Abstract
During endoplasmic-reticulum-associated protein degradation (ERAD), misfolded proteins are polyubiquitinated, extracted from the ER membrane and degraded by the proteasome1,2,3,4. In a process called retrotranslocation, misfolded luminal proteins first need to traverse the ER membrane before ubiquitination can occur in the cytosol. It was suggested that the membrane-embedded ubiquitin ligase Hrd1 forms a retrotranslocation pore regulated by cycles of auto- and deubiquitination5,6,7,8. However, the mechanism by which auto-ubiquitination affects Hrd1 and allows polypeptides to cross the membrane and whether Hrd1 forms a membrane-spanning pore remained unknown. Here, using purified Hrd1 incorporated into different model membranes, we show that Hrd1 auto-ubiquitination leads to the opening of a pore. Substrate binding increases the pore size and its activity, whereas deubiquitination closes the pore and renders it unresponsive to substrate. We identify two binding sites for misfolded proteins in Hrd1, a low-affinity luminal site and a high-affinity cytoplasmic site formed following auto-ubiquitination of specific lysine residues in Hrd1’s RING domain. We propose that the affinity difference between the luminal and cytoplasmic binding sites provides the initial driving force for substrate movement through Hrd1.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Christianson, J. C. & Ye, Y. Cleaning up in the endoplasmic reticulum: ubiquitin in charge. Nat. Struct. Mol. Biol. 21, 325–335 (2014).
Ruggiano, A., Foresti, O. & Carvalho, P. Quality control: ER-associated degradation: protein quality control and beyond. J. Cell Biol. 204, 869–879 (2014).
Mehrtash, A. B. & Hochstrasser, M. Ubiquitin-dependent protein degradation at the endoplasmic reticulum and nuclear envelope. Semin. Cell Dev. Biol. 93, 111–124 (2019).
Wu, X. & Rapoport, T. A. Mechanistic insights into ER-associated protein degradation. Curr. Opin. Cell Biol. 53, 22–28 (2018).
Carvalho, P., Stanley, A. M. & Rapoport, T. A. Retrotranslocation of a misfolded luminal ER protein by the ubiquitin-ligase Hrd1p. Cell 143, 579–591 (2010).
Stein, A., Ruggiano, A., Carvalho, P. & Rapoport, T. A. Key steps in ERAD of luminal ER proteins reconstituted with purified components. Cell 158, 1375–1388 (2014).
Baldridge, R. D. & Rapoport, T. A. Autoubiquitination of the Hrd1 ligase triggers protein retrotranslocation in ERAD. Cell 166, 394–407 (2016).
Peterson, B. G., Glaser, M. L., Rapoport, T. A. & Baldridge, R. D. Cycles of autoubiquitination and deubiquitination regulate the ERAD ubiquitin ligase Hrd1. eLife 8, e50903 (2019).
Hampton, R. Y., Gardner, R. G. & Rine, J. Role of 26S proteasome and HRD genes in the degradation of 3-hydroxy-3-methylglutaryl-CoA reductase, an integral endoplasmic reticulum membrane protein. Mol. Biol. Cell 7, 2029–2044 (1996).
Knop, M., Finger, A., Braun, T., Hellmuth, K. & Wolf, D. H. Der1, a novel protein specifically required for endoplasmic reticulum degradation in yeast. EMBO J. 15, 753–763 (1996).
Bordallo, J., Plemper, R. K., Finger, A. & Wolf, D. H. Der3p/Hrd1p is required for endoplasmic reticulum-associated degradation of misfolded lumenal and integral membrane proteins. Mol. Biol. Cell 9, 209–222 (1998).
Bays, N. W., Gardner, R. G., Seelig, L. P., Joazeiro, C. A. & Hampton, R. Y. Hrd1p/Der3p is a membrane-anchored ubiquitin ligase required for ER-associated degradation. Nat. Cell Biol. 3, 24–29 (2001).
Gauss, R., Jarosch, E., Sommer, T. & Hirsch, C. A complex of Yos9p and the HRD ligase integrates endoplasmic reticulum quality control into the degradation machinery. Nat. Cell Biol. 8, 849–854 (2006).
Carvalho, P., Goder, V. & Rapoport, T. A. Distinct ubiquitin-ligase complexes define convergent pathways for the degradation of ER proteins. Cell 126, 361–373 (2006).
