKHG4B, ARHGEF11 and ARHGEF12 interact via their N-termini. Immunoprecipitation assays (IP) performed in HEK293T cells expressing YFP-PLEKHG4B together with the indicated full-length or truncated FLAG-ARHGEF11 or FLAG-ARHGEF12 constructs (left panel) or FLAG-ARHGEF11 or FLAG-ARHGEF12 together with the indicated full-length or truncated YFP-PLEKHG4B constructs (right panel). Data are representative of 2 independent experiments. b, PLEKHG4B autoinhibition is released by ARHGEF11 and ARHGEF12. Anti-FLAG Western Blot corresponding to the SRE-luciferase reporter activation data presented in Fig. 4e, left panel, showing the expression of the transfected constructs. Mean ± SD (n=3 independent samples of one experiment, representative out of three experiments with similar results). Significance was determined using One-way ANOVA, followed by Tukey’s multiple comparisons. Significance was ranked as *** p<0.001. Numerical source data including p-values is available online. See Source Data_Extended Data Fig. 5 for unprocessed blots.
Extended Data Fig. 6 Cytochalasin D screen reveals actin-associated RhoGEFs and RhoGAPs.
Confocal images of MDCK cells transiently transfected with YFP fusion constructs of the indicated a, positive (upper panel) or negative (lower panel) control proteins, or b, YFP-tagged RhoGEFs and RhoGAPs. Cells were treated for 30 min with cytochalasin D, fixed and stained for actin with phalloidin. YFP signals were enhanced by anti-GFP immunofluorescence (green: YFP, red: actin in merged images). All 34 actin-associated regulators are displayed. Representative images of two independent experiments (with five images obtained for each experiment) with similar results are shown. Scale bars: 10 μm. The assay not only reliably identified all 12 regulators that we found to colocalize with actin in the primary confocal microscopy screen but also a set of known actin-associated proteins. No coaggregation was observed for negative control proteins known either to be cytosolic or to localize to other compartments. In addition, ARHGEF11, a RhoGEF that associates with actin filaments but is not detectable through microscopy on this structure, was found to coaggregate with actin, while a mutant deficient in actin binding64 (ARHGEF11abm) did not.
Extended Data Fig. 7 RhoGEFs and RhoGAPs provide positional information to Rho signalling regulation.
a, Images from genome-wide siRNA knockdown screen in MDA-MB-231 cells, related to Fig. 6a. Shown are representative examples of abnormal nuclei in cells treated with siRNA against the indicated eleven RhoGEFs/RhoGAPs identified in this study to localize in the nucleus. Experiment was done in quadruplicates. Scale bars: 50 μm. b, Live confocal micrographs of HeLa cells coexpressing EGFR-CFP, mRFP-GRB2 and the indicated YFP-tagged RhoGEFs/RhoGAPs before and 1 min after EGF stimulation (100 ng/ml), related to Fig. 6b. 25 candidate GRB2-interactors were tested: eight regulators identified in our interactome analysis (Supplementary Table 3) and additional proteins listed in the BioGRID database (https://thebiogrid.org/). HeLa cells were chosen because of their robust responsiveness to growth factor stimulation, resulting in an almost complete GRB2 recruitment to the plasma membrane. Only direct interactors of GRB2 co-translocate to the plasma membrane to the same extent. Note, that ARHGEF5 isoform 2, lacking a large N-terminal portion, does not bind GRB2 and remains cytosolic. Representative examples of three independent stimulation experiments with similar results are shown. Scale bars: 10 μm.
Extended Data Fig. 8 C-DOCK subfamily RhoGEFs interact with LRCH family proteins.