Kanehara, K., Xie, W. & Ng, D. T. Modularity of the Hrd1 ERAD complex underlies its diverse client range. J. Cell Biol. 188, 707–716 (2010).
Mehnert, M., Sommer, T. & Jarosch, E. Der1 promotes movement of misfolded proteins through the endoplasmic reticulum membrane. Nat. Cell Biol. 16, 77–86 (2014).
Hiller, M. M., Finger, A., Schweiger, M. & Wolf, D. H. ER degradation of a misfolded luminal protein by the cytosolic ubiquitin-proteasome pathway. Science 273, 1725–1728 (1996).
Biederer, T., Volkwein, C. & Sommer, T. Role of Cue1p in ubiquitination and degradation at the ER surface. Science 278, 1806–1809 (1997).
Wilhovsky, S., Gardner, R. & Hampton, R. HRD gene dependence of endoplasmic reticulum-associated degradation. Mol. Biol. Cell 11, 1697–1708 (2000).
Twomey, E. C. et al. Substrate processing by the Cdc48 ATPase complex is initiated by ubiquitin unfolding. Science 365, eaax1033 (2019).
Bays, N. W., Wilhovsky, S. K., Goradia, A., Hodgkiss-Harlow, K. & Hampton, R. Y. HRD4/NPL4 is required for the proteasomal processing of ubiquitinated ER proteins. Mol. Biol. Cell 12, 4114–4128 (2001).
Neuber, O., Jarosch, E., Volkwein, C., Walter, J. & Sommer, T. Ubx2 links the Cdc48 complex to ER-associated protein degradation. Nat. Cell Biol. 7, 993–998 (2005).
Jarosch, E. et al. Protein dislocation from the ER requires polyubiquitination and the AAA-ATPase Cdc48. Nat. Cell Biol. 4, 134–139 (2002).
Ye, Y., Meyer, H. H. & Rapoport, T. A. The AAA ATPase Cdc48/p97 and its partners transport proteins from the ER into the cytosol. Nature 414, 652–656 (2001).
Schuberth, C. & Buchberger, A. Membrane-bound Ubx2 recruits Cdc48 to ubiquitin ligases and their substrates to ensure efficient ER-associated protein degradation. Nat. Cell Biol. 7, 999–1006 (2005).
Plemper, R. K. et al. Genetic interactions of Hrd3p and Der3p/Hrd1p with Sec61p suggest a retro-translocation complex mediating protein transport for ER degradation. J. Cell Sci. 112, 4123–4134 (1999).
Finger, A., Knop, M. & Wolf, D. H. Analysis of two mutated vacuolar proteins reveals a degradation pathway in the endoplasmic reticulum or a related compartment of yeast. Eur. J. Biochem. 218, 565–574 (1993).
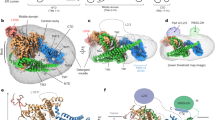
Schoebel, S. et al. Cryo-EM structure of the protein-conducting ERAD channel Hrd1 in complex with Hrd3. Nature 548, 352–355 (2017).
Meinecke, M. et al. Tim50 maintains the permeability barrier of the mitochondrial inner membrane. Science 312, 1523–1526 (2006).
Saparov, S. M. et al. Determining the conductance of the SecY protein translocation channel for small molecules. Mol. Cell 26, 501–509 (2007).
Truscott, K. N. et al. A presequence- and voltage-sensitive channel of the mitochondrial preprotein translocase formed by Tim23. Nat. Struct. Biol. 8, 1074–1082 (2001).
Hospenthal, M. K., Mevissen, T. E. T. & Komander, D. Deubiquitinase-based analysis of ubiquitin chain architecture using Ubiquitin Chain Restriction (UbiCRest). Nat. Protoc. 10, 349–361 (2015).
Meinecke, M. et al. The peroxisomal importomer constitutes a large and highly dynamic pore. Nat. Cell Biol. 12, 273–277 (2010).
Wirth, A. et al. The Sec61p complex is a dynamic precursor activated channel. Mol. Cell 12, 261–268 (2003).
Spear, E. D. & Ng, D. T. Single, context-specific glycans can target misfolded glycoproteins for ER-associated degradation. J. Cell Biol. 169, 73–82 (2005).
Vashistha, N., Neal, S. E., Singh, A., Carroll, S. M. & Hampton, R. Y. Direct and essential function for Hrd3 in ER-associated degradation. Proc. Natl Acad. Sci. USA 113, 5934–5939 (2016).
Neal, S. et al. The Dfm1 derlin is required for ERAD retrotranslocation of integral membrane proteins. Mol. Cell 69, 306–320 (2018).
Horn, S. C. et al. Usa1 functions as a scaffold of the HRD-ubiquitin ligase. Mol. Cell 36, 782–793 (2009).
Gardner, R. G. et al. Endoplasmic reticulum degradation requires lumen to cytosol signaling. Transmembrane control of Hrd1p by Hrd3p. J. Cell Biol. 151, 69–82 (2000).
Carroll, S. M. & Hampton, R. Y. Usa1p is required for optimal function and regulation of the Hrd1p endoplasmic reticulum-associated degradation ubiquitin ligase. J. Biol. Chem. 285, 5146–5156 (2010).
Popp, M. W. & Ploegh, H. L. Making and breaking peptide bonds: protein engineering using sortase. Angew. Chem. Int. Ed. Engl. 50, 5024–5032 (2011).
Chen, I., Dorr, B. M. & Liu, D. R. A general strategy for the evolution of bond-forming enzymes using yeast display. Proc. Natl Acad. Sci. USA 108, 11399–11404 (2011).
Hernandez, J. M. et al. Membrane fusion intermediates via directional and full assembly of the SNARE complex. Science 336, 1581–1584 (2012).
Bello, O. D., Auclair, S. M., Rothman, J. E. & Krishnakumar, S. S. Using ApoE nanolipoprotein particles to analyze SNARE-induced fusion pores. Langmuir 32, 3015–3023 (2016).
Harsman, A., Bartsch, P., Hemmis, B., Kruger, V. & Wagner, R. Exploring protein import pores of cellular organelles at the single molecule level using the planar lipid bilayer technique. Eur. J. Cell Biol. 90, 721–730 (2011).
Denkert, N. et al. Cation selectivity of the presequence translocase channel Tim23 is crucial for efficient protein import. eLife 6, e28324 (2017).
Reinhold, R. et al. The channel-forming Sym1 protein is transported by the TIM23 complex in a presequence-independent manner. Mol. Cell. Biol. 32, 5009–5021 (2012).
Cohen, F. S., Niles, W. D. & Akabas, M. H. Fusion of phospholipid vesicles with a planar membrane depends on the membrane permeability of the solute used to create the osmotic pressure. J. Gen. Physiol. 93, 201–210 (1989).
Smart, O. S., Breed, J., Smith, G. R. & Sansom, M. S. A novel method for structure-based prediction of ion channel conductance properties. Biophys. J. 72, 1109–1126 (1997).
Hodgkin, A. L. & Katz, B. The effect of sodium ions on the electrical activity of giant axon of the squid. J. Physiol. 108, 37–77 (1949).
Hotz, T. et al. Idealizing ion channel recordings by a jump segmentation multiresolution filter. IEEE Trans. Nanobioscience 12, 376–386 (2013).
Acknowledgements
We thank O. D. Bello and J. E. Rothman for providing the construct for ApoE422K, I. Bickmeyer and N. Nupur for technical assistance and T. Rapoport and R. Jahn for comments on the manuscript. This work was supported by the European Research Council (ERC) under the Horizon2020 research and innovation programme (grant no. 677770) to A.S., by the Deutsche Forschungsgemeinschaft SFB1190, grant nos. P12 (to M.M.) and P15 (to A.S.), and a Boehringer Ingelheim Fonds PhD Fellowship (to V.V.).
Author information
Authors and Affiliations
Contributions
A.S. and M.M. conceived the experiments and wrote the manuscript. V.V., N.D., D.R. and A.S. carried out the experiments: N.D. performed the electrophysiology; V.V. and A.S. performed the biochemistry; C.C.S. provided reagents and established the Ubc6-related experiments; and D.R. performed the electron microscopy. All authors contributed to data analysis.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of Hrd1 liposomes and channel properties.