a, Domain architecture of all human LRCH family proteins and their isoforms and all eleven DOCK family proteins. Asterisks (*) mark LRCH protein isoforms used in this study. LRR: Leucin rich repeats; CH: Calponin homology; TMR: transmembrane region; SH3: Src homology 3; PH: Pleckstrin homology; DHR: DOCK homology region. b, Live confocal micrographs of MDCK cells expressing all four LRCH proteins (YFP). Note the compromised CH domain in LRCH3 isoform 3 used in this study which may account for its cytosolic localization. c, The A-DOCK family protein DOCK2 (YFP) is not recruited by LRCH2 (CFP) to the periphery of MDCK cells. Live confocal micrographs related to Fig. 6d. d, Live confocal micrographs of MDCK cells coexpressing CFP-DOCK8 and the indicated LRCH proteins and fragments thereof, showing the recruitment of DOCK8 to the endoplasmic reticulum by LRCH1 and LRCH4, or to the cell periphery by LRCH4-ΔTMR. LRCH2-CH, lacking the Leucine rich repeats, cannot recruit DOCK8 to the cell periphery. e, Live confocal micrographs of MDCK cells coexpressing CFP-LRCH1 and the ER marker PTP1B-YFP, revealing their colocalization (related to Fig. 6h). f, Cytochalasin D experiment related to Fig. 6i, revealing actin association of full-length LRCH2. Scale bars: 10 μm. All confocal images are representative of three independent experiments with similar results.
Extended Data Fig. 9 Spatial segregation of RAC1-specific GEFs and GAPs on integrin adhesions in spreading cells.
a, Quantification of RhoGEF/RhoGAP distribution on integrin adhesions. REF52 fibroblasts were transfected with YFP-tagged regulators and the adhesion marker mScarlet-dSH2, together with the plasma membrane marker miRFP-KRas-HVR, or with iRFP-RAC1-Q61L for RAC1-specific GAPs, to balance the GAP phenotype (see (b-e)). Normalized intensity at adhesion complexes is false colour-coded as indicated. Graph shows normalized mean intensity over all pixels of each sampling region ± SD (n=number of pixels in each sampling region) of the example cell on the left. See Methods for details. b, The spreading phenotype induced by RAC1-specific GAPs can be re-balanced by coexpression of low levels of constitutively active RAC1 (RAC1-Q61L). Dominant negative RAC1 (RAC1-T17N) causes a spreading phenotype similar to RAC1-specific GAPs. REF52 cells were transfected with YFP-tagged Paxillin control, the exemplary RAC1-specific GAPs ARHGAP22, CHN2 or SYDE2, or dominant negative RAC1 (RAC1-T17N)), together with mScarlet-dSH2 and miRFP-KRas-HVR (control, left panel) or iRFP-RAC1-Q61L (right panel). Experiment was repeated three times with similar results. c, Expression of RAC1-Q61L does not alter the relative distribution of actin, paxillin and phospho-tyrosine on integrin adhesions in isotropically spreading cells. REF52 cells were transfected with mEGFP-LifeAct or mEGFP-Paxillin, together with mScarlet-dSH2 (phospho-tyrosine adhesion marker) and miRFP-KRas-HVR (control) or iRFP-RAC1-Q61L. d, Expression of RAC1-Q61L does not alter the relative distribution of ARHGAP9 in isotropically spreading cells. ARHGAP9 is a RAC1-specific GAP showing only a mild spreading phenotype. REF52 cells in (b), (c), and (d) were treated as in (a). n in (c) and (d) is given as number of analyzed cells inside the graph. e, Expression of RAC1-GEFs, or of RAC1-GAPs together with RAC1-Q61L, does not alter the relative distribution of dSH2 on integrin adhesions in isotropically spreading cells. Left panel: Quantification of GEF/GAP distributions as shown in Fig. 7e, right panel: corresponding distributions of dSH2. Means of n=9–23 cells from one experiment are shown (for details on n see Supplementary Information Fig. 3). All scale bars: 10 µm. Boxplot centre lines in (c) and (d) represent the median values, box limits the 25th and 75th percentiles; whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles.
Extended Data Fig. 10 RhoGEF/RhoGAP re-distribution on focal adhesions in isotropically spreading cells upon Y-27632 addition.
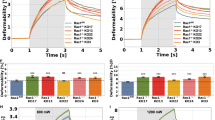
Timelapse images showing RhoGEF/RhoGAP re-distribution on adhesions in isotropically spreading cells upon Y-27632 addition (corresponding to Supplementary Videos 3–6). REF52 cells were treated as in Fig. 7d. Left panel: representative timepoints before and 30 min after addition of inhibitor. Right panel: kymographs of boxed regions with the cell edge marked in red. Note, that the RAC1-specific GEF ARHGEF6, as well as the RHOA-specific GAPs DLC1 and STARD13, localize to early nascent adhesions after Y-27632 treatment (as indicated by dSH2 close to the cell edge), whilst the RAC1-specific GAP SYDE2 does not. Data shown represent four, five, three and four independent experiments for ARHGEF6, DLC1, SYDE2 and STARD13, respectively. All scale bars: 10 µm.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3, Supplementary Notes and Supplementary Methods.