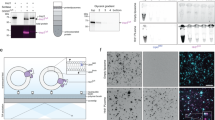
a, Liposomes containing C-terminally fluorescently labelled Hrd1 were floated in a Nycodenz step gradient. The gradient was fractionated and samples were analyzed by SDS-PAGE and fluorescence scanning. b, Fluorescently labeled Hrd1 in liposomes were incubated with Tobacco Etch Virus (TEV) protease that cleaves off the C-terminal SBP tag and the fluorescent dye. As a control, detergent-solubilized liposomes were incubated with TEV protease. Samples were analyzed by SDS PAGE and fluorescence scanning. c, Conductance state histogram zoom plot of Fig. 1e. The arrow and number indicate the highest observed conductance state for ubiquitinated Hrd1. To focus on large conductance states, the zoom plot starts at 100 pS. d, Current-voltage relationship of ubiquitinated Hrd1 at asymmetric salt. Arrows indicate the various reversal potentials and the numbers give the corresponding relative selectivities of potassium over chloride as calculated from the Goldman-Hodgkin-Katz equation. The red line represents linear regression (least-squares) of indicated data regions (length of red line on x-axis). Shown is a representative trace of three independent experiments. e, Time course of deubiquitination of Hrd1 using indicated concentrations of Usp2. Liposomes containing fluorescently labeled Hrd1 were immobilized onto streptavidin magnetic beads and incubated with ubiquitination mix in the presence or absence of ATP. After washing, Hrd1 liposomes were eluted with 2 mM biotin and incubated with the indicated amount of Usp2. The reaction was stopped by addition of SDS sample buffer. Samples were analyzed by SDS-PAGE and fluorescence scanning. This particular experiment was performed once. Other related deubiquitination experiments are shown in Fig. 3a and Extended Data Fig. 3a. In a-b, representative images of three independent experiments are shown. Source data and unprocessed gels are provided in Source Data Extended Data Fig. 1.
Extended Data Fig. 2 Interaction of misfolded substrates with Hrd1 reconstituted in liposomes or nanodiscs.
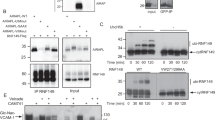
a, PrA*, but not PrA WT interacts with ubiquitinated Hrd1. Indicated amounts of Hrd1 liposomes were immobilized onto streptavidin magnetic beads via the C-terminal SBP tag on Hrd1. After incubation with ubiquitination mix in the presence or absence of ATP, beads were washed and incubated with 20 nM PrA* or 20 nM PrA WT at the indicated Hrd1 concentrations. The fraction of bound PrA* or PrA WT was determined from the supernatants. mean ± s.d (n = 3 independent experiments). b, Negative stain electron micrographs of glutaraldehyde-fixed Hrd1 nanodiscs. c, Size distribution from n = 738 Hrd1 nanodiscs. d, CPY*, but not CPY WT is efficiently ubiquitinated when added to the outside of Hrd1 in liposomes. Left: Fluorescently labelled CPY* or CPY WT (100 nM) was added to liposomes containing Hrd1 (200 nM) and incubated with ubiquitination mix with or without ATP. Samples from indicated time points were analyzed by SDS-PAGE and fluorescence scanning. Right: Quantification of three ubiquitination experiments. These data are also presented in Fig. 3h (CPY* lipos) and Extended Data Fig. 3c, d (CPY WT, WT Hrd1). mean ± s.d (n = 3 independent experiments). e, PrA*, but not PrA WT is efficiently ubiquitinated when added to the outside of Hrd1 in liposomes. Left: Fluorescently labelled PrA* or PrA WT (100 nM) was added to liposomes containing Hrd1 (200 nM) and incubated with ubiquitination mix with or without ATP. Samples from indicated time points were analyzed by SDS-PAGE and fluorescence scanning. Right: Quantification of three ubiquitination experiments. mean ± s.d (n = 3 independent experiments). Source data and unprocessed gels are provided in Source Data Extended Data Fig. 2.
Extended Data Fig. 3 Characterization of the interaction of CPY* with autoubiquitinated Hrd1.