Supplementary Tables
Supplementary Table 1: Information on the RhoGEF and RhoGAP cDNAs included in our cDNA library. Detailed description of the RhoGEF and RhoGAP cDNA expression library, including information on synonyms, Human Ensembl and Entrez GeneID, species in library, Protein ID, size of construct in the library, mCitrine-YFP expression construct ID, cDNA source and cDNA sequence. Supplementary Table 2: RhoGEF and RhoGAP specificities identified in this study and in the literature. Data from FRET biosensor screen and from literature review are shown, including information about other substrate GTPases and reference PubMed IDs. Due to the high degree of conflicting literature data, four separate lists were compiled with different emphases: ‘integrated’, ‘in vitro’, ‘in vivo’ and ‘reference’, see Methods for detail. Supplementary Table 3: The RhoGEF and RhoGAP interactome. The gold, silver and bronze interactome lists are included. All baits were ran at least four times (Citrine and 3-FLAG-tagged, transient transfection and stable expression ×2 replicates). The samples ran in the Q Exactive HF-X instrument only had 2 replicates and were therefore analysed differently (see Methods). The precise number of samples used for each bait can be found in Biostudies:S-BSST160, the raw data table in the-rhome.com database and in the raw MS files deposited in PRIDE. The sample number in which each interaction was found is in column G of this table. Supplementary Table 4: Localization reported in the literature and determined in this study. Overview of RhoGEF and RhoGAP subcellular localization screen including data from confocal, TIRF and cytochalasin D screen with additional notes and data from literature review with reference Pubmed IDs. Supplementary Table 5: FA-associated RhoGEFs and RhoGAPs: substrate specificities and notes. Overview of the 37 FA-associated RhoGEFs and RhoGAPs identified in this study, including their substrate specificities determined in the FRET biosensor screen, specificites reported in the literature and further notes.
Supplementary Video 1
Y-27632 treatment of YFP-DOCK3-expressing REF52 cells. The experiment was repeated four times with similar results. This video is associated with Fig. 8a.
Supplementary Video 2
Y-27632 treatment of YFP-ARHGAP31-expressing REF52 cells. The experiment was repeated six times with similar results. This video is associated with Fig. 8b.
Supplementary Video 3
Y-27632 treatment of YFP-ARHGEF6-expressing REF52 cells. The experiment was repeated four times with similar results. This video is associated with Extended Data Fig. 10.
Supplementary Video 4
Y-27632 treatment of YFP-SYDE1-expressing REF52 cells. The experiment was repeated three times with similar results. This video is associated with Extended Data Fig. 10.
Supplementary Video 5
Y-27632 treatment of YFP-DLC1-expressing REF52 cells. The experiment was repeated five times with similar results. This video is associated with Extended Data Fig. 10.
Supplementary Video 6
Y-27632 treatment of YFP-STARD13-expressing REF52 cells. The experiment was repeated four times with similar results. This video is associated with Extended Data Fig. 10.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 6
Unprocessed western blots.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Müller, P.M., Rademacher, J., Bagshaw, R.D. et al. Systems analysis of RhoGEF and RhoGAP regulatory proteins reveals spatially organized RAC1 signalling from integrin adhesions. Nat Cell Biol 22, 498–511 (2020). https://doi.org/10.1038/s41556-020-0488-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-0488-x
This article is cited by
-
Patterning of the cell cortex by Rho GTPases
Nature Reviews Molecular Cell Biology (2024)
-
Extracellular matrix stiffness mediates uterine repair via the Rap1a/ARHGAP35/RhoA/F-actin/YAP axis
Cell Communication and Signaling (2023)
-
The multiple links between actin and mitochondria
Nature Reviews Molecular Cell Biology (2023)
-
Rho GTPase activity crosstalk mediated by Arhgef11 and Arhgef12 coordinates cell protrusion-retraction cycles
Nature Communications (2023)
-
CNK2 promotes cancer cell motility by mediating ARF6 activation downstream of AXL signalling
Nature Communications (2023)