a, Release of CPY* from liposomes containing ubiquitinated Ubc6 or Hrd1 upon deubiquitination. Beads with immobilized liposomes containing ubiquitinated Hrd1 (250 nM) or ubiquitinated Ubc6 (250 nM) were incubated with CPY* (50 nM). After washing, an aliquot of beads were incubated with SDS sample buffer to determine the total bound CPY*. Usp2 (3 µM) was added to the beads and the supernatant was collected. The Usp2 treated samples were then eluted with SDS sample buffer. Samples were analyzed by SDS PAGE and fluorescence scanning. In: CPY* input to the beads, Unb: unbound CPY* after incubation with ubiquitinated Hrd1 or Ubc6. b, Quantification (mean ± s.d.) of the fraction of CPY* released upon deubiquitination by Usp2 from three experiments as in a. c, CPY WT (100 nM) was incubated with fluorescently labeled WT Hrd1 or indicated mutants in liposomes (200 nM), and ubiquitination mix with or without ATP. Samples from indicated time points were analyzed by SDS-PAGE and fluorescence scanning. d, Quantification (mean ± s.d.) of three experiments as in c. e, Increasing concentrations of ubiquitinated, bead-immobilized WT Hrd1 or indicated Hrd1 mutants in liposomes were incubated with fluorescently labeled CPY WT (20 nM). The bound fraction was quantified from supernatants. mean ± s.d (n = 3 independent experiments). f, Liposomes containing fluorescently labeled Hrd1 were incubated with ubiquitination mix containing the ubiquitin K48R mutant. Samples were analyzed by SDS PAGE and fluorescence scanning. Shown is a representative image of two independent experiments. g, Bead-immobilized Hrd1 liposomes were incubated with ubiquitination mix containing WT or K48R ubiquitin with or without ATP. Beads were then subsequently incubated with fluorescently labelled CPY* (40 nM). Samples were analyzed by SDS PAGE and fluorescence scanning. h, Quantification (mean ± s.d.) of three experiments as in g. i, Constant-voltage recordings of ubiquitinated Hrd1 KRK mutant at indicated voltages in the absence (left) and presence of CPY* (right). Shown are representative traces of three independent experiments. Source data and unprocessed gels are provided in Source Data Extended Data Fig. 3.
Extended Data Fig. 4 Molecular mechanism for Hrd1-dependent retrotranslocation of misfolded proteins from the ER lumen to the cytosol.
A Misfolded substrate binds to the luminal face of Hrd1 (1). Hrd1 auto-ubiquitination opens the retrotranslocation pore (2) which is further expanded by substrate insertion. A high-affinity binding site on the cytoplasmic face of Hrd1 drives initial substrate translocation (3). The substrate is ubiquitinated by Hrd1 on the cytoplasmic side of the membrane and recruits the Cdc48 complex (4). The Cdc48 complex segregates substrate and Hrd1, and extracts the ubiquitinated substrate from the membrane through rounds of ATP hydrolysis. Hrd1 is deubiquitinated by a DUB, closing the retrotranslocation pore (5).
Supplementary Information
Source data
Source Data Fig. 1
Statistical source data
Source Data Fig. 1
Unprocessed gels
Source Data Fig. 2
Statistical source data
Source Data Fig. 3
Statistical source data
Source Data Fig. 3
Unprocessed gels
Source Data Fig. 4
Statistical source data
Source Data Fig. 4
Unprocessed gels
Source Data Extended Data Fig. 1
Statistical source data
Source Data Extended Data Fig. 1
Unprocessed gels
Source Data Extended Data Fig. 2
Statistical source data
Source Data Extended Data Fig. 2
Unprocessed gels
Source Data Extended Data Fig. 3
Statistical source data
Source Data Extended Data Fig. 3
Unprocessed gels
Rights and permissions
About this article
Cite this article
Vasic, V., Denkert, N., Schmidt, C.C. et al. Hrd1 forms the retrotranslocation pore regulated by auto-ubiquitination and binding of misfolded proteins. Nat Cell Biol 22, 274–281 (2020). https://doi.org/10.1038/s41556-020-0473-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-0473-4
This article is cited by
-
Direct observation of autoubiquitination for an integral membrane ubiquitin ligase in ERAD
Nature Communications (2024)
-
Casting Light on the Janus-Faced HMG-CoA Reductase Degradation Protein 1: A Comprehensive Review of Its Dualistic Impact on Apoptosis in Various Diseases
Molecular Neurobiology (2024)
-
Mechanisms of substrate processing during ER-associated protein degradation
Nature Reviews Molecular Cell Biology (2023)
-
Involvement of Proteasomal and Endoplasmic Reticulum Stress in Neurodegeneration After Global Brain Ischemia
Molecular Neurobiology (2023)
-
A positive genetic selection for transmembrane domain mutations in HRD1 underscores the importance of Hrd1 complex integrity during ERAD
Current Genetics (2022